Abstract
Background
Tumor growth is mediated in part by glutamine, and glutaminase is an enzyme necessary for glutamine catabolism. We studied glutaminase (GLS1) gene expression in primary breast cancer to determine correlations with clinical and tumor characteristics, and gene associations in publicly available databases. A better understanding of glutaminase gene expression may help guide further exploration of glutaminase inhibitors in breast cancer.
Methods
GLS1 mRNA levels were evaluated in The Cancer Genome Atlas (n = 817) and METABRIC (n = 1992) datasets. Associations between GLS1 and tumor subtype (ANOVA followed by post-hoc Tukey test for pairwise comparisons) and selected genes involved in the pathogenesis of breast cancer (Pearson’s correlations) were determined in both datasets. In METABRIC, associations with overall survival (Cox proportional hazard model) were determined. For all analyses, p < 0.05 was the threshold for statistical significance.
Results
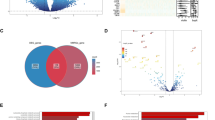
GLS1 expression was significantly higher in triple negative breast cancer (TNBC) than hormone receptor (HR) +/HER2− and HER2+ breast cancer (p < 0.001) and basal versus luminal A, luminal B, and HER2 enriched breast cancer (p < 0.001) in both datasets. In METABRIC, higher GLS1 expression was associated with improved overall survival (HR 0.91, 95% CI: 0.85–0.97, p = 0.005) and this association remained significant in the TNBC subset (HR 0.83, 95% CI: 0.71–0.98, p = 0.032). GLS1 had significant positive gene correlations with immune, proliferative, and basal genes, and inverse correlations with luminal genes and genes involved in metabolism.
Conclusion
GLS1 expression is highest in TNBC and basal breast cancer, supporting ongoing clinical investigation of GLS1 inhibition in TNBC. GLS1 may have prognostic implications but further research is needed to validate this finding. GLS1 had significant positive gene correlations with immune genes, which may have implications for potential combinations of glutaminase inhibition and immunotherapy.





Similar content being viewed by others
References
Jiang J, Srivastava S, Zhang J. Starve cancer cells of glutamine: break the spell or make a hungry monster? Cancers. 2019;11(6):804.
Masisi BK, El Ansari R, Alfarsi L, Rakha EA, Green AR, Craze ML. The role of glutaminase in cancer. Histopathology. 2020;76(4):498–508.
Mates JM, Campos-Sandoval JA, Santos-Jimenez JL, Marquez J. Dysregulation of glutaminase and glutamine synthetase in cancer. Cancer Lett. 2019;467:29–39.
Lampa M, Arlt H, He T, Ospina B, Reeves J, Zhang B, et al. Glutaminase is essential for the growth of triple-negative breast cancer cells with a deregulated glutamine metabolism pathway and its suppression synergizes with mTOR inhibition. PLoS ONE. 2017;12(9): e0185092.
Singleton DC, Dechaume AL, Murray PM, Katt WP, Baguley BC, Leung EY. Pyruvate anaplerosis is a mechanism of resistance to pharmacological glutaminase inhibition in triple-receptor negative breast cancer. BMC Cancer. 2020;20(1):470.
Gross MI, Demo SD, Dennison JB, Chen L, Chernov-Rogan T, Goyal B, et al. Antitumor activity of the glutaminase inhibitor CB-839 in triple-negative breast cancer. Mol Cancer Ther. 2014;13(4):890–901.
Kalinsky K, Harding JJ, DeMichele A, Infante JR, Gogineni K, Owaonikoko TK, et al. Abstract PD3–13: phase I study of CB-839, a first-in-class oral inhibitor of glutaminase, in combination with paclitaxel in patients with advanced triple negative breast cancer. Cancer Res. 2018;78(4_supplement):PD3-13.
Ciriello G, Gatza ML, Beck AH, Wilkerson MD, Rhie SK, Pastore A, et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell. 2015;163(2):506–19.
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012;486(7403):346–52.
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Investig. 2011;121(7):2750–67.
Burstein MD, Tsimelzon A, Poage GM, Covington KR, Contreras A, Fuqua SA, et al. Comprehensive genomic analysis identifies novel subtypes and targets of triple-negative breast cancer. Clin Cancer Res. 2015;21(7):1688–98.
