Abstract
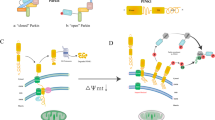
Lipid droplet (LD) is a kind of subcellular organelle, which originates from the endoplasmic reticulum (ER). LDs can move flexibly between other organelles and store energy in the cells. In recent years, LDs and lipid droplet–associated proteins have attracted added attention at home and abroad, especially in cardiovascular diseases. Cardiovascular diseases, especially ischemic heart disease (IHD), have always been the focus of attention because of their high morbidity and mortality. Atherosclerosis and myocardial remodeling are two important pathologic processes of IHD, and LDs and other organelles are involved in the development of the disease. The interaction between LDs and ER is involved in the formation of foam cells in atherosclerosis. And LDs, mitochondria, and lysosomes also affect the remodeling of cardiomyocytes by affecting ROS production and regulating PI3K/AKT pathways. In this article, we will review the role of LDs in IHD.
Graphical abstract


Similar content being viewed by others
Availability of Data and Material
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
Code Availability
No
References
van Meer, G. (2005). Cellular lipidomics. EMBO Journal, 24(18), 3159–3165. https://doi.org/10.1038/sj.emboj.7600798
Plakkal Ayyappan, J., Paul, A., & Goo, Y. H. (2016). Lipid droplet-associated proteins in atherosclerosis (Review). Molecular Medicine Reports, 13(6), 4527–4534. https://doi.org/10.3892/mmr.2016.5099
Saliba, A. E., Vonkova, I., & Gavin, A. C. (2015). The systematic analysis of protein-lipid interactions comes of age. Nature Reviews Molecular Cell Biology, 16(12), 753–761. https://doi.org/10.1038/nrm4080
Bosch M, Sanchez-Alvarez M, Fajardo A, et al. Mammalian lipid droplets are innate immune hubs integrating cell metabolism and host defense. Science 2020;370(6514)https://doi.org/10.1126/science.aay8085
Stanley, W. C., Recchia, F. A., & Lopaschuk, G. D. (2005). Myocardial substrate metabolism in the normal and failing heart. Physiological Reviews, 85(3), 1093–1129. https://doi.org/10.1152/physrev.00006.2004
Lopaschuk, G. D., Belke, D. D., Gamble, J., Itoi, T., & Schonekess, B. O. (1994). Regulation of fatty acid oxidation in the mammalian heart in health and disease. Biochimica et Biophysica Acta, 1213(3), 263–276. https://doi.org/10.1016/0005-2760(94)00082-4
Mortality, G. B. D., Causes of Death C. (2016). Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980–2015: A systematic analysis for the Global Burden of Disease Study 2015. Lancet, 388(10053), 1459–1544. https://doi.org/10.1016/S0140-6736(16)31012-1
Gidda, S. K., Park, S., Pyc, M., et al. (2016). Lipid droplet-associated proteins (LDAPs) are required for the dynamic regulation of neutral lipid compartmentation in plant cells. Plant Physiology, 170(4), 2052–2071. https://doi.org/10.1104/pp.15.01977
Xu, W., Wu, L., Yu, M., et al. (2016). Differential roles of cell death-inducing DNA fragmentation factor-alpha-like effector (CIDE) proteins in promoting lipid droplet fusion and growth in subpopulations of hepatocytes. Journal of Biological Chemistry, 291(9), 4282–4293. https://doi.org/10.1074/jbc.M115.701094
Bickel, P. E., Tansey, J. T., & Welte, M. A. (2009). PAT proteins, an ancient family of lipid droplet proteins that regulate cellular lipid stores. Biochimica Et Biophysica Acta-Molecular and Cell Biology of Lipids, 1791(6), 419–440. https://doi.org/10.1016/j.bbalip.2009.04.002
