Abstract
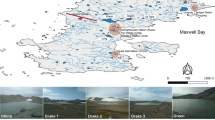
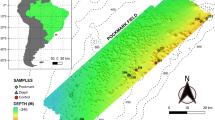
We have previously identified a sulfate methane transition zone (SMTZ) within the methane hydrate-bearing sediment in the Ulleung Basin, East Sea of Korea, and the presence of ANME-1b group in the sediment has been shown by phylogenetic analysis of a 16S rRNA gene. Herein, we describe taxonomic and functional profiling in the SMTZ sample by metagenomic analysis, comparing with that of surface sediment. Metagenomic sequences of 115 Mbp and 252 Mbp were obtained from SMTZ and surface sediments, respectively. The taxonomic profiling using BLASTX against the SEED within MG-RAST showed the prevalence of methanogens (19.1%), such as Methanosarcinales (12.0%) and Methanomicrobiales (4.1%) predominated within the SMTZ metagenome. A number of 185,200 SMTZ reads (38.9%) and 438,484 surface reads (62.5%) were assigned to functional categories, and methanogenesis-related reads were statistically significantly overrepresented in the SMTZ metagenome. However, the mapping analysis of metagenome reads to the reference genomes, most of the sequences of the SMTZ metagenome were mapped to ANME-1 draft genomes, rather than those of methanogens. Furthermore, the two copies of the methyl-coenzyme M reductase gene (mcrA) segments of the SMTZ metagenome were clustered with ANME-1b in the phylogenetic cluster. These results indicate that ANME-1b reads were miss-annotated to methanogens due to limitation of database. Many of key genes necessary for reverse methanogenesis were present in the SMTZ metagenome, except for N 5,N 10-methenyl-H4MPT reductase (mer) and CoB-CoM heterodisulfide reductase subunits D and E (hdrDE). These data suggest that the ANME-1b represents the primary player the anaerobic methane oxidation in the SMTZ, of the methane hydrate-bearing sediment at the Ulleung Basin, East Sea of Korea.
Similar content being viewed by others
References
Bahk, J.J., Kim, J.H., Kong, G.S., Park, Y., Lee, H., Park, Y., and Park, K.P. 2010. Occurrence of near-seafloor gas hydrates and associated cold vents in the Ulleung Basin, East Sea. Geosci. J. 13, 371–385.
Bäumer, S., Murakami, E., Brodersen, J., Gottschalk, G., Ragsdale, S.W., and Deppenmeier, U. 1998. The F420H2:heterodisulfide oxidoreductase system from Methanosarcina species. 2-Hydroxyphenazine mediates electron transfer from F420H2 dehydrogenase to heterodisulfide reductase. FEBS Lett. 428, 295–298.
Benjamini, Y. and Hochberg, Y. 1995. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy. Stat. Soc. B. Met. 57, 289–300.
Blumenberg, M., Seifert, R., Reitner, J., Pape, T., and Michaelis, W. 2004. Membrane lipid patterns typify distinct anaerobic methanotrophic consortia. Proc. Natl. Acad. Sci. USA 101, 11111–11116.
Boetius, A., Ravenschlag, K., Schubert, C.J., Rickert, D., Widdel, F., Gieseke, A., Amann, R., Jørgensen, B.B., Witte, U., and Pfannkuche, O. 2000. A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 407, 623–626.
Brochier-Armanet, C., Boussau, B., Gribaldo, S., and Forterre, P. 2008. Mesophilic Crenarchaeota: proposal for a third archaeal phylum, the Thaumarchaeota. Nat. Rev. Microbiol. 6, 245–252.
Cui, M., Ma, A., Qi, H., Zhuang, X., and Zhuang, G. 2015. Anaerobic oxidation of methane: an “active” microbial process. Microbiology Open 4, 1–11.
Ettwig, K.F., Butler, M.K., Le Paslier, D., Pelletier, E., Mangenot, S., Kuypers, M.M.M., Schreiber, F., Dutilh, B.E., Zedelius, J., de Beer, D., et al. 2010. Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 464, 543–548.
Fisher, W. 1958. On grouping for maximum homogeneity. J. Amer. Statist. Assoc. 53, 789–798.
Guy, L. and Ettema, T.J.G. 2011. The archaeal ‘TACK’ superphylum and the origin of eukaryotes. Trends Microbiol. 19, 580–587.
