Abstract
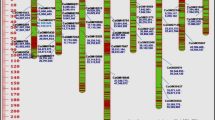
Chickpea (Cicer arietinum L.) is one of the most important legumes worldwide. We addressed this study to the genetic characterization of a germplasm collection from main chickpea growing countries. Several Italian traditional landraces at risk of genetic erosion were included in the analysis. Twenty-two simple sequence repeat (SSR) markers, widely used to explore genetic variation in plants, were selected and yielded 218 different alleles. Structure analysis and hierarchical clustering indicated that a model with three distinct subpopulations best fits the data. The composition of two subpopulations, named K1 and K2, broadly reflects the commercial classification of chickpea in the two types desi and kabuli, respectively. The third subpopulation (K3) is composed by both desi and kabuli genotypes. Italian accessions group both in K2 and K3. Interestingly, this study highlights genetic distance between desi genotypes cultivated in Asia and Ethiopia, which respectively represent the chickpea primary and the secondary centres of diversity. Moreover, European desi are closer to the Ethiopian gene pool. Overall, this study will be of importance for chickpea conservation genetics and breeding, which is limited by the poor characterization of germplasm collection.


Similar content being viewed by others
References
Abbo S, Berger J, Turner NC (2003) Evolution of cultivated chickpea: four bottlenecks limit diversity and constrain adaptation. Funct Plant Biol 30:1081–1087
Amin M, Melkamu F (2014) Management of Ascochyta Blight (Ascochyta rabiei) in chickpea using a new fungicide. Res Plant Sci 2:27–32
Choudhary S, Sethy NK, Shokeen B, Bhatia S (2006) Development of sequence tagged microsatellite site markers for chickpea (Cicer arietinum L.). Mol Ecol Notes 6:93–95
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Faostat (2013) Agriculture data. http://faostat.fao.org/site/567/default.aspx# ancor. Accessed Nov 2013
Fernández M, Figueiras A, Benito C (2002) The use of ISSR and RAPD markers for detecting DNA polymorphism, genotype identification and genetic diversity among barley cultivars with known origin. TAG Theor Appl Genet 104(5):845–851
Hajibarat Z, Saidi A, Hajibarat Z, Taleb R (2015) Characterization of genetic diversity in chickpea using SSR markers, Start Codon Targeted Polymorphism (SCoT) and Conserved DNA-Derived Polymorphism (CDDP). Physiol Mol Biol Plant 21:365–373
Hüttel B, Winter P, Weising K, Choumane W, Weigand F, Kahl G (1999) Sequence-tagged microsatellite site markers for chickpea (Cicer arietinum L.). Genome 42:210–217
Jamalabadi JG, Saidi A, Karami A, Kharkesh M, Talebi R (2013) Molecular Mapping and Characterization of Genes Governing Time to Flowering, Seed Weight, and Plant Height in an Intraspecific Genetic Linkage Map of Chickpea (Cicer arietinum). Biochem Genet 51(5–6):387–397
Kassie M, Shiferaw B, Asfaw S, Abate T, Geoffrey M, Setotaw F, Eshete M, Assefa K (2009) Current situation and future outlooks of the chickpea sub-sector in Ethiopia. ICRISAT working paper. ICRISAT and EIAR. 39 pp. www.icrisat.org/tropicallegumesII
Khajuria YP, Saxena MS, Gaur R, Chattopadhyay D, Jain M, Parida SK, Bhatia S (2015) Development and integration of genome-wide polymorphic microsatellite markers onto a reference linkage map for constructing a high-density genetic map of chickpea. PLoS ONE. doi:10.1371/journal.pone.0125583
Kloosterman AD, Budowle B, Daselaar P (1993) PCR amplification and detection of the human D1S80 VNTR locus: amplification conditions, population genetics and application forensic analysis. Int J Leg Med 105:257–264
Lichtenzveig J, Scheuring C, Dodge J, Abbo S, Zhang HB (2005) Construction of BAC and BIBAC libraries and their applications for generation of SSR markers for genome analysis of chickpea, Cicer arietinum L. Theor Appl Genet 110:492–510
Nayak SN, Zhu H, Varghese N, Datta S, Choi HK, Horres R, Jüngling R, Singh J, Kishor PB, Sivaramakrishnan S, Hoisington DA, Kahl G, Winter P, Cook DR, Varshney RK (2010) Integration of novel SSR and gene-based SNP marker loci in the chickpea genetic map and establishment of new anchor points with Medicago truncatula genome. Theor Appl Genet 120:1415–1441