Varghese M, Kimler VA, Ghazi FR, et al. Adipocyte lipolysis affects Perilipin 5 and cristae organization at the cardiac lipid droplet-mitochondrial interface. Scientific Reports. Mar 2019;9ARTN 4734. https://doi.org/10.1038/s41598-019-41329-4
Karanasios, E., Han, G. S., Xu, Z., Carman, G. M., & Siniossoglou, S. (2010). A phosphorylation-regulated amphipathic helix controls the membrane translocation and function of the yeast phosphatidate phosphatase. Proceedings of the National Academy of Sciences of the United States of America, 107(41), 17539–17544. https://doi.org/10.1073/pnas.1007974107
Adeyo, O., Horn, P. J., Lee, S. K., et al. (2011). The yeast lipin orthologue Pah1p is important for biogenesis of lipid droplets. Journal of Cell Biology, 192(6), 1043–1055. https://doi.org/10.1083/jcb.201010111
Szymanski, K. M., Binns, D., Bartz, R., et al. (2007). The lipodystrophy protein seipin is found at endoplasmic reticulum lipid droplet junctions and is important for droplet morphology. Proceedings of the National Academy of Sciences of the United States of America, 104(52), 20890–20895. https://doi.org/10.1073/pnas.0704154104
Gross, D. A., Zhan, C. Y., & Silver, D. L. (2011). Direct binding of triglyceride to fat storage-inducing transmembrane proteins 1 and 2 is important for lipid droplet formation. Proceedings of the National Academy of Sciences of the United States of America, 108(49), 19581–19586. https://doi.org/10.1073/pnas.1110817108
Henne, W. M., Reese, M. L., & Goodman, J. M. (2019). The assembly of lipid droplets and their roles in challenged cells (vol 37, e98947, 2018). Embo Journal, 38(9), ARTN e101816. https://doi.org/10.15252/embj.2019101816
Wang HJ, Becuwe M, Housden BE, et al. Seipin is required for converting nascent to mature lipid droplets. Elife 2016;5ARTN e16582 https://doi.org/10.7554/eLife.16582
Jackson, C. L. (2019). Lipid droplet biogenesis. Current Opinion in Cell Biology, 59, 88–96. https://doi.org/10.1016/j.ceb.2019.03.018
Talukder, M. M. U., Sim, M. F. M., O’Rahilly, S., Edwardson, J. M., & Rochford, J. J. (2015). Seipin oligomers can interact directly with AGPAT2 and lipin 1, physically scaffolding critical regulators of adipogenesis. Molecular Metabolism, 4(3), 199–209. https://doi.org/10.1016/j.molmet.2014.12.013
Hayes, M., Choudhary, V., Ojha, N., et al. (2017). Fat storage-inducing transmembrane (FIT or FITM) proteins are related to lipid phosphatase/phosphotransferase enzymes. Microbial Cell, 5(2), 88–103. https://doi.org/10.15698/mic2018.02.614
Choudhary, V., Ojha, N., Golden, A., & Prinz, W. A. (2015). A conserved family of proteins facilitates nascent lipid droplet budding from the ER. Journal of Cell Biology, 211(2), 261–271. https://doi.org/10.1083/jcb.201505067
Joshi, A. S., Nebenfuehr, B., Choudhary, V., et al. (2018). Lipid droplet and peroxisome biogenesis occur at the same ER subdomains. Nature communications, 9(1), 2940. https://doi.org/10.1038/s41467-018-05277-3
Wang, S., Idrissi, F. Z., Hermansson, M., Grippa, A., Ejsing, C. S., & Carvalho, P. (2018). Seipin and the membrane-shaping protein Pex30 cooperate in organelle budding from the endoplasmic reticulum. Nature communications, 9(1), 2939. https://doi.org/10.1038/s41467-018-05278-2
Gao, G., Chen, F. J., Zhou, L., et al. (2017). Control of lipid droplet fusion and growth by CIDE family proteins. Biochimica et Biophysica Acta - Molecular and Cell Biology of Lipids, 1862(10 Pt B), 1197–1204. https://doi.org/10.1016/j.bbalip.2017.06.009
Farese, R. V., Jr., & Walther, T. C. (2009). Lipid droplets finally get a little R-E-S-P-E-C-T. Cell, 139(5), 855–860. https://doi.org/10.1016/j.cell.2009.11.005