Hallam, S.J., Girguis, P.R., Preston, C.M., Richardson, P.M., and DeLong, E.F. 2003. Identification of methyl coenzyme M reductase A (mcrA) genes associated with methane-oxidizing archaea. Appl. Environ. Microbiol. 69, 5483–5491.
Hallam, S.J., Putnam, N., Preston, C.M., Detter, J.C., Richardson, P.M., Rokhsar, D., and Delong, E.F. 2004. Reverse methanogenesis: testing the hypothesis with environmental genomics. Science (New York, N.Y.) 305, 1457–1462.
Haroon, M.F., Hu, S., Shi, Y., Imelfort, M., Keller, J., Hugenholtz, P., Yuan, Z., and Tyson, G.W. 2013. Anaerobic oxidation of methane coupled to nitrate reduction in a novel archaeal lineage. Nature 500, 567–570.
Hedderich, R., Berkessel, A., and Thauer, R.K. 1990. Purification and properties of heterodisulfide reductase from Methanobacterium thermoautotrophicum (Strain Marburg). Eur. J. Biochem. 193, 255–261.
Heiden, S., Hedderich, R., Setzke, E., and Thauer, R.K. 1994. Purification of a two-subunit cytochrome-b-containing heterodisulfide reductase from methanol-grown Methanosarcina barkeri. Eur. J. Biochem. 221, 855–861.
Hinrichs, K.U., Hayes, J.M., Sylva, S.P., Brewer, P.G., and DeLong, E.F. 1999. Methane-consuming archaebacteria in marine sediments. Nature 398, 802–805.
Hoehler, T.M., Alperin, M.J., Albert, D.B., and Martens, C.S. 1994. Field and laboratory studies of methane oxidation in an anoxic marine sediment: Evidence for a methanogen-sulfate reducer consortium. Glob. Biogeochem. Cycles 8, 451–463.
Hong, W.L., Torres, M.E., Kim, J.H., Choi, J., and Bahk, J.J. 2013. Carbon cycling within the sulfate-methane-transition-zone in marine sediments from the Ulleung Basin. Biogeochemistry 115, 129–148.
Joye, S.B. 2012. Microbiology: a piece of the methane puzzle. Nature 491, 538–539.
Knittel, K. and Boetius, A. 2009. Anaerobic oxidation of methane: progress with an unknown process. Annu. Rev. Microbiol. 63, 311–334.
Knittel, K., Lösekann, T., Boetius, A., Kort, R., and Amann, R. 2005. Diversity and distribution of methanotrophic archaea at cold seeps. Appl. Environ. Microbiol. 71, 467–479.
Krüger, M., Meyerdierks, A., Glöckner, F.O., Amann, R., Widdel, F., Kube, M., Reinhardt, R., Kahnt, J., Böcher, R., Thauer, R.K., et al. 2003. A conspicuous nickel protein in microbial mats that oxidize methane anaerobically. Nature 426, 878–881.
Lee, S.H. and Chough, S.K. 2002. Distribution and origin of shallow gas in deep-sea sediments of the Ulleung Basin, East Sea (Sea of Japan). Geo-Marine Lett. 22, 204–209.
Lee, J.W., Kwon, K.K., Azizi, A., Oh, H.M., Kim, W., Bahk, J.J., Lee, D.H., and Lee, J.H. 2013. Microbial community structures of methane hydrate-bearing sediments in the Ulleung Basin, East Sea of Korea. Mar. Petrol. Geol. 47, 136–146.
Lim, D., Choi, J., Xu, Z., Kim, M., Choi, D., Jung, H., and Lee, P. 2009. Methane-derived authigenic carbonates from the Ulleung basin sediments, East Sea of Korea. Cont. Shelf. Res. 29, 1588–1596.
Meyer, F., Paarmann, D., D’Souza, M., Olson, R., Glass, E.M., Kubal, M., Paczian, T., Rodriguez, A., Stevens, R., Wilke, A., et al. 2008. The metagenomics RAST server - a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics 9, 1–8.
Meyerdierks, A., Kube, M., Kostadinov, I., Teeling, H., Glöckner, F.O., Reinhardt, R., and Amann, R. 2010. Metagenome and mRNA expression analyses of anaerobic methanotrophic archaea of the ANME-1 group. Environ. Microbiol. 12, 422–439.
Newcombe, R.G. 1998. Improved confidence intervals for the difference between binomial proportions based on paired data. Stat. Med. 17, 2635–2650.
Orphan, V.J., House, C.H., Hinrichs, K.U., McKeegan, K.D., and De-Long, E.F. 2001. Methane-consuming archaea revealed by directly coupled isotopic and phylogenetic analysis. Science 293, 484–487.
