Abstract
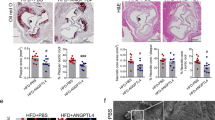
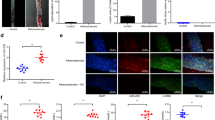
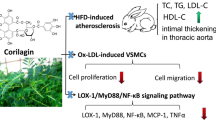
We aimed to explore the effect of chaetocin on atherosclerosis and its possible mechanism. In vitro, we observed that chaetocin treatment significantly inhibited the proliferation of VSMCs in concentration- and time-dependent manner. We also found that chaetocin suppressed the migration of VSMCs. Moreover, chaetocin treatment induced a contractile phenotype in VSMCs by increasing α-SMA and SM22α expression. In addition, chaetocin treatment attenuated the accumulation of H3K9me3 on VSMCs contractile gene promoters, which promoted the expression of α-SMA and SM22α. In vivo, chaetocin treatment decreased the H3K9me3 expression, diminished atherosclerotic plaque formation, and increased plaque stability by decreasing necrotic core area and lipid accumulation and increasing collagen content and contractile VSMC phenotype. We demonstrated a new function of chaetocin in inhibiting atherosclerosis progression and increasing plaque stability partly by inhibiting pathological phenotypic switching of VSMCs. These newly identified roles of chaetocin might provide a novel therapeutic target in atherosclerosis.





Similar content being viewed by others
Abbreviations
- VSMCs:
-
Vascular smooth muscle cells
- ApoE:
-
Apolipoprotein E
- ECM:
-
Extracellular matrix
- α-SMA:
-
α-Smooth muscle actin
- SFM:
-
Serum-free media
References s
Shankman, L. S., Gomez, D., Cherepanova, O. A., Salmon, M., Alencar, G. F., Haskins, R. M., et al. (2015). KLF4-dependent phenotypic modulation of smooth muscle cells has a key role in atherosclerotic plaque pathogenesis. Nature Medicine, 21(6), 628–637.
Shah, A., Gray, K., Figg, N., Finigan, A., Starks, L., & Bennett, M. (2018). Defective base excision repair of oxidative DNA damage in vascular smooth muscle cells promotes atherosclerosis. Circulation, 138(14), 1446–1462.
Ding, Y., Zhang, M., Zhang, W., Lu, Q., Cai, Z., Song, P., et al. (2016). AMP-activated protein kinase alpha 2 deletion induces VSMC phenotypic switching and reduces features of atherosclerotic plaque stability. Circulation Research, 119(6), 718–730.
Alexander, M. R., & Owens, G. K. (2012). Epigenetic control of smooth muscle cell differentiation and phenotypic switching in vascular development and disease. Annual Review of Physiology, 74, 13–40.
Liu, R., Jin, Y., Tang, W. H., Qin, L., Zhang, X., Tellides, G., et al. (2013). Ten-eleven translocation-2 (TET2) is a master regulator of smooth muscle cell plasticity. Circulation, 128(18), 2047–2057.
Guo, X., Shi, N., Cui, X. B., Wang, J. N., Fukui, Y., & Chen, S. Y. (2015). Dedicator of cytokinesis 2, a novel regulator for smooth muscle phenotypic modulation and vascular remodeling. Circulation Research, 116(10), e71–e80.
Hutter, R., Huang, L., Speidl, W. S., Giannarelli, C., Trubin, P., Bauriedel, G., et al. (2013). Novel small leucine-rich repeat protein podocan is a negative regulator of migration and proliferation of smooth muscle cells, modulates neointima formation, and is expressed in human atheroma. Circulation, 128(22), 2351–2363.
Yu, Q., Li, W., Jin, R., Yu, S., Xie, D., Zheng, X., et al. (2019). PI3Kgamma (phosphoinositide 3-kinase gamma) regulates vascular smooth muscle cell phenotypic modulation and neointimal formation through CREB (cyclic AMP-response element binding protein)/YAP (yes-associated protein) signaling. Arteriosclerosis, Thrombosis, and Vascular Biology, 39(3), e91–e105.