Dupont, N., Chauhan, S., Arko-Mensah, J., et al. (2014). Neutral lipid stores and lipase PNPLA5 contribute to autophagosome biogenesis. Current Biology, 24(6), 609–620. https://doi.org/10.1016/j.cub.2014.02.008
Valm, A. M., Cohen, S., Legant, W. R., et al. (2017). Applying systems-level spectral imaging and analysis to reveal the organelle interactome. Nature, 546(7656), 162–167. https://doi.org/10.1038/nature22369
Olzmann, J. A., & Carvalho, P. (2019). Dynamics and functions of lipid droplets. Nature Reviews Molecular Cell Biology, 20(3), 137–155. https://doi.org/10.1038/s41580-018-0085-z
Aviram, R., Manella, G., Kopelman, N., et al. (2016). Lipidomics analyses reveal temporal and spatial lipid organization and uncover daily oscillations in intracellular organelles. Molecular Cell, 62(4), 636–648. https://doi.org/10.1016/j.molcel.2016.04.002
Agmon, E., & Stockwell, B. R. (2017). Lipid homeostasis and regulated cell death. Current Opinion in Chemical Biology, 39, 83–89. https://doi.org/10.1016/j.cbpa.2017.06.002
Williams, G. S. B., Boyman, L., & Lederer, W. J. (2015). Mitochondrial calcium and the regulation of metabolism in the heart. Journal of Molecular and Cellular Cardiology, 78, 35–45. https://doi.org/10.1016/j.yjmcc.2014.10.019
Neubauer, S. (2007). The failing heart–an engine out of fuel. New England Journal of Medicine, 356(11), 1140–1151. https://doi.org/10.1056/NEJMra063052
Long, Q., Yang, K., & Yang, Q. (2015). Regulation of mitochondrial ATP synthase in cardiac pathophysiology. American Journal of Cardiovascular Diseases, 5(1), 19–32.
Gemmink, A., Daemen, S., Kuijpers, H. J. H., et al. (2018). Super-resolution microscopy localizes perilipin 5 at lipid droplet-mitochondria interaction sites and at lipid droplets juxtaposing to perilipin 2. Biochimica Et Biophysica Acta-Molecular and Cell Biology of Lipids, 1863(11), 1423–1432. https://doi.org/10.1016/j.bbalip.2018.08.016
Granneman, J. G., Moore, H. P. H., Mottillo, E. P., Zhu, Z. X., & Zhou, L. (2011). Interactions of Perilipin-5 (Plin5) with adipose triglyceride lipase. Journal of Biological Chemistry, 286(7), 5126–5135. https://doi.org/10.1074/jbc.M110.180711
Wang, H., & Sztalryd, C. (2011). Oxidative tissue: Perilipin 5 links storage with the furnace. Trends in Endocrinology and Metabolism, 22(6), 197–203. https://doi.org/10.1016/j.tem.2011.03.008
Sanders, M. A., Madoux, F., Mladenovic, L., et al. (2015). Endogenous and synthetic ABHD5 ligands regulate ABHD5-perilipin interactions and lipolysis in fat and muscle. Cell Metabolism, 22(5), 851–860. https://doi.org/10.1016/j.cmet.2015.08.023
Gallardo-Montejano, V. I., Saxena, G., Kusminski, C. M., et al. (2016). Nuclear perilipin 5 integrates lipid droplet lipolysis with PGC-1alpha/SIRT1-dependent transcriptional regulation of mitochondrial function. Nature communications, 7, 12723. https://doi.org/10.1038/ncomms12723
Wang, H., Sreenivasan, U., Gong, D. W., et al. (2013). Cardiomyocyte-specific perilipin 5 overexpression leads to myocardial steatosis and modest cardiac dysfunction. Journal of Lipid Research., 54(4), 953–965. https://doi.org/10.1194/jlr.M032466
Benador, I. Y., Veliova, M., Mahdaviani, K., et al. (2018). Mitochondria bound to lipid droplets have unique bioenergetics, composition, and dynamics that support lipid droplet expansion. Cell Metabolism, 27(4), 869. https://doi.org/10.1016/j.cmet.2018.03.003
