Abstract
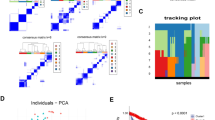
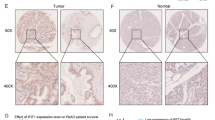
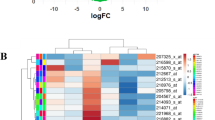
The aim was to expound the pathogenesis of prostate cancer and to identify the potentially biomarkers for prostate cancer (PC). DNA methylation microarray data GSE38240 containing 8 prostate cancer metastases and 4 normal prostate samples as well as gene expression profile data GSE26910 containing 6 prostate primary tumors and 6 normal samples were used. Differentially expressed genes (DEGs) and differently methylated sites of PC were screened and the regulatory network was constructed with DEGs-related transcription factors (TFs). The obtained hub genes were subjected to protein-protein interaction network analysis. Enrichment analysis of down-regulated DEGs were performed. Total 351 DEGs including 190 down-regulated and 161 up-regulated genes and 3234 differently methylated sites were identified. In total 69 DEGs-related TFs were found. Regulatory network contained 1301 nodes and 2527 connection pairs and that FOXA1 (forkhead box A1), BZRAP1-AS1 (benzodiazapine receptor associated protein 1 antisense RNA 1) and KRT8 (keratin 8) were the top three nodes of it. The enriched GO terms were mainly biological activity of the blood and cells-related. Total 29 DEGs (such as AGTR1, angiotensin II receptor, type 1) and 57 none-DEGs involved in the PPI network. Biological functions in blood circulation and the involved AGTR1 may play important roles in PC by gene-methylation. Besides, BZRAP1-AS1 may be novel biomarker related with PC.


Similar content being viewed by others
References
Aryee MJ, Liu W, Engelmann JC, Nuhn P, Gurel M, Haffner MC, Esopi D, Irizarry RA, Getzenberg RH, Nelson WG, Luo J, Xu J, Isaacs WB, Bova GS, Yegnasubramanian S (2013) DNA methylation alterations exhibit intraindividual stability and interindividual heterogeneity in prostate cancer metastases. Sci Transl Med 5(169):169ra110. https://doi.org/10.1126/scitranslmed.3005211
Bonci D, Coppola V, Musumeci M, Addario A, Giuffrida R, Memeo L, D'Urso L, Pagliuca A, Biffoni M, Labbaye C (2008) The miR-15a–miR-16-1 cluster controls prostate cancer by targeting multiple oncogenic activities. Nat Med 14(11):1271–1277
Currie D, McKnight A, Patterson C, Sadlier D, Maxwell A (2010) Investigation of ACE, ACE2 and AGTR1 genes for association with nephropathy in Type 1 diabetes mellitus. Diabet Med 27(10):1188–1194
Das PM, Singal R (2004) DNA methylation and cancer. J Clin Oncol 22(22):4632–4642
Dedeurwaerder S, Defrance M, Calonne E, Denis H, Sotiriou C, Fuks F (2011) Evaluation of the Infinium Methylation 450K technology. Epigenomics 3(6):771–784. https://doi.org/10.2217/epi.11.105
Elbaz N, Bedecs K, Masson M, Sutren M, Strosberg AD, Nahmias C (2000) Functional trans-inactivation of insulin receptor kinase by growth-inhibitory angiotensin II AT2 receptor. Mol Endocrinol 14(6):795–804
Gakhar G, YuLee G, Seandel M, Bander N, Nanus D (2013) Prostate circulating tumor cells metastasize to bone via E-selectin expressed on endothelial cells. Can Res 73(8 Supplement):5130
Gautier L, Cope L, Bolstad BM, Irizarry RA (2004) affy--analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20(3):307–315. https://doi.org/10.1093/bioinformatics/btg405
Karolchik D, Barber GP, Casper J, Clawson H, Cline MS, Diekhans M, Dreszer TR, Fujita PA, Guruvadoo L, Haeussler M, Harte RA, Heitner S, Hinrichs AS, Learned K, Lee BT, Li CH, Raney BJ, Rhead B, Rosenbloom KR, Sloan CA, Speir ML, Zweig AS, Haussler D, Kuhn RM, Kent WJ (2014) The UCSC Genome Browser database: 2014 update. Nucleic Acids Res 42(Database issue):D764–D770. https://doi.org/10.1093/nar/gkt1168
Kwabi-Addo B, Wang S, Furbert-Harris P, Yegnasubramanian S, Devaney J (2013) Functional characterization of Basonuclin 1 (BNC1): a novel tumor suppressor gene commonly downregulated in human prostate cancer. Can Res 73(8 Supplement):1974
Lin RJ, Xiao DW, Liao LD, Chen T, Xie ZF, Huang WZ, Wang WS, Jiang TF, Wu BL, Li EM (2012) MiR-142-3p as a potential prognostic biomarker for esophageal squamous cell carcinoma. J Surg Oncol 105(2):175–182