Parks, D.H. and Beiko, R.G. 2010. Identifying biologically relevant differences between metagenomic communities. Bioinformatics 26, 715–721.
Pereira, I.A.C., Ramos, A.R., Grein, F., Marques, M.C., da Silva, S.M., and Venceslau, S.S. 2011. A comparative genomic analysis of energy metabolism in sulfate reducing bacteria and archaea. Front. Microbiol. 2, 69.
Pires, R.H., Lourenç o, A.I., Morais, F., Teixeira, M., Xavier, A.V., Saraiva, L.M., and Pereira, I.A.C. 2003. A novel membrane-bound respiratory complex from Desulfovibrio desulfuricans ATCC 27774. Biochim. Biophys. Acta. 1605, 67–82.
Reeve, J.N., Nölling, J., Morgan, R.M., and Smith, D.R. 1997. Methanogenesis: genes, genomes, and who’s on first? J.Bacteriol. 179, 5975–5986.
Roalkvam, I., Jørgensen, S.L., Chen, Y., Stokke, R., Dahle, H., Hocking, W.P., Lanzén, A., Haflidason, H., and Steen, I.H. 2011. New insight into stratification of anaerobic methanotrophs in cold seep sediments. FEMS Microbiol. Ecol. 78, 233–243.
Ruff, S.E., Biddle, J.F., Teske, A.P., Knittel, K., Boetius, A., and Ramette, A. 2015. Global dispersion and local diversification of the methane seep microbiome. Proc. Natl. Acad. Sci. USA 112, 4015–4020.
Ryu, B.J., Collett, T.S., Riedel, M., Kim, G.Y., Chun, J.H., Bahk, J.J., Lee, J.Y., Kim, J.H., and Yoo, D.G. 2013. Scientific results of the Second Gas Hydrate Drilling Expedition in the Ulleung Basin (UBGH2). Mar. Pet. Geol. 47, 1–20.
Ryu, B.J., Riedel, M., Kim, J.H., Hyndman, R.D., Lee, Y.J., Chung, B.H., and Kim, I.S. 2009. Gas hydrates in the western deep-water Ulleung Basin, East Sea of Korea. Mar. Pet. Geol. 26, 1483–1498.
Stokke, R., Roalkvam, I., Lanzen, A., Haflidason, H., and Steen, I.H. 2012. Integrated metagenomic and metaproteomic analyses of an ANME-1-dominated community in marine cold seep sediments. Environ. Microbiol. 14, 1333–1346.
Thauer, R.K., Kaster, A.K., Goenrich, M., Schick, M., Hiromoto, T., and Shima, S. 2010. Hydrogenases from methanogenic archaea, nickel, a novel cofactor, and H2 storage. Annu. Rev. Biochem. 79, 507–536.
Valentine, D.L. 2002. Biogeochemistry and microbial ecology of methane oxidation in anoxic environments: a review. Antonie van Leeuwenhoek 81, 271–282.
Valentine, D.L. and Reeburgh, W.S. 2000. New perspectives on anaerobic methane oxidation. Environ. Microbiol. 2, 477–484.
Vigneron, A., Cruaud, P., Pignet, P., Caprais, J.C., Cambon-Bonavita, M.A., Godfroy, A., and Toffin, L. 2013. Archaeal and anaerobic methane oxidizer communities in the Sonora Margin cold seeps, Guaymas Basin (Gulf of California). ISME J. 7, 1595–1608.
Wang, F.P., Zhang, Y., Chen, Y., He, Y., Qi, J., Hinrichs, K.U., Zhang, X.X., Xiao, X., and Boon, N. 2014. Methanotrophic archaea possessing diverging methane-oxidizing and electron-transporting pathways. ISME J. 8, 1069–1078.
Welander, P.V. and Metcalf, W.W. 2008. Mutagenesis of the C1 oxidation pathway in Methanosarcina barkeri: new insights into the Mtr/Mer bypass pathway. J. Bacteriol. 190, 1928–1936.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplemental material for this article may be found at http://www.springerlink.com/content/120956.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lee, JW., Kwon, K.K., Bahk, JJ. et al. Metagenomic analysis reveals the contribution of anaerobic methanotroph-1b in the oxidation of methane at the Ulleung Basin, East Sea of Korea. J Microbiol. 54, 814–822 (2016). https://doi.org/10.1007/s12275-016-6379-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-016-6379-y