Bleck CKE, Kim Y, Willingham TB, Glancy B. Subcellular connectomic analyses of energy networks in striated muscle. Nature Communications 2018;9ARTN 5111 https://doi.org/10.1038/s41467-018-07676-y
Herms A, Bosch M, Reddy BJN, et al. AMPK activation promotes lipid droplet dispersion on detyrosinated microtubules to increase mitochondrial fatty acid oxidation. Nature Communications 2015;6ARTN 7176. https://doi.org/10.1038/ncomms8176
Yu, J. H., Zhang, S. Y., Cui, L. J., et al. (2015). Lipid droplet remodeling and interaction with mitochondria in mouse brown adipose tissue during cold treatment. Biochimica Et Biophysica Acta-Molecular Cell Research, 1853(5), 918–928. https://doi.org/10.1016/j.bbamcr.2015.01.020
Hu, R., Chen, B., Wang, Z. M., et al. (2019). Intriguing “chameleon” fluorescent bioprobes for the visualization of lipid droplet-lysosome interplay. Biomaterials, 203, 43–51. https://doi.org/10.1016/j.biomaterials.2019.03.002
Mizushima, N., & Komatsu, M. (2011). Autophagy: Renovation of cells and tissues. Cell, 147(4), 728–741. https://doi.org/10.1016/j.cell.2011.10.026
Kaushik, S., & Cuervo, A. M. (2018). The coming of age of chaperone-mediated autophagy. Nature Reviews Molecular Cell Biology, 19(6), 365–381. https://doi.org/10.1038/s41580-018-0001-6
Kaushik, S., & Cuervo, A. M. (2015). Degradation of lipid droplet-associated proteins by chaperone-mediated autophagy facilitates lipolysis. Nature Cell Biology, 17(6), 759–770. https://doi.org/10.1038/ncb3166
Singh, R., Kaushik, S., Wang, Y. J., et al. (2009). Autophagy regulates lipid metabolism. Nature, 458(7242), 1131-U64. https://doi.org/10.1038/nature07976
Settembre, C., Fraldi, A., Medina, D. L., & Ballabio, A. (2013). Signals from the lysosome: A control centre for cellular clearance and energy metabolism. Nature Reviews Molecular Cell Biology, 14(5), 283–296. https://doi.org/10.1038/nrm3565
Settembre, C., & Ballabio, A. (2011). TFEB regulates autophagy An integrated coordination of cellular degradation and recycling processes. Autophagy, 7(11), 1379–1381. https://doi.org/10.4161/auto.7.11.17166
Cohn, J. N., Ferrari, R., Sharpe, N., & Remodeling, I. F. C. (2000). Cardiac remodeling-concepts and clinical implications: A consensus paper from an international forum on cardiac remodeling. Journal of the American College of Cardiology, 35(3), 569–582. https://doi.org/10.1016/S0735-1097(99)00630-0
Khafagy, R., & Dash, S. (2021). Obesity and cardiovascular disease: The emerging role of inflammation. Frontiers in Cardiovascular Medicine, 8, 768119. https://doi.org/10.3389/fcvm.2021.768119
Bermudez, V., Duran, P., Rojas, E., et al. (2021). The sick adipose tissue: New insights into defective signaling and crosstalk with the myocardium. Front Endocrinol (Lausanne), 12, 735070. https://doi.org/10.3389/fendo.2021.735070
Joshi, P. H., & de Lemos, J. A. (2021). Diagnosis and management of stable angina: A review. JAMA, 325(17), 1765–1778. https://doi.org/10.1001/jama.2021.1527
Li, X., Yang, Y., Wang, Z., et al. (2021). Targeting non-coding RNAs in unstable atherosclerotic plaques: Mechanism, regulation, possibilities, and limitations. International Journal of Biological Sciences, 17(13), 3413–3427. https://doi.org/10.7150/ijbs.62506
Lubrano, V., & Balzan, S. (2021). Status of biomarkers for the identification of stable or vulnerable plaques in atherosclerosis. Clinical Science (London, England), 135(16), 1981–1997. https://doi.org/10.1042/CS20210417
Moore, K. J., & Tabas, I. (2011). Macrophages in the pathogenesis of atherosclerosis. Cell, 145(3), 341–55. https://doi.org/10.1016/j.cell.2011.04.005