Chappell, J., Harman, J. L., Narasimhan, V. M., Yu, H., Foote, K., Simons, B. D., et al. (2016). Extensive proliferation of a subset of differentiated, yet plastic, medial vascular smooth muscle cells contributes to neointimal formation in mouse injury and atherosclerosis models. Circulation Research, 119(12), 1313–1323.
Yu, H., Clarke, M. C., Figg, N., Littlewood, T. D., & Bennett, M. R. (2011). Smooth muscle cell apoptosis promotes vessel remodeling and repair via activation of cell migration, proliferation, and collagen synthesis. Arteriosclerosis, Thrombosis, and Vascular Biology, 31(11), 2402–2409.
Kim, J., & Movassaghi, M. (2015). Biogenetically-inspired total synthesis of epidithiodiketopiperazines and related alkaloids. Accounts of Chemical Research, 48(4), 1159–1171.
Nakajo, H., Ishibashi, K., Aoyama, K., Kubota, S., Hasegawa, H., Yamaguchi, N., et al. (2019). Role for tyrosine phosphorylation of SUV39H1 histone methyltransferase in enhanced trimethylation of histone H3K9 via neuregulin-1/ErbB4 nuclear signaling. Biochemical and Biophysical Research Communications, 511(4), 765–771.
Mozzetta, C., Boyarchuk, E., Pontis, J., & Ait-Si-Ali, S. (2015). Sound of silence: The properties and functions of repressive Lys methyltransferases. Nature Reviews Molecular Cell Biology, 16(8), 499–513.
Greissel, A., Culmes, M., Burgkart, R., Zimmermann, A., Eckstein, H. H., Zernecke, A., et al. (2016). Histone acetylation and methylation significantly change with severity of atherosclerosis in human carotid plaques. Cardiovascular Pathology, 25(2), 79–86.
Chen, J., Zhang, J., Yang, J., Xu, L., Hu, Q., Xu, C., et al. (2017). Histone demethylase KDM3a, a novel regulator of vascular smooth muscle cells, controls vascular neointimal hyperplasia in diabetic rats. Atherosclerosis, 257, 152–163.
Li, M. F., Zhang, R., Li, T. T., Chen, M. Y., Li, L. X., Lu, J. X., et al. (2016). High glucose increases the expression of inflammatory cytokine genes in macrophages through H3K9 methyltransferase mechanism. Journal of Interferon and Cytokine Research, 36(1), 48–61.
Chen, M. Y., Ke, J. F., Zhang, Z. H., Li, M. F., Wang, J. W., Lu, J. X., et al. (2021). Deletion of Fam172a accelerates advanced atherosclerosis and induces plaque instability. Atherosclerosis, 333, 39–47.
Sekita, S., Yoshihira, K., Natori, S., Udagawa, S., Muroi, T., Sugiyama, Y., et al. (1981). Mycotoxin production by Chaetomium spp. and related fungi. Canadian Journal of Microbiology, 27(8), 766–772.
Song, X., Zhao, Z., Qi, X., Tang, S., Wang, Q., Zhu, T., et al. (2015). Identification of epipolythiodioxopiperazines HDN-1 and chaetocin as novel inhibitor of heat shock protein 90. Oncotarget, 6(7), 5263–5274.
Tibodeau, J. D., Benson, L. M., Isham, C. R., Owen, W. G., & Bible, K. C. (2009). The anticancer agent chaetocin is a competitive substrate and inhibitor of thioredoxin reductase. Antioxidants & Redox Signaling, 11(5), 1097–1106.
Lai, Y. S., Chen, J. Y., Tsai, H. J., Chen, T. Y., Hung, W. C. (2015). The SUV39H1 inhibitor chaetocin induces differentiation and shows synergistic cytotoxicity with other epigenetic drugs in acute myeloid leukemia cells. Blood Cancer Journal, 5, e313.
Lee, Y. M., Lim, J. H., Yoon, H., Chun, Y. S., & Park, J. W. (2011). Antihepatoma activity of chaetocin due to deregulated splicing of hypoxia-inducible factor 1 alpha pre-mRNA in mice and in vitro. Hepatology, 53(1), 171–180.