Negri V (2003) Landraces in central Italy: where and why they are conserved and perspectives for their on-farm conservation. Genet Resour Crop Evol 50:877–888
Nybom H, Weising K, Rotter B (2014) DNA fingerprinting in botany: past, present, future. Investig Genet 5(1):1. doi:10.1186/2041-2223-5-1
Parida SK, Verma M, Yadav SK, Ambawat S, Das S, Garg R, Jain M (2015) Development of genome-wide informative simple sequence repeat markers for large scale genotyping application sin chickpea and development of web resource. Front in Plant Sci 6(645):1–12
Patanè C (2006) Variation and relationships among some nutritional traits in Sicilian genotypes of chickpea (Cicer arietinum L.). J Food Qual 29:282–293
Penmetsa VR, Carrasquilla-Garcia N, Bergmann EM, Vance L, Castro B, Kassa MT, Sarma BK, Datta S, Farmer AD, Baek JM, Coyne CJ, Varshney RK, von Wettberg EJB, Cook DR (2016) Multiple post-domestication origins of Kabuli chickpea through allelic variation in a diversification-associated transcription factor. New Phytol 211:1440–1451
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98:107–112
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rohlf FJ (1998) NTSYS-pc. Numerical taxonomy and multivariate analysis system, version 2.02. Exeter Software, New York
Saxena MS, Bajaj D, Kujur A, Das S, Badoni S, Kumar V, Singh M, Bansal KC, Tyagi AK, Parida SK (2014) Natural allelic diversity, genetic structure and linkage disequilibrium pattern in wild chickpea. PLoS One. doi:10.1371/journal.pone.0107484
Sethy NK, Shokeen B, Edwards KJ, Bhatia S (2006) Development of microsatellite markers and analysis of intraspecific genetic variability in chickpea (Cicer arietinum L.). Theor Appl Genet 112:1416–1428
Singh R, Prasad CD, Singhal V, Randhawa GJ (2003) Assessment of genetic diversity in chickpea cultivars using RAPD, AFLP and STMS markers. J Gen Breed 57:165–174
Tautz D (1989) Hypervariability of simple sequences as a general source of polymorphic DNA markers. Nucleic Acids Res 17:6463–6471
Udupa SM, Baum M (2001) High mutation rare and mutational bias at (TAA)n microsatellite loci in chickpea (Cicer arietinum L.). Mol Genet Genomics 266:343–344
Upadhyaya HD, Dwivedi SL, Baum M, Varshney RK, Udupa SM, Gowda Cholenahalli LL, Hoisington D, Singh S (2008) Genetic structure, diversity, and allelic richness in composite collection and reference set in chickpea (Cicer arietinum L.). BMC Plant Biol 8:106
Van Der Maesen LJG (1987) Origin, history and taxonomy of chickpea. In: Saxena MJ, Singh KB (eds) The chickpea. CAB International, Cambridge, pp 11–34
Vavilov NI (1926) Studies on the origin of cultivated plants. Inst Appl Bot Plant Breed, Leningrad
Weising K, Nybom H, Pfenninger M, Wolff K, Kahl G (2005) DNA fingerprinting in plants: principles, methods, and applications, 2nd edn. CRC Press, Boca Raton
Winter P, Benko-Iseppon AM, Hüttel B, Ratnaparkhe M, Tullu A, Sonnante G, Pfaff T, Tekeoglu M, Santra D, Sant VJ, Rajesh PN, Kahl GF, Muehlbauer J (2000) A linkage map of the chickpea (Cicer arietinum L.) genome based on recombinant inbred lines from a C. arietinum×C. reticulatum cross: localization of resistance genes for fusarium wilt races 4 and 5. Theor Appl Genet 101(7):1155–1163
Acknowledgements
This work was supported by the SaVeGraINPuglia Programme (Reg.CE n. 1698/2005 Programma di Sviluppo rurale per la Puglia 2007/2013. Misura 214—Azione 4 Sub azione a). The authors would like to acknowledge Dr. Venturino Bisignano for kindly providing some Apulian genotypes characterized in this study.
Author information
Authors and Affiliations
Corresponding authors
Additional information
C. De Giovanni and S. Pavan have contributed equally to this work.
Rights and permissions
About this article
Cite this article
De Giovanni, C., Pavan, S., Taranto, F. et al. Genetic variation of a global germplasm collection of chickpea (Cicer arietinum L.) including Italian accessions at risk of genetic erosion. Physiol Mol Biol Plants 23, 197–205 (2017). https://doi.org/10.1007/s12298-016-0397-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-016-0397-4