Zhu, Z., Li, J., & Zhang, X. (2019). Astragaloside IV protects against oxidized low-density lipoprotein (ox-LDL)-induced endothelial cell injury by reducing oxidative stress and inflammation. Medical Science Monitor Basic Research, 25, 2132–2140. https://doi.org/10.12659/MSM.912894
Liao, Y., Zhu, E., & Zhou, W. (2021). Ox-LDL aggravates the oxidative stress and inflammatory responses of THP-1 macrophages by reducing the inhibition effect of miR-491-5p on MMP-9. Frontiers in Cardiovascular Medicine, 8, 697236. https://doi.org/10.3389/fcvm.2021.697236
Chellan, B., Reardon, C. A., Getz, G. S., & Hofmann Bowman, M. A. (2016). Enzymatically modified low-density lipoprotein promotes foam cell formation in smooth muscle cells via macropinocytosis and enhances receptor-mediated uptake of oxidized low-density lipoprotein. Arteriosclerosis, Thrombosis, and Vascular Biology, 36(6), 1101–13. https://doi.org/10.1161/ATVBAHA.116.307306
Li, W., Zhi, W., Liu, F., He, Z., Wang, X., & Niu, X. (2017). Atractylenolide I restores HO-1 expression and inhibits Ox-LDL-induced VSMCs proliferation, migration and inflammatory responses in vitro. Experimental Cell Research, 353(1), 26–34. https://doi.org/10.1016/j.yexcr.2017.02.040
Allahverdian, S., Chehroudi, A. C., McManus, B. M., Abraham, T., & Francis, G. A. (2014). Contribution of intimal smooth muscle cells to cholesterol accumulation and macrophage-like cells in human atherosclerosis. Circulation, 129(15), 1551–9. https://doi.org/10.1161/CIRCULATIONAHA.113.005015
Pi, H., Wang, Z., Liu, M., et al. (2019). SCD1 activation impedes foam cell formation by inducing lipophagy in oxLDL-treated human vascular smooth muscle cells. Journal of Cellular and Molecular Medicine, 23(8), 5259–5269. https://doi.org/10.1111/jcmm.14401
Hsieh, C. C., Li, C. Y., Hsu, C. H., et al. (2019). Mitochondrial protection by simvastatin against angiotensin II-mediated heart failure. British Journal of Pharmacology, 176(19), 3791–3804. https://doi.org/10.1111/bph.14781
Perrotta, I. (2017). Interaction between lipid droplets and endoplasmic reticulum in human atherosclerotic plaques. Ultrastructural Pathology, 41(1), 1–9. https://doi.org/10.1080/01913123.2016.1269861
Masumoto, N., Lanyon-Hogg, T., Rodgers, U. R., Konitsiotis, A. D., Magee, A. I., & Tate, E. W. (2015). Membrane bound O-acyltransferases and their inhibitors. Biochemical Society Transactions, 43, 246–252. https://doi.org/10.1042/Bst20150018
Rogers, M. A., Liu, J., Song, B. L., Li, B. L., Chang, C. C. Y., & Chang, T. Y. (2015). Acyl-CoA:Cholesterol acyltransferases (ACATs/SOATs): Enzymes with multiple sterols as substrates and as activators. Journal of Steroid Biochemistry and Molecular Biology, 151, 102–107. https://doi.org/10.1016/j.jsbmb.2014.09.008
Xu, Y. Q., Du, X. M., Turner, N., Brown, A. J., & Yang, H. Y. (2019). Enhanced acyl-CoA: Cholesterol acyltransferase activity increases cholesterol levels on the lipid droplet surface and impairs adipocyte function. Journal of Biological Chemistry, 294(50), 19306–19321. https://doi.org/10.1074/jbc.RA119.011160
Yagyu, H., Kitamine, T., Osuga, J., et al. (2000). Absence of ACAT-1 attenuates atherosclerosis but causes dry eye and cutaneous xanthomatosis in mice with congenital hyperlipidemia. Journal of Biological Chemistry, 275(28), 21324–21330. https://doi.org/10.1074/jbc.M002541200
Nissen, S. E. (2006). Effect of ACAT inhibition on the progression of coronary atherosclerosis (vol 354, pg 1253, 2006). New England Journal of Medicine, 355(6), 638–638.