Bennett, M. R., Sinha, S., & Owens, G. K. (2016). Vascular smooth muscle cells in atherosclerosis. Circulation Research, 118(4), 692–702.
Yang, L., Gao, L., Nickel, T., Yang, J., Zhou, J., Gilbertsen, A., et al. (2017). Lactate promotes synthetic phenotype in vascular smooth muscle cells. Circulation Research, 121(11), 1251–1262.
Lu, Q. B., Wan, M. Y., Wang, P. Y., Zhang, C. X., Xu, D. Y., Liao, X., et al. (2018). Chicoric acid prevents PDGF-BB-induced VSMC dedifferentiation, proliferation and migration by suppressing ROS/NFkappaB/mTOR/P70S6K signaling cascade. Redox Biology, 14, 656–668.
Wu, B., Zhang, L., Zhu, Y. H., Zhang, Y. E., Zheng, F., Yang, J. Y., et al. (2018). Mesoderm/mesenchyme homeobox gene l promotes vascular smooth muscle cell phenotypic modulation and vascular remodeling. International Journal of Cardiology, 251, 82–89.
Gao, S., Xu, L., Zhang, Y., Yu, Q., Li, J., Guan, H., et al. (2018). Salusin-alpha Inhibits Proliferation and Migration of Vascular Smooth Muscle Cell via Akt/mTOR Signaling. Cellular Physiology and Biochemistry, 50(5), 1740–1753.
Li, P., Zhu, N., Yi, B., Wang, N., Chen, M., You, X., et al. (2013). MicroRNA-663 regulates human vascular smooth muscle cell phenotypic switch and vascular neointimal formation. Circulation Research, 113(10), 1117–1127.
Lin, H., Ni, T., Zhang, J., Meng, L., Gao, F., Pan, S., et al. (2018). Knockdown of Herp alleviates hyperhomocysteinemia mediated atherosclerosis through the inhibition of vascular smooth muscle cell phenotype switching. International Journal of Cardiology, 269, 242–249.
Chiba, T., Saito, T., Yuki, K., Zen, Y., Koide, S., Kanogawa, N., et al. (2015). Histone lysine methyltransferase SUV39H1 is a potent target for epigenetic therapy of hepatocellular carcinoma. International Journal of Cancer, 136(2), 289–298.
Yokoyama, Y., Hieda, M., Nishioka, Y., Matsumoto, A., Higashi, S., Kimura, H., et al. (2013). Cancer-associated upregulation of histone H3 lysine 9 trimethylation promotes cell motility in vitro and drives tumor formation in vivo. Cancer Science, 104(7), 889–895.
Sepsa, A., Levidou, G., Gargalionis, A., Adamopoulos, C., Spyropoulou, A., Dalagiorgou, G., et al. (2015). Emerging role of linker histone variant H1x as a biomarker with prognostic value in astrocytic gliomas. A multivariate analysis including trimethylation of H3K9 and H4K20. PLOS ONE, 10(1), e115101.
McDonald, O. G., & Owens, G. K. (2007). Programming smooth muscle plasticity with chromatin dynamics. Circulation Research, 100(10), 1428–1441.
Liu, R., Leslie, K. L., & Martin, K. A. (2015). Epigenetic regulation of smooth muscle cell plasticity. Biochimica et Biophysica Acta, 1849(4), 448–453.
Manabe, I., & Owens, G. K. (2001). Recruitment of serum response factor and hyperacetylation of histones at smooth muscle-specific regulatory regions during differentiation of a novel P19-derived in vitro smooth muscle differentiation system. Circulation Research, 88(11), 1127–1134.
McDonald, O. G., Wamhoff, B. R., Hoofnagle, M. H., & Owens, G. K. (2006). Control of SRF binding to CArG box chromatin regulates smooth muscle gene expression in vivo. The Journal of Clinical Investigation, 116(1), 36–48.