Prat A, Perou CM. Deconstructing the molecular portraits of breast cancer. Mol Oncol. 2011;5(1):5–23.
Vidula N, Bardia A. Targeted therapy for metastatic triple negative breast cancer: the next frontier in precision oncology. Oncotarget. 2017;8(63):106167–8.
Edwards DN, Ngwa VM, Raybuck AL, Wang S, Hwang Y, Kim LC, et al. Selective glutamine metabolism inhibition in tumor cells improves antitumor T lymphocyte activity in triple-negative breast cancer. J Clin Invest. 2021;131(4):140100.
Fu A, Yu Z, Song Y, Zhang E. Silencing of glutaminase 1 resensitizes Taxol-resistant breast cancer cells to Taxol. Mol Med Rep. 2015;11(6):4727–33.
NCT03057600. Study of CB-839 in combination with paclitaxel in patients of African ancestry and Non-African ancestry with advanced TNBC. Available online: https://clinicaltrials.gov/ct2/show/NCT03057600. Accessed 15 Dec 2022.
Craze ML, Cheung H, Jewa N, Coimbra NDM, Soria D, El-Ansari R, et al. MYC regulation of glutamine-proline regulatory axis is key in luminal B breast cancer. Br J Cancer. 2018;118(2):258–65. https://doi.org/10.1038/bjc.2017.387.
Cacace A, Sboarina M, Vazeille T, Sonveaux P. Glutamine activates STAT3 to control cancer cell proliferation independently of glutamine metabolism. Oncogene. 2017;36(15):2074–84. https://doi.org/10.1038/onc.2016.364.
Mates JM, Campos-Sandoval JA, de Los Santos-Jimenez J, Marquez J. Glutaminases regulate glutathione and oxidative stress in cancer. Arch Toxicol. 2020.
Wang T, Cai B, Ding M, Su Z, Liu Y, Shen L. c-Myc overexpression promotes oral cancer cell proliferation and migration by enhancing glutaminase and glutamine synthetase activity. Am J Med Sci. 2019;358(3):235–42.
Acknowledgements
While this study was not funded, the I-SPY 1 clinical trial (CALGB 150007/150012; ACRIN 6657) was funded through the Alliance grant U10CA180821.
Author information
Authors and Affiliations
Contributions
All authors contributed to this study’s design, data analysis, and manuscript preparation. NV wrote the first draft of the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
NV: Research support (not related to this work) for clinical trials to the institution (MGH): Pfizer, Daehwa, Radius, Merck, Novartis. Advisory Board participation (not related to this work): OncoSec, AbbVie, Gilead, TerSera, Stemline. CY: Research funding (not related to this work) to the institution (UCSF): NCI and Quantum Leap Healthcare Collaborative. HSR: Institutional research support (not related to this work) from AMBRX; Astellas Pharma Inc.; AstraZeneca; Daiichi Sankyo, Inc.; F. Hoffmann-La Roche AG/Genentech, Inc.; Gilead Sciences, Inc.; GlaxoSmithKline; Lilly; Merck & Co., Inc.; Novartis Pharmaceuticals Corporation; OBI Pharma; Pfizer; Pionyr Immunotherapeutics; and Seattle Genetics, Inc; Sermonix Pharmaceuticals Inc.; Taiho Oncology, Inc. and Veru Inc. Travel support to academic meetings (not related to this work) from Merck, Astra Zeneca and Gilead. Consultancy/advisory support (not related to this work) from Puma, NAPO and Blueprint.
Ethical approval
This study was conducted in accordance with the ethical standards of the institutional research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. As this was an analysis of existing datasets, formal consent was not required. This article does not contain any studies with animals performed by any of the authors.
Research involving human participants
This research only involved de-identified patient data from existing datasets.
Informed consent
Institutional Review Board review and consent of patients was not required for this study as it involved analysis of publicly available de-identified datasets.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Vidula, N., Yau, C. & Rugo, H.S. Glutaminase (GLS1) gene expression in primary breast cancer. Breast Cancer 30, 1079–1084 (2023). https://doi.org/10.1007/s12282-023-01502-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12282-023-01502-0