Song, M. J., & Malhi, H. (2019). The unfolded protein response and hepatic lipid metabolism in non alcoholic fatty liver disease. Pharmacology & Therapeutics, 203, 107401. https://doi.org/10.1016/j.pharmthera.2019.107401
Borradaile, N. M., Han, X., Harp, J. D., Gale, S. E., Ory, D. S., & Schaffer, J. E. (2006). Disruption of endoplasmic reticulum structure and integrity in lipotoxic cell death. Journal of Lipid Research, 47(12), 2726–37. https://doi.org/10.1194/jlr.M600299-JLR200
Akoumi, A., Haffar, T., Mousterji, M., Kiss, R. S., & Bousette, N. (2017). Palmitate mediated diacylglycerol accumulation causes endoplasmic reticulum stress, Plin2 degradation, and cell death in H9C2 cardiomyoblasts. Experimental Cell Research, 354(2), 85–94. https://doi.org/10.1016/j.yexcr.2017.03.032
Ouimet, M., Franklin, V., Mak, E., Liao, X. H., Tabas, I., & Marcel, Y. L. (2011). Autophagy regulates cholesterol efflux from macrophage foam cells via lysosomal acid lipase. Cell Metabolism., 13(6), 655–667. https://doi.org/10.1016/j.cmet.2011.03.023
Tsai, T. H., Chen, E., Li, L., et al. (2017). The constitutive lipid droplet protein PLIN2 regulates autophagy in liver. Autophagy, 13(7), 1130–1144. https://doi.org/10.1080/15548627.2017.1319544
Ueno, M., Suzuki, J., Hirose, M., et al. (2017). Cardiac overexpression of perilipin 2 induces dynamic steatosis: Prevention by hormone-sensitive lipase. American Journal of Physiology-Endocrinology and Metabolism, 313(6), E699–E709. https://doi.org/10.1152/ajpendo.00098.2017
Mardani I, Dalen KT, Drevinge C, et al. Plin2-deficiency reduces lipophagy and results in increased lipid accumulation in the heart. Scientific Reports 2019;9ARTN 6909. https://doi.org/10.1038/s41598-019-43335-y
Garcia, E. J., Vevea, J. D., & Pon, L. A. (2018). Lipid droplet autophagy during energy mobilization, lipid homeostasis and protein quality control. Front Biosci (Landmark Ed), 23, 1552–1563. https://doi.org/10.2741/4660
Vazquez, M. M., Gutierrez, M. V., Salvatore, S. R., et al. (2020). Nitro-oleic acid, a ligand of CD36, reduces cholesterol accumulation by modulating oxidized-LDL uptake and cholesterol efflux in RAW264.7 macrophages. Redox Biology, 36, 101591. https://doi.org/10.1016/j.redox.2020.101591
Robichaud, S., Fairman, G., Vijithakumar, V., et al. (2021). Identification of novel lipid droplet factors that regulate lipophagy and cholesterol efflux in macrophage foam cells. Autophagy. https://doi.org/10.1080/15548627.2021.1886839
Ouimet, M., Ediriweera, H., Afonso, M. S., et al. (2017). microRNA-33 regulates macrophage autophagy in atherosclerosis. Arteriosclerosis Thrombosis and Vascular Biology, 37(6), 1058. https://doi.org/10.1161/Atvbaha.116.308916
Ishimaru K, Yoshioka K, Kano K, et al. Sphingosine kinase-2 prevents macrophage cholesterol accumulation and atherosclerosis by stimulating autophagic lipid degradation. Scientific Reports 2019;9ARTN 18329 https://doi.org/10.1038/s41598-019-54877-6
Groenewegen, A., Rutten, F. H., Mosterd, A., & Hoes, A. W. (2020). Epidemiology of heart failure. European Journal of Heart Failure, 22(8), 1342–1356. https://doi.org/10.1002/ejhf.1858
Nishihama, N., Nagayama, T., Makino, S., & Koishi, R. (2019). Mice lacking fat storage-inducing transmembrane protein 2 show improved profiles upon pressure overload-induced heart failure. Heliyon, 5(3), e01292. https://doi.org/10.1016/j.heliyon.2019.e01292