Villeneuve, L. M., Reddy, M. A., Lanting, L. L., Wang, M., Meng, L., & Natarajan, R. (2008). Epigenetic histone H3 lysine 9 methylation in metabolic memory and inflammatory phenotype of vascular smooth muscle cells in diabetes. Proceedings of the National Academy of Sciences of the United States America, 105(26), 9047–9052.
Weng, X., Cheng, X., Wu, X., Xu, H., Fang, M., & Xu, Y. (2014). Sin3B mediates collagen type I gene repression by interferon gamma in vascular smooth muscle cells. Biochemical and Biophysical Research Communications, 447(2), 263–270.
Schweizer, S., Harms, C., Lerch, H., Flynn, J., Hecht, J., Yildirim, F., et al. (2015). Inhibition of histone methyltransferases SUV39H1 and G9a leads to neuroprotection in an in vitro model of cerebral ischemia. Journal of Cerebral Blood Flow and Metabolism, 35(10), 1640–1647.
Yang, G., Weng, X., Zhao, Y., Zhang, X., Hu, Y., Dai, X., et al. (2017). The histone H3K9 methyltransferase SUV39H links SIRT1 repression to myocardial infarction. Nature Communications, 8, 14941.
Ono, T., Kamimura, N., Matsuhashi, T., Nagai, T., Nishiyama, T., Endo, J., et al. (2017). The histone 3 lysine 9 methyltransferase inhibitor chaetocin improves prognosis in a rat model of high salt diet-induced heart failure. Science and Reports, 7, 39752.
Acknowledgements
We thank the other investigators, the staff, and the participants of the present study for their valuable contributions.
Funding
This work was supported by grants from the National Key Research and Development Plan (2018YFC1314900 and 2018YFC1314905), the National Natural Science Foundation of China (81170759, 81770813, 82070866, and 82100876), the Science and Technology Commission of Shanghai Municipality (15411960600), and Joint Funds for the Innovation of Science and Technology, Fujian Province (No. 2020Y9106).
Shanghai Municipal Key Clinical Specialty which granted to Dr. Lian-Xi Li.
Author information
Authors and Affiliations
Contributions
Li LX designed the study, supervised the work, and reviewed and edited the manuscript. Chen MY, Zhang ZH, and Ke JF performed the studies. Chen MY wrote the manuscript. Li TT, Li MF, and Lu JX researched the data and reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval
All institutional and national guidelines for the care and use of laboratory animals were followed and approved by the Institutional Animal Care and Use Committees at Shanghai Jiao Tong University Affiliated Sixth People’s Hospital. This article does not contain any studies with humans.
Conflict of Interest
The authors declare no competing interests.
Additional information
Associate Editor Joost Sluijter oversaw the review of this article.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12265_2022_10258_MOESM1_ESM.jpg
Supplementary file1 (JPG 217 kb) The comparison of lipid profile between two groups a Quantification of serum total cholesterol levels. b Quantification of serum triglyceride levels. c Quantification of serum free fatty acid levels. d Quantification of serum low density lipoprotein-cholesterol levels. f Quantification of serum high density lipoprotein-cholesterol levels. n = 9 for each group. Data was expressed as mean ± SEM. Statistical comparisons were made using t-test. *p < 0.05 vs control group.
12265_2022_10258_MOESM2_ESM.jpg
Supplementary file2 (JPG 1410 kb) Chaetocin reduces VSMCs apoptosis in plaque a Representative image of apoptosis was assessed by staining for TUNEL (green) in cross-sections from the aortic root. b Quantification assessment of apoptosis rate. n=6 for control group and n=5 for chaetocin treatment group. Data was expressed as mean ± SEM. Statistical comparisons were made using t-test. **p < 0.01 vs control group.
Rights and permissions
About this article
Cite this article
Chen, MY., Zhang, ZH., Ke, JF. et al. Chaetocin attenuates atherosclerosis progression and inhibits vascular smooth muscle cell phenotype switching. J. of Cardiovasc. Trans. Res. 15, 1270–1282 (2022). https://doi.org/10.1007/s12265-022-10258-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12265-022-10258-5