Abstract
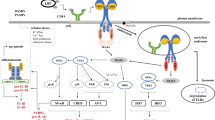
Oligodeoxynucleotides containing unmethylated CpG dinucleotides (CpG-ODN) can be specifically recognized by Toll-like receptor 9 (TLR9), provoking innate immune responses. Designed according to this structural feature, many synthetic phosphorothioate CpG-ODNs successfully activate macrophages. However, it is difficult to find potent stimulatory CpG-DNA fragments in microbial genomes. Therefore, whether microbial CpG-DNA substantially contributes to infectious and immune diseases remains controversial. In this study, high-throughput scanning was carried out for thousands of bacterial genomes with bioinformatics tools to comprehensively evaluate the distribution of CpG-DNA fragments. A random sampling test was then performed to verify their immunostimulatory properties by experiments in vitro and in vivo. Natural TLR9-dependent and potent stimulatory CpG-DNA fragments were found in microbial genomes. Interestingly, highly conserved stimulatory CpG-DNA fragments were found in 16S and 23S rDNA sequences with multiple copies, while others were species-specific. Additionally, we found that the reported active motifs were mostly non-stimulatory in natural CpG fragments. This evidence indicates that the previous structural descriptions of functional CpG-ODNs are incomplete. Our study has assessed the distribution of microbial CpG-DNA fragments, and identified natural stimulatory CpG-DNA fragments. These findings provide a deeper understanding of CpG-ODN structures and new evidence for microbial DNA inflammatory function and pathogenicity.
Similar content being viewed by others
Change history
01 April 2020
In the article by Liu et al. published in Journal of Microbiology 2020; 58, 153���162, 1# The Supplementary data���s consecutive numbers (Supplementary data Fig. S2) on 9th line of 4th paragraph in the section of ���Discussion.��� on page 160 should be corrected in (Supplementary data Fig. S1). The sentence should have read: The results showed that the nonspecific internalization of T03 had only a slight competition with CpG-ODNs (Supplementary data Fig. S1).
References
Anders, H.J. and Lech, M. 2013. NOD-like and Toll-like receptors or inflammasomes contribute to kidney disease in a canonical and a non-canonical manner. Kidney Int.84, 225–228.
Bauer, S., Kirschning, C.J., Häcker, H., Redecke, V., Hausmann, S., Akira, S., Wagner, H., and Lipford, G.B. 2001. Human TLR9 confers responsiveness to bacterial DNA via species-specific CpG motif recognition. Proc. Natl. Acad. Sci. USA98, 9237–9242.
Cai, Y.H., Lu, Z.Y., Shi, R.F., Xue, F., Chen, X.Y., Pan, M., Yuan, W.R., Xu, H., Li, W.P., and Zheng, J. 2009. Enhanced proliferation and activation of peripheral blood mononuclear cells in patients with psoriasis vulgaris mediated by streptococcal antigen with bacterial DNA. J. Invest. Dermatol.129, 2653–2660.
Cárdenas-Reyna, T., Angulo, C., Hori-Oshima, S., Velázquez-Lizárraga, E., and Reyes-Becerril, M. 2016. B-cell activating CpG ODN 1668 enhance the immune response of Pacific red snapper (Lutjanus peru) exposed to Vibrio parahaemolyticus. Dev. Comp. Immunol.62, 72–81.
Carpentier, A.F., Auf, G., and Delattre, J.Y. 2003. CpG-oligonucleotides for cancer immunotherapy: review of the literature and potential applications in malignant glioma. Front. Biosci.8, e115–127.
Chakravorty, S., Helb, D., Burday, M., Connell, N., and Alland, D. 2007. A detailed analysis of 16S ribosomal RNA gene segments for the diagnosis of pathogenic bacteria. J. Microbiol. Methods69, 330–339.
Clarridge, J.E.3rd. 2004. Impact of 16S rRNA gene sequence analysis for identification of bacteria on clinical microbiology and infectious diseases. Clin. Microbiol. Rev.17, 840–862.
Duriez, M., Quillay, H., Madec, Y., El Costa, H., Cannou, C., Marlin, R., de Truchis, C., Rahmati, M., Barré-Sinoussi, F., Nugeyre, M.T.et al. 2014. Human decidual macrophages and NK cells differentially express Toll-like receptors and display distinct cytokine profiles upon TLR stimulation. Front. Microbiol.5, 316.
Fearon, K., Marshall, J.D., Abbate, C., Subramanian, S., Yee, P., Gregorio, J., Teshima, G., Ott, G., Tuck, S., Van, N.G., et al. 2003. A minimal human immunostimulatory CpG motif that potently induces IFN-gamma and IFN-alpha production. Eur. J. Immunol.33, 2114–2122.
Hemmi, H., Takeuchi, O., Kawai, T., Kaisho, T., Sato, S., Sanjo, H., Matsumoto, M., Hoshino, K., Wagner, H., Takeda, K., et al. 2000. A Toll-like receptor recognizes bacterial DNA. Nature408, 740–745.
Huang, X., Palmer, S.R., Ahn, S.J., Richards, V.P., Williams, M.L., Nascimento, M.M., and Burne, R.A. 2016. A highly arginolytic Streptococcus species that potently antagonizes Streptococcus mutans. Appl. Environ. Microbiol.82, 2187–2201.
Iho, S., Yamamoto, T., Takahashi, T., and Yamamoto, S. 1999. Oligodeoxynucleotides containing palindrome sequences with internal 5′-CpG-3′ act directly on human NK and activated T cells to induce IFN-γ production in vitro. J. Immunol.163, 3642–3652.