Schulze, P. C., Drosatos, K., & Goldberg, I. J. (2016). Lipid use and misuse by the heart. Circulation Research, 118(11), 1736–1751. https://doi.org/10.1161/Circresaha.116.306842
Benjamin, E. J., Blaha, M. J., Chiuve, S. E., et al. (2017). Heart Disease and Stroke Statistics-2017 Update: A Report From the American Heart Association. Circulation, 135(10), e146–e603. https://doi.org/10.1161/CIR.0000000000000485
Ljubkovic, M., Gressette, M., Bulat, C., et al. (2019). Disturbed fatty acid oxidation, endoplasmic reticulum stress, and apoptosis in left ventricle of patients with type 2 diabetes. Diabetes, 68(10), 1924–1933. https://doi.org/10.2337/db19-0423
Mizuno, M., Kuno, A., Yano, T., et al. (2018). Empagliflozin normalizes the size and number of mitochondria and prevents reduction in mitochondrial size after myocardial infarction in diabetic hearts. Physiological Reports, 6(12), e13741. https://doi.org/10.14814/phy2.13741
Holzem, K. M., Vinnakota, K. C., Ravikumar, V. K., et al. (2016). Mitochondrial structure and function are not different between nonfailing donor and end-stage failing human hearts. The FASEB Journal, 30(8), 2698–707. https://doi.org/10.1096/fj.201500118R
Andersson, L., Drevinge, C., Mardani, I., et al. (2017). Deficiency in perilipin 5 reduces mitochondrial function and membrane depolarization in mouse hearts. International Journal of Biochemistry & Cell Biology, 91, 9–13. https://doi.org/10.1016/j.biocel.2017.07.021
Liu, Z., Hua, J., Cai, W., et al. (2018). Nterminal truncated peroxisome proliferatoractivated receptorgamma coactivator1alpha alleviates phenylephrineinduced mitochondrial dysfunction and decreases lipid droplet accumulation in neonatal rat cardiomyocytes. Molecular Medicine Reports, 18(2), 2142–2152. https://doi.org/10.3892/mmr.2018.9158
Wang, C., Zhao, Y. L., Gao, X., et al. (2015). Perilipin 5 improves hepatic lipotoxicity by inhibiting lipolysis. Hepatology, 61(3), 870–882. https://doi.org/10.1002/hep.27409
Rababa’h, A. M., Guillory, A. N., Mustafa, R., & Hijjawi, T. (2018). Oxidative stress and cardiac remodeling: An updated edge. Current Cardiology Reviews, 14(1), 53–59. https://doi.org/10.2174/1573403X14666180111145207
Koncsos, G., Varga, Z. V., Baranyai, T., et al. (2016). Diastolic dysfunction in prediabetic male rats: Role of mitochondrial oxidative stress. American Journal of Physiology. Heart and Circulatory Physiology, 311(4), H927–H943. https://doi.org/10.1152/ajpheart.00049.2016
Zheng PF, Xie ZL, Yuan Y, et al. Plin5 alleviates myocardial ischaemia/reperfusion injury by reducing oxidative stress through inhibiting the lipolysis of lipid droplets. Scientific Reports 2017;7ARTN 42574. https://doi.org/10.1038/srep42574
Wang, C., Yuan, Y., Wu, J., et al. (2019). Plin5 deficiency exacerbates pressure overload-induced cardiac hypertrophy and heart failure by enhancing myocardial fatty acid oxidation and oxidative stress. Free Radical Biology and Medicine, 141, 372–382. https://doi.org/10.1016/j.freeradbiomed.2019.07.006
Wang ZG, Wang Y, Huang Y, et al. bFGF regulates autophagy and ubiquitinated protein accumulation induced by myocardial ischemia/reperfusion via the activation of the PI3K/Akt/mTOR pathway. Scientific Reports 2015;5ARTN 9287. https://doi.org/10.1038/srep09287
Wu, H., Ye, M., Yang, J., et al. (2015). Nicorandil protects the heart from ischemia/reperfusion injury by attenuating endoplasmic reticulum response-induced apoptosis through PI3K/Akt signaling pathway. Cellular Physiology and Biochemistry, 35(6), 2320–2332. https://doi.org/10.1159/000374035