Lin Q, Tan HT, Lim HS, Chung MC (2013) Sieving through the cancer secretome. Biochim Biophys Acta 1834(11):2360–2371. https://doi.org/10.1016/j.bbapap.2013.01.030
Logothetis CJ, Gallick GE, Maity SN, Kim J, Aparicio A, Efstathiou E, Lin S-H (2013) Molecular classification of prostate cancer progression: foundation for marker-driven treatment of prostate cancer. Cancer Discov 3(8):849–861
Mitchell C, Zakar T, Sykes S, Pringle K, Lumbers E (2010) 141. Methylation of genes of the renin angiotensin system (ras) in early human amnion. Reprod Fertil Dev 22(9):59–59
Modarresi F, Faghihi MA, Lopez-Toledano MA, Fatemi RP, Magistri M, Brothers SP, van der Brug MP, Wahlestedt C (2012) Inhibition of natural antisense transcripts in vivo results in gene-specific transcriptional upregulation. Nat Biotechnol 30(5):453–459
Oh S, Shin S, Lightfoot SA, Janknecht R (2013) 14-3-3 proteins modulate the ETS transcription factor ETV1 in prostate cancer. Can Res 73(16):5110–5119
Planche A, Bacac M, Provero P, Fusco C, Delorenzi M, Stehle JC, Stamenkovic I (2011) Identification of prognostic molecular features in the reactive stroma of human breast and prostate cancer. PLoS One 6(5):e18640. https://doi.org/10.1371/journal.pone.0018640
Prasad NB, Somervell H, Tufano RP, Dackiw AP, Marohn MR, Califano JA, Wang Y, Westra WH, Clark DP, Umbricht CB (2008) Identification of genes differentially expressed in benign versus malignant thyroid tumors. Clin Cancer Res 14(11):3327–3337
Rhodes DR, Ateeq B, Cao Q, Tomlins SA, Mehra R, Laxman B, Kalyana-Sundaram S, Lonigro RJ, Helgeson BE, Bhojani MS (2009) AGTR1 overexpression defines a subset of breast cancer and confers sensitivity to losartan, an AGTR1 antagonist. Proc Natl Acad Sci U S A 106(25):10284–10289
Richiardi L, Fiano V, Grasso C, Zugna D, Delsedime L, Gillio-Tos A, Merletti F (2013) Methylation of APC and GSTP1 in non-neoplastic tissue adjacent to prostate tumour and mortality from prostate cancer. PLoS One 8(7):e68162
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Sherman BT, Huang d W, Tan Q, Guo Y, Bour S, Liu D, Stephens R, Baseler MW, Lane HC, Lempicki RA (2007) DAVID Knowledgebase: a gene-centered database integrating heterogeneous gene annotation resources to facilitate high-throughput gene functional analysis. BMC Bioinf. 8:426. https://doi.org/10.1186/1471-2105-8-426
Skårn M, Barøy T, Stratford EW, Myklebost O (2013) Epigenetic Regulation and Functional Characterization of MicroRNA-142 in Mesenchymal Cells. PLoS One 8(11):e79231
Szklarczyk D, Franceschini A, Kuhn M, Simonovic M, Roth A, Minguez P, Doerks T, Stark M, Muller J, Bork P (2011) The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res 39(suppl 1):D561–D568
Tjensvoll K, Nordgard O, Smaaland R (2014) Circulating tumor cells in pancreatic cancer patients: methods of detection and clinical implications. Int J Cancer 134(1):1–8. https://doi.org/10.1002/ijc.28134
Verma A, Facchina SL, Hirsch DJ, Song S-Y, Dillahey LF, Williams JR, Snyder SH (1998) Photodynamic tumor therapy: mitochondrial benzodiazepine receptors as a therapeutic target. Mol Med 4(1):40
Viola MV, Fromowitz F, Oravez S, Deb S, Finkel G, Lundy J, Hand P, Thor A, Schlom J (1986) Expression of ras oncogene p21 in prostate cancer. N Engl J Med 314(3):133–137
Voeller H, Sugars L, Pretlow T, Gelmann E (1994) p53 oncogene mutations in human prostate cancer specimens. J Urol 151(2):492–495
Wang F, Wang X-S, Yang G-H, Zhai P-F, Xiao Z, Xia L-Y, Chen L-R, Wang Y, Wang X-Z, Bi L-X (2012) miR-29a and miR-142-3p downregulation and diagnostic implication in human acute myeloid leukemia. Mol Biol Rep 39(3):2713–2722
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Tan, J., Jin, X. & Wang, K. Integrated Bioinformatics Analysis of Potential Biomarkers for Prostate Cancer. Pathol. Oncol. Res. 25, 455–460 (2019). https://doi.org/10.1007/s12253-017-0346-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-017-0346-8