Kembel, S.W., Wu, M., Eisen, J.A., and Green, J.L. 2012. Incorporating 16S gene copy number information improves estimates of microbial diversity and abundance. PLoS Comput. Biol.8, e1002743.
Krieg, A.M. 2002. CpG motifs in bacterial DNA and their immune effects. Annu. Rev. Immunol.20, 709–760.
Krieg, A.M., Yi, A.K., Matson, S., Waldschmidt, T.J., Bishop, G.A., Teasdale, R., Koretzky, G.A., and Klinman, D.M. 1995. CpG motifs in bacterial DNA trigger direct B-cell activation. Nature374, 546–549.
Li, Y., Cao, H., Wang, N., Xiang, Y., Lu, Y., Zhao, K., Zheng, J., and Zhou, H. 2011. A novel antagonist of TLR9 blocking all classes of immunostimulatory CpG-ODNs. Vaccine29, 2193–2198.
Liu, W., Yang, X., Wang, N., Fan, S., Zhu, Y., Zheng, X., and Li, Y. 2017. Multiple immunosuppressive effects of CpG-c41 on intracellular TLR-mediated inflammation. Mediators Inflamm.2017, 6541729.
Louca, S., Doebeli, M., and Parfrey, L.W. 2018. Correcting for 16S rRNA gene copy numbers in microbiome surveys remains an unsolved problem. Microbiome6, 41.
Mende, M., Hopert, A., Wünsche, W., Overhoff, M., Detzer, A., Börngen, K., Schlenke, P., Kirchner, H., and Sczakiel, G. 2007. A hexanucleotide selected for increased cellular uptake in cis contains a highly active CpG-motif in human B cells and primary peripheral blood mononuclear cells. Immunology120, 261–272.
Novák, K. 2014. Functional polymorphisms in Toll-like receptor genes for innate immunity in farm animals. Vet. Immunol. Immunopathol.157, 1–11.
Ohto, U., Ishida, H., Shibata, T., Sato, R., Miyake, K., and Shimizu, T. 2018. Toll-like receptor 9 contains two DNA binding sites that function cooperatively to promote receptor dimerization and activation. Immunity48, 649–658.e4.
Qian, C. and Cao, X. 2013. Regulation of Toll-like receptor signaling pathways in innate immune responses. Ann. N.Y. Acad. Sci.1283, 67–74.
Roberts, T.L., Sweet, M.J., Hume, D.A., and Stacey, K.J. 2005. Cutting edge:species-specific TLR9-mediated recognition of CpG and non-CpG phosphorothioate-modified oligonucleotides. J. Immunol.174, 605–608.
Rocha, D.J., Santos, C.S., and Pacheco, L.G. 2015. Bacterial reference genes for gene expression studies by RT-qPCR: survey and analysis. Antonie van Leeuwenhoek108, 685–693.
Sato, Y., Roman, M., Tighe, H., Lee, D., Corr, M., Nguyen, M.D., Silverman, G.J., Lotz, M., Carson, D.A., and Raz, E. 1996. Immunostimulatory DNA sequences necessary for effective intradermal gene immunization. Science273, 352–354.
Shimada, S., Yano, O., Inoue, H., Kuramoto, E., Fukuda, T., Yamamoto, H., Kataoka, T., and Tokunaga, T. 1985. Antitumor activity of the DNA fraction from Mycobacterium bovis BCG. II. Effects on various syngeneic mouse tumors. J. Natl. Cancer Inst.74, 681–688.
Sun, D.L., Jiang, X., Wu, Q.L., and Zhou, N.Y. 2013. Intragenomic heterogeneity of 16S rRNA genes causes overestimation of prokaryotic diversity. Appl. Environ. Microbiol.79, 5962–5969.
Talati, A.J., Kim, H.J., Kim, Y.I., Yi, A.K., and English, B.K. 2008. Role of bacterial DNA in macrophage activation by group B streptococci. Microbes Infect.10, 1106–1113.
Tokunaga, T., Yamamoto, T., and Yamamoto, S. 1999. How BCG led to the discovery of immunostimulatory DNA. Jpn. J. Infect. Dis.52, 1–11.
Verthelyi, D., Ishii, K.J., Gursel, M., Takeshita, F., and Klinman, D.M. 2001. Human peripheral blood cells differentially recognize and respond to two distinct CpG motifs. J. Immunol.166, 2372–2377.
Wang, J., Zhang, W., He, A., Zhao, W., and Cao, X. 2011. Screening of novel immunostimulatory CpG ODNs and their anti-leukemic effects as immunoadjuvants of tumor vaccines in murine acute lymphoblastic leukemia. Oncol. Rep.25, 519–529.
Yang, B., Wang, Y., and Qian, P.Y. 2016. Sensitivity and correlation of hypervariable regions in 16S rRNA genes in phylogenetic analysis. BMC Bioinformatics17, 135.
Acknowledgments
Thanks to Professor Jianping Xie for providing the experimental training for Juan Liu. This work was supported by the National Natural Science Foundation of China (Grant no. 81373133).
Author information
Authors and Affiliations
Corresponding author
Additional information
Conflicts of Interest
The authors declare that they have no competing interests.
Supplemental material for this article may be found at http://www.springerlink.com/content/120956.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Liu, J., Wei, Y., Lu, Y. et al. The discovery of potent immunostimulatory CpG-ODNs widely distributed in bacterial genomes. J Microbiol. 58, 153–162 (2020). https://doi.org/10.1007/s12275-020-9289-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-020-9289-y