Abstract
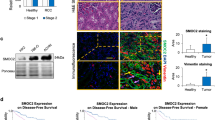
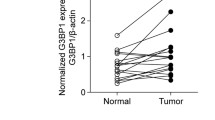
Suppressor of cancer cell invasion (SCAI) has been originally characterized as a tumor suppressor inhibiting metastasis in different human cancer cells, and it has been suggested that SCAI expression declines in tumors. The expression patterns and role of SCAI during physiological and pathophysiological processes is still poorly understood. Earlier we demonstrated that SCAI is regulating the epithelial-mesenchymal transition of proximal tubular epithelial cells, it is downregulated during renal fibrosis and it is overexpressed in Wilms’ tumors. Here we bring further evidence for the involvement of SCAI during cell plasticity and we examine the prognostic value and expression patterns of SCAI in various tumors. SCAI prevented the activation of the SMA promoter induced by angiotensin II. SCAI expression decreased in a model of endothelial-mesenchymal transition and increased during iPS reprogramming of fibroblasts. During renal fibrosis SCAI expression declined, as evidenced in a rat model of renal transplant rejection and in TGF-β1 overexpressing transgenic mice. High expression of SCAI correlated with better survival in patients with breast and lung cancers. Intriguingly, in the case of other cancers (gastric, prostate, colorectal) high SCAI expression correlated with poor survival of patients. Finally, we bring evidence for SCAI overexpression in colorectal cancer patients, irrespective of stage or metastatic status of the disease, suggesting a diverse role of SCAI in various diseases and cancer.




Similar content being viewed by others
References
Medjkane S, Perez-Sanchez C, Gaggioli C, Sahai E, Treisman R (2009) Myocardin-related transcription factors and SRF are required for cytoskeletal dynamics and experimental metastasis. Nat Cell Biol 11(3):257–268. doi:10.1038/ncb1833
Fan L, Sebe A, Peterfi Z, Masszi A, Thirone AC, Rotstein OD, Nakano H, McCulloch CA, Szaszi K, Mucsi I, Kapus A (2007) Cell contact-dependent regulation of epithelial-myofibroblast transition via the rho-rho kinase-phospho-myosin pathway. Mol Biol Cell 18(3):1083–1097. doi:10.1091/mbc.E06-07-0602
Sebe A, Masszi A, Zulys M, Yeung T, Speight P, Rotstein OD, Nakano H, Mucsi I, Szaszi K, Kapus A (2008) Rac, PAK and p38 regulate cell contact-dependent nuclear translocation of myocardin-related transcription factor. FEBS Lett 582(2):291–298. doi:10.1016/j.febslet.2007.12.021
Brandt DT, Baarlink C, Kitzing TM, Kremmer E, Ivaska J, Nollau P, Grosse R (2009) SCAI acts as a suppressor of cancer cell invasion through the transcriptional control of beta1-integrin. Nat Cell Biol 11(5):557–568. doi:10.1038/ncb1862
Fintha A, Gasparics A, Fang L, Erdei Z, Hamar P, Mozes MM, Kokeny G, Rosivall L, Sebe A (2013) Characterization and role of SCAI during renal fibrosis and epithelial-to-mesenchymal transition. Am J Pathol 182(2):388–400. doi:10.1016/j.ajpath.2012.10.009
Chen X, Hu W, Xie B, Gao H, Xu C, Chen J (2014) Downregulation of SCAI enhances glioma cell invasion and stem cell like phenotype by activating Wnt/beta-catenin signaling. Biochem Biophys Res Commun 448(2):206–211. doi:10.1016/j.bbrc.2014.04.098
Kressner C, Nollau P, Grosse R, Brandt DT (2013) Functional interaction of SCAI with the SWI/SNF complex for transcription and tumor cell invasion. PLoS One 8(8):e69947. doi:10.1371/journal.pone.0069947
Ishikawa M, Nishijima N, Shiota J, Sakagami H, Tsuchida K, Mizukoshi M, Fukuchi M, Tsuda M, Tabuchi A (2010) Involvement of the serum response factor coactivator megakaryoblastic leukemia (MKL) in the activin-regulated dendritic complexity of rat cortical neurons. J Biol Chem 285(43):32734–32743. doi:10.1074/jbc.M110.118745
Burns WC, Velkoska E, Dean R, Burrell LM, Thomas MC (2010) Angiotensin II mediates epithelial-to-mesenchymal transformation in tubular cells by ANG 1-7/MAS-1-dependent pathways. Am J Physiol Ren Physiol 299(3):F585–F593. doi:10.1152/ajprenal.00538.2009
Li R, Liang J, Ni S, Zhou T, Qing X, Li H, He W, Chen J, Li F, Zhuang Q, Qin B, Xu J, Li W, Yang J, Gan Y, Qin D, Feng S, Song H, Yang D, Zhang B, Zeng L, Lai L, Esteban MA, Pei D (2010) A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 7(1):51–63. doi:10.1016/j.stem.2010.04.014
Samavarchi-Tehrani P, Golipour A, David L, Sung HK, Beyer TA, Datti A, Woltjen K, Nagy A, Wrana JL (2010) Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell 7(1):64–77. doi:10.1016/j.stem.2010.04.015
Evelyn CR, Wade SM, Wang Q, Wu M, Iniguez-Lluhi JA, Merajver SD, Neubig RR (2007) CCG-1423: a small-molecule inhibitor of RhoA transcriptional signaling. Mol Cancer Ther 6(8):2249–2260. doi:10.1158/1535-7163.MCT-06-0782
Bodor C, Nagy JP, Vegh B, Nemeth A, Jenei A, MirzaHosseini S, Sebe A, Rosivall L (2012) Angiotensin II increases the permeability and PV-1 expression of endothelial cells. Am J Physiol Cell Physiol 302(1):C267–C276. doi:10.1152/ajpcell.00138.2011
Krizbai IA, Gasparics A, Nagyoszi P, Fazakas C, Molnar J, Wilhelm I, Bencs R, Rosivall L, Sebe A (2015) Endothelial-mesenchymal transition of brain endothelial cells: possible role during metastatic extravasation. PLoS One 10(3):e0119655. doi:10.1371/journal.pone.0119655
Grabundzija I, Wang J, Sebe A, Erdei Z, Kajdi R