Abstract
Background
Deciphering new molecules related to the breast cancer subtypes is crucial for prognosis and determining a better strategy for targeted therapy. In this study, we aimed to model ceRNAs networks in luminal A and luminal B subtypes of breast cancer and then delve deeper into the role of two candidate lncRNAs in breast tumors.
Methods
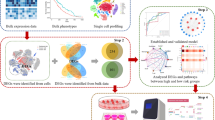
We constructed two networks as a regulatory model based on our previously identified transcription factors (TFs) and miRNAs with associated lncRNAs. Then, we highlighted the role of some lncRNAs in luminal subtypes of breast cancer using available online databases. Furthermore, we empirically quantified the expression levels of two candidate lncRNAs (DRAIC and TP53TG1) in breast tumors and normal tissues.
Results
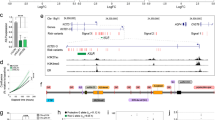
Here, we proposed a regulatory model for TFs–miRNAs–lncRNAs in luminal subtypes of breast cancer. We found 18 and 17 differentially expressed lncRNAs in luminal A and luminal B subtypes, respectively. Of these lncRNAs, 16 were associated with breast cancer patients' RFS and/or OS rates. Well-known lncRNAs like HOTAIR and MALAT1 were identified as central factors associated with patients’ survival rates in both networks. Based on the results acquired from our comprehensive in-silico data analysis, we carried out clinical experiments on two less-known lncRNAs, DRAIC and TP53TG1, and found a significant association between them with luminal subtypes of breast cancer. Interestingly, we discovered a significant association between DRAIC and TP53TG1 lncRNAs with ER- and PR-positive samples and lymph-node invasion in breast cancer patients.
Conclusion
According to the results, DRAIC and TP53TG1 lncRNAs are overexpressed in breast tumors and may play an oncogenic role with a moderate value of prognosis for luminal subtypes of breast cancer.









Similar content being viewed by others
Data availability
The data that support the findings of this study are available.
References
Pal B, et al. A single-cell RNA expression atlas of normal, preneoplastic and tumorigenic states in the human breast. EMBO J. 2021;40(11): e107333.
Yersal O, Barutca S. Biological subtypes of breast cancer: Prognostic and therapeutic implications. World J Clin Oncol. 2014;5(3):412.
Reis-Filho JS, et al. Molecular profiling: moving away from tumor philately. Sci Transl Med. 2010;2(47):43–7.
Creighton CJ. The molecular profile of luminal B breast cancer. Biologics. 2012;6:289.
Li A, et al. H19, a long non-coding RNA, mediates transcription factors and target genes through interference of microRNAs in pan-cancer. Mol Ther-Nucleic Acids. 2020;21:180–91.
Wang J, et al. Construction and comprehensive analysis of dysregulated long non-coding RNA-associated competing endogenous RNA network in clear cell renal cell carcinoma. J Cell Biochem. 2019;120(2):2576–93.
Bai H, et al. Comprehensive analysis of lncRNA–miRNA–mRNA during proliferative phase of rat liver regeneration. J Cell Physiol. 2019;234(10):18897–905.
Eskandari E, Motalebzadeh J. Transcriptomics-based screening of molecular signatures associated with patients overall survival and their key regulators in subtypes of breast cancer. Cancer Genet. 2019;239:62–74.
Zhao H, et al. LncTarD: a manually-curated database of experimentally-supported functional lncRNA–target regulations in human diseases. Nucleic Acids Res. 2020;48(D1):D118–26.
Cheng L, et al. LncRNA2Target v2. 0: a comprehensive database for target genes of lncRNAs in human and mouse. Nucleic Acids Res. 2019;47:D140–4.
Kang J, et al. RNAInter v4. 0: RNA interactome repository with redefined confidence scoring system and improved accessibility. Nucleic Acids Res. 2021;50:D326-332.
Gong J, et al. RISE: a database of RNA interactome from sequencing experiments. Nucleic Acids Res. 2018;46(D1):D194–201.
Tang Z, et al. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019;47(W1):W556–60.
Chandrashekar DS, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19(8):649–58.
Lánczky A, et al. miRpower: a web-tool to validate survival-associated miRNAs utilizing expression data from 2178 breast cancer patients. Breast Cancer Res Treat. 2016;160(3):439–46.
Perron U, Provero P, Molineris I. In silico prediction of lncRNA function using tissue specific and evolutionary conserved expression. BMC Bioinform. 2017;18(5):29–39.
Kun C, Lei Y. Microscopic traffic-emission simulation and case study for evaluation of traffic control strategies. J Transp Syst Eng Inf Technol. 2007;7(1):93–9.
Motalebzadeh J, Eskandari E. Transcription factors linked to the molecular signatures in the development of hepatocellular carcinoma on a cirrhotic background. Med Oncol. 2021;38(10):1–12.
Sivadas A, Kok VC, Ng K-L. Multi-omics analyses provide novel biological insights to distinguish lobular ductal types of invasive breast cancers. Breast Cancer Res Treat. 2022;193(2):361–79.