Zhu, Z. D., Ye, J. Y., Niu, H., et al. (2018). Effects of microRNA-292-5p on myocardial ischemia-reperfusion injury through the peroxisome proliferator-activated receptor-alpha/-gamma signaling pathway. Gene Therapy, 25(3), 234–248. https://doi.org/10.1038/s41434-018-0014-y
Zhao, Y. B., Zhao, J., Zhang, L. J., et al. (2019). MicroRNA-370 protects against myocardial ischemia/reperfusion injury in mice following sevoflurane anesthetic preconditioning through PLIN5-dependent PPAR signaling pathway. Biomedicine & Pharmacotherapy, 113, 108697. https://doi.org/10.1016/j.biopha.2019.108697
Wang, Y., Chen, J., Cowan, D. B., & Wang, D. Z. (2021). Non-coding RNAs in cardiac regeneration: Mechanism of action and therapeutic potential. Seminars in Cell & Developmental Biology, 118, 150–162. https://doi.org/10.1016/j.semcdb.2021.07.007
Costantino, S., Akhmedov, A., Melina, G., et al. (2019). Obesity-induced activation of JunD promotes myocardial lipid accumulation and metabolic cardiomyopathy. European Heart Journal, 40(12), 997–1008. https://doi.org/10.1093/eurheartj/ehy903
Drevinge, C., Dalen, K. T., Mannila, M. N., et al. (2016). Perilipin 5 is protective in the ischemic heart. International Journal of Cardiology., 219, 446–454. https://doi.org/10.1016/j.ijcard.2016.06.037
Toyama, E. Q., Herzig, S., Courchet, J., et al. (2016). Metabolism. AMP-activated protein kinase mediates mitochondrial fission in response to energy stress. Science, 351(6270), 275–281. https://doi.org/10.1126/science.aab4138
Kolleritsch, S., Kien, B., Schoiswohl, G., et al. (2020). Low cardiac lipolysis reduces mitochondrial fission and prevents lipotoxic heart dysfunction in Perilipin 5 mutant mice. Cardiovascular Research, 116(2), 339–352. https://doi.org/10.1093/cvr/cvz119
Funding
This work was supported by the National Natural Science Foundation of China (81774232); National Youth Natural Science Foundation of China (82104721); the Second Batch of National “Ten Thousand Talents Program” Project Leading Talents (20160621); and Scientific Research Program of Tianjin Education Commission (2019KJ059).
Author information
Authors and Affiliations
Contributions
Xiaoying Guo and Qi Shi contributed to the conception of and writing of the manuscript.
Wanqin Zhang, Zhongwen Qi, Hao Lv, and Fujing Man contributed significantly to analysis and manuscript preparation.
Yingyu Xie, Yaping Zhu, and Junping Zhang helped perform the analysis with constructive discussions.
Corresponding authors
Ethics declarations
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Research involving human participants and/or animals
Not applicable
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Consent to Participate
Informed consent was obtained from all individual participants included in the study.
Consent for Publication
The participant has consented to the submission of the case report to the journal.
Competing Interests
The authors declare no competing interests.
Additional information
Associate Editor: Nicola Smart oversaw the review of this article.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yingyu Xie and Junping Zhang contributed equally to this work.
Rights and permissions
About this article
Cite this article
Guo, X., Shi, Q., Zhang, W. et al. Lipid Droplet—a New Target in Ischemic Heart Disease. J. of Cardiovasc. Trans. Res. 15, 730–739 (2022). https://doi.org/10.1007/s12265-021-10204-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12265-021-10204-x