Abstract
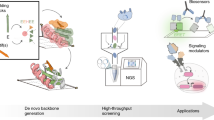
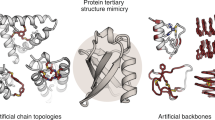
Proteins evolved to be very diverse and adapted to a multitude of central cellular functions. This impressive repertoire of today’s proteins developed through duplication and recombination of smaller protein and peptide building blocks. Based on such evolutionary observations we proposed a rational design strategy in which new functional hybrids can be build from fragments of existing proteins. With this approach we are tackling the long-standing goal of complex custom-made protein design.
Similar content being viewed by others
Literatur
Söding J, Lupas AN (2003) More than the sum of their parts: on the evolution of proteins from peptides. Bioessays 25:837–846
Söding J (2005) Protein homology detection by HMM-HMM comparison. Bioinformatics 21:951–960
Farias-Rico JA, Schmidt S, Höcker B (2014) Evolutionary relationship of two ancient protein superfolds. Nat Chem Biol 10:710–715
Höcker B, Schmidt S, Sterner R (2002) A common evolutionary origin of two elementary enzyme folds. FEBS Lett 510:133–135
Höcker B (2014) Design of proteins from smaller fragments–learning from evolution. Curr Opin Struct Biol 27:56–62
Bharat TA, Eisenbeis S, Zeth K et al. (2008) A beta alpha-barrel build by the combination of fragments from different folds. Proc Natl Acad Sci USA 105:9942–9947
Eisenbeis S, Proffitt W, Coles M et al. (2012) Potential of fragment recombination for rational design of proteins. J Am Chem Soc 134:4019–4022
Shanmugaratnam S, Eisenbeis S, Höcker B (2012) A highly stable protein chimera built from fragments of different folds. Protein Eng Des Sel 25:699–703
Alva V, Söding J, Lupas AN (2015) A vocabulary of ancient peptides at the origin of folded proteins. Elife 4:e09410
Farias-Rico JA, Höcker B (2013). Design of chimeric proteins by combination of subdomain-sized fragments. Methods Enzymol 523:389–405
Huang PS, Feldmeier K, Parmeggiani F et al. (2016) De novo design of a fourfold symmetric TIM-barrel protein with atomic level accuracy. Nat Chem Biol 12:29–34
Author information
Authors and Affiliations
Corresponding author
Additional information
Saacnicteh Toledo-Patino Biochemie- und Pharmaziestudium an der Michoacana de San Nicolas de Hidalgo Universität Morelia, Mexico, sowie Biochemie an der Universität Tübingen. 2013 Diplomarbeit am Max-Planck-Institut für Entwicklungsbiologie, Tübingen, dort seit 2014 Doktorandin in der AG „Protein-design“ und seit 2016 an der Universität Bayreuth.
Francisco Lobos 2011 Bachelor in Bioengineering an der University of Concepcion, Chile, dort 2014 Master in Biochemie und Bioinformatik. Seit 2015 Doktorand der International Max Planck Research School (IMPRS) Tübingen in der AG „Proteindesign“ am Max-Planck-Institut für Entwicklungsbiologie, Tübingen, und seit 2016 an der Universität Bayreuth.
Birte Höcker Biologiestudium an den Universitäten Göttingen und Ottawa, Kanada. 2003 Promotion im Fach Biochemie an der Universität Köln. 2003–2005 Postdoc am Duke Univer sity Medical Center, Durham, NC, USA. 2006–2016 Gruppenleiterin am Max-Planck-Institut für Entwicklungsbiologie, Tübingen. Seit 2016 W3-Professorin für Biochemie an der Universität Bayreuth. 2015 ERC Consolidator Award und Otto-Meyerhof-Preis der GBM.
Rights and permissions
About this article
Cite this article
Toledo-Patino, S., Lobos, F. & Höcker, B. Baukasten der Natur: neue Proteine aus konservierten Fragmenten. Biospektrum 23, 630–633 (2017). https://doi.org/10.1007/s12268-017-0847-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12268-017-0847-8