Abstract
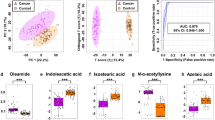
A metabolomic study for determination of certain urinary metabolomes, 1-methyladenosine (1-MA), 1-methylguanosine (1-MG), and 8-hydroxy-2′ deoxyguanosine (8-OHdG) in urine specimens of breast cancer patients. The accuracy of these metabolites and their combined score with cancer antigen 15-3 (CA15-3) was developed to improve the early detection of breast cancer. This study recruited 52 healthy individuals, 47 benign breast tumors, and 167 malignant breast tumor patients. Urine samples were handled to adjust the creatinine concentrations to 8 mg/dL (0.7 mmol/L) and analyzed using GC–MS to detect and quantify the selected urinary metabolomes in urine samples of all participants. The accuracy of individual urinary metabolomes and their combination with CA15-3 were evaluated using multivariate statistical analysis. The cutoff value of CA15-3 was 32.5 U/mL. Cutoff values of 1-MA, 1-MG, and 8-OHdG were 2.19, 2.1, and 7.3 µmol/mmol creatinine, respectively. The concentrations of 1-MA, 1-MG, and 8-OHdG were significantly higher in breast cancer patients, especially in the early-stage. The combination of three urinary metabolomes with CA15-3 improves the diagnostic sensitivity of breast cancer. For the combined score, the area under the curve (AUC) value of combined score ranged from 0.820 to 0.950, with high accuracy, ranged from 77.0 to 95.5%. The most significant AUC (0.973), sensitivity (90.1%), selectivity (94.0%) was recorded at comparing the healthy control with the early-stage of malignant breast cancer. In conclusion, the combination of three urinary metabolomes with serum CA15-3 improves the diagnostic sensitivity of breast cancer.



Similar content being viewed by others
References
DeSantis CE, Fedewa SA, Goding Sauer A, Kramer JL, Smith RA, Jemal A. Breast cancer statistics, 2015: convergence of incidence rates between black and white women. CA Cancer J Clin. 2016;66(1):31–42.
Struck-Lewicka W, Kordalewska M, Bujak R, Mpanga AY, Markuszewski M, Jacyna J, et al. Urine metabolic fingerprinting using LC–MS and GC–MS reveals metabolite changes in prostate cancer: a pilot study. J Pharm Biomed Anal. 2015;111:351–61.
Günther UL. Metabolomics biomarkers for breast cancer. Pathobiology. 2015;82(3–4):153–65.
Asiago VM, Alvarado LZ, Shanaiah N, Gowda GN, Owusu-Sarfo K, Ballas RA, et al. Early detection of recurrent breast cancer using metabolite profiling. Cancer Res. 2010;70(21):8309–18.
Nagrath D, Caneba C, Karedath T. Bellance N (2011) Metabolomics for mitochondrial and cancer studies. Biochim Biophys Acta (BBA) Bioenerg. 1807;6:650–63.
Armitage EG, Barbas C. Metabolomics in cancer biomarker discovery: current trends and future perspectives. J Pharm Biomed Anal. 2014;87:1–11.
Seidel A, Brunner S, Seidel P, Fritz G, Herbarth O. Modified nucleosides: an accurate tumour marker for clinical diagnosis of cancer, early detection and therapy control. Br J Cancer. 2006;94(11):1726.
Zhang S, Raftery D. Headspace SPME-GC–MS metabolomics analysis of urinary volatile organic compounds (VOCs). Mass Spectrom Metab Methods Protoc. 2014;1198(17):265–72.
Jobu K, Sun C, Yoshioka S, Yokota J, Onogawa M, Kawada C, et al. Metabolomics study on the biochemical profiles of odor elements in urine of human with bladder cancer. Biol Pharm Bull. 2012;35(4):639–42.
Safaei A, Oskouie AA, Mohebbi SR, Rezaei-Tavirani M, Mahboubi M, Peyvandi M, et al. Metabolomic analysis of human cirrhosis, hepatocellular carcinoma, non-alcoholic fatty liver disease and non-alcoholic steatohepatitis diseases. Gastroenterol Hepatol Bed Bench. 2016;9(3):158–65.
Omran MM, Farid K, Omar MA, Emran TM, El-Taweel FM, Tabll AA. A Combination of α-fetoprotein, midkine, thioredoxin and metabolite for predicting hepatocellular carcinoma. Ann Hepatol. 2019;19:179–85.
Struck-Lewicka W, Kaliszan R, Markuszewski MJ. Analysis of urinary nucleosides as potential cancer markers determined using LC–MS technique. J Pharm Biomed Anal. 2014;101:50–7.
Shulaev V. Metabolomics technology and bioinformatics. Brief Bioinform. 2006;7(2):128–39.
Weckwerth W. Metabolomics: an integral technique in systems biology. Bioanalysis. 2010;2(4):829–36.
Blau N, Duran M, Gibson KM. Laboratory guide to the methods in biochemical genetics. Berlin: Springer; 2008.