Sørensen KP, et al. Long non-coding RNA HOTAIR is an independent prognostic marker of metastasis in estrogen receptor-positive primary breast cancer. Breast Cancer Res Treat. 2013;142(3):529–36.
Li Z-X, et al. MALAT1: a potential biomarker in cancer. Cancer Manag Res. 2018;10:6757.
Chen Q, Zhu C, Jin Y. The oncogenic and tumor suppressive functions of the long noncoding RNA MALAT1: an emerging controversy. Front Genet. 2020;11:93.
Elias-Rizk T, et al. The long non coding RNA H19 as a biomarker for breast cancer diagnosis in Lebanese women. Sci Rep. 2020;10(1):1–7.
Li W, et al. The FOXN3-NEAT1-SIN3A repressor complex promotes progression of hormonally responsive breast cancer. J Clin Investig. 2017;127(9):3421–40.
Guo F, et al. miR-589-3p sponged by the lncRNA TINCR inhibits the proliferation, migration and invasion and promotes the apoptosis of breast cancer cells by suppressing the Akt pathway via IGF1R. Int J Mol Med. 2020;46(3):989–1002.
Zhang J, Guo S, Jia B. Down-regulation of long non-coding RNA MEG3 serves as an unfavorable risk factor for survival of patients with breast cancer. Eur Rev Med Pharmacol Sci. 2016;20(24):5143–7.
Cheng K, et al. lncRNA GAS5 inhibits colorectal cancer cell proliferation via the miR-182-5p/FOXO3a axis. Oncol Rep. 2018;40(4):2371–80.
Sakurai K, et al. The lncRNA DRAIC/PCAT29 locus constitutes a tumor-suppressive nexus. Mol Cancer Res. 2015;13(5):828–38.
Sun M, et al. Discovery, annotation, and functional analysis of long noncoding RNAs controlling cell-cycle gene expression and proliferation in breast cancer cells. Mol Cell. 2015;59(4):698–711.
Zhao D, Dong J-T. Upregulation of long non-coding RNA DRAIC correlates with adverse features of breast cancer. Non-coding RNA. 2018;4(4):39.
Kok VC, et al. Cross-platform in-silico analyses exploring tumor immune microenvironment with prognostic value in triple-negative breast cancer. Breast Cancer (Dove Medical Press). 2022;14:85–99.
Saha S, et al. Long noncoding RNA DRAIC inhibits prostate cancer progression by interacting with IKK to inhibit NF-κB activation. Can Res. 2020;80(5):950–63.
Zhang Z, et al. LncRNA DRAIC inhibits proliferation and metastasis of gastric cancer cells through interfering with NFRKB deubiquitination mediated by UCHL5. Cell Mol Biol Lett. 2020;25(1):1–17.
Diaz-Lagares A, et al. Epigenetic inactivation of the p53-induced long noncoding RNA TP53 target 1 in human cancer. Proc Natl Acad Sci. 2016;113(47):E7535–44.
Shao M, et al. Survival analysis for long noncoding RNAs identifies TP53TG1 as an antioncogenic target for the breast cancer. J Cell Physiol. 2020;235(10):6574–81.
Xiao H, et al. TP53TG1 enhances cisplatin sensitivity of non-small cell lung cancer cells through regulating miR-18a/PTEN axis. Cell Biosci. 2018;8(1):1–13.
Xue X, et al. LncRNA HOTAIR enhances ER signaling and confers tamoxifen resistance in breast cancer. Oncogene. 2016;35(21):2746–55.
Lu Q, et al. LncRNA TP53TG1 promotes the growth and migration of hepatocellular carcinoma cells via activation of ERK signaling. Non-coding RNA. 2021;7(3):52.
Wang H, et al. Long non-coding RNA TP53TG1 upregulates shcbp1 to promote retinoblastoma progression by sponging miR-33b. Cell Transplant. 2021;30:09636897211025223.
Zhang Y, et al. Long noncoding RNA TP53TG1 promotes pancreatic ductal adenocarcinoma development by acting as a molecular sponge of microRNA-96. Cancer Sci. 2019;110(9):2760–72.
Acknowledgements
This research was conducted using the authors’ personal money.
Author information
Authors and Affiliations
Contributions
JM: conceptualization, investigation, formal analysis, writing original draft, figures illustration, and writing—review and editing. EE: investigation, formal analysis, and writing—original draft.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical standards
This study was approved by the ethics committee of Iran National Tumor Bank (INTB). All procedures performed in the study involving human participants were in accordance with the ethical standards and with the Helsinki Declaration and its later amendments.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
About this article
Cite this article
Motalebzadeh, J., Eskandari, E. Comprehensive analysis of DRAIC and TP53TG1 in breast cancer luminal subtypes through the construction of lncRNAs regulatory model. Breast Cancer 29, 1050–1066 (2022). https://doi.org/10.1007/s12282-022-01385-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12282-022-01385-7