Omran MM, Rashed RE, Darwish H, Belal AA, Mohamed FZ. Development of a gas chromatography–mass spectrometry method for breast cancer diagnosis based on nucleoside metabolomes 1-methyl adenosine, 1-methylguanosine, and 8-hydroxy-2’-deoxyguanosine. Biomed Chromatogr. 2019;34:e4713.
Chen Y, Xu J, Zhang R, Abliz Z. Methods used to increase the comprehensive coverage of urinary and plasma metabolomes by MS. Bioanalysis. 2016;8(9):981–97.
Raftery DM, Pan Z, Gu H. Breast cancer biomarkers and identification methods using NMR and gas chromatography-mass spectrometry. Google Patents; 2015.
Oakman C, Tenori L, Biganzoli L, Santarpia L, Cappadona S, Luchinat C, et al. Uncovering the metabolomic fingerprint of breast cancer. Int J Biochem Cell Biol. 2011;43(7):1010–20.
Mishra P, Ambs S. Metabolic signatures of human breast cancer. Mol Cell Oncol. 2015;2(3):e992217.
Huang J-H, Fu L, Li B, Xie H-L, Zhang X, Chen Y, et al. Distinguishing the serum metabolite profiles differences in breast cancer by gas chromatography mass spectrometry and random forest method. RSC Adv. 2015;5(73):58952–8.
Beretov J, Wasinger V, Graham P, Millar E, Kearsley J, Li Y. Proteomics for breast cancer urine biomarkers. Adv Clin Chem. 2014;63:123–67.
Zhang A, Sun H, Wu X, Wang X. Urine metabolomics. Clin Chim Acta. 2012;414(5):65–9.
Zhang A, Sun H, Yan G, Wang P, Wang X. Mass spectrometry-based metabolomics: applications to biomarker and metabolic pathway research. Biomed Chromatogr. 2016;30(1):7–12.
Zhang A-H, Sun H, Qiu S, Wang X-J. Metabolomics in noninvasive breast cancer. Clin Chim Acta. 2013;424:3–7.
Chen J, Hou W, Han B, Liu G, Gong J, Li Y, et al. Target-based metabolomics for the quantitative measurement of 37 pathway metabolites in rat brain and serum using hydrophilic interaction ultra-high-performance liquid chromatography–tandem mass spectrometry. Anal Bioanal Chem. 2016;408(10):2527–42.
Pinu FR, Beale DJ, Paten AM, Kouremenos K, Swarup S, Schirra HJ, et al. Systems biology and multi-omics integration: viewpoints from the metabolomics research community. Metabolites. 2019;9(4):76–83.
Hedrich CM. Mechanistic aspects of epigenetic dysregulation in SLE. Clin Immunol. 2018;196:3–11.
Zhang C, Jia G. Reversible RNA Modification N1-methyladenosine (m1A) in mRNA and tRNA. Genomics Proteomics Bioinform. 2018;16(3):155–61.
Bayo J, Castaño M, Rivera F, Navarro F. Analysis of blood markers for early breast cancer diagnosis. Clin Transl Oncol. 2018;20(4):467–75.
Cala M, Aldana J, Sánchez J, Guio J, Meesters RJ. Urinary metabolite and lipid alterations in Colombian Hispanic women with breast cancer: A pilot study. J Pharm Biomed Anal. 2018;152:234–41.
Jasbi P, Wang D, Cheng SL, Fei Q, Cui JY, Liu L, et al. Breast cancer detection using targeted plasma metabolomics. J Chromatogr B. 2019;1105:26–37.
Dowling P, Henry M, Meleady P, Clarke C, Gately K, O’Byrne K, et al. Metabolomic and proteomic analysis of breast cancer patient samples suggests that glutamate and 12-HETE in combination with CA15-3 may be useful biomarkers reflecting tumour burden. Metabolomics. 2015;11(3):620–35.
Hsu W-Y, Chen C-J, Huang Y-C, Tsai F-J, Jeng L-B, Lai C-C. Urinary nucleosides as biomarkers of breast, colon, lung, and gastric cancer in Taiwanese. PLoS ONE. 2013;8(12):e81701.
Dutertre M, Lambert S, Carreira A, Amor-Guéret M, Vagner S. DNA damage: RNA-binding proteins protect from near and far. Trends Biochem Sci. 2014;39(3):141–9.
Beger RD. A review of applications of metabolomics in cancer. Metabolites. 2013;3(3):552–74.
Dudley E, Bond L. Mass spectrometry analysis of nucleosides and nucleotides. Mass Spectrom Rev. 2014;33(4):302–31.
Liebich H, Di Stefano C, Wixforth A, Schmid H. Quantitation of urinary nucleosides by high-performance liquid chromatography. J Chromatogr A. 1997;763(1–2):193–7.
Pande D, Negi R, Karki K, Khanna S, Khanna RS, Khanna H. Oxidative damage markers as possible discriminatory biomarkers in breast carcinoma. Transl Res. 2012;160(6):411–8.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zahran, F., Rashed, R., Omran, M. et al. Study on Urinary Candidate Metabolome for the Early Detection of Breast Cancer. Ind J Clin Biochem 36, 319–329 (2021). https://doi.org/10.1007/s12291-020-00905-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12291-020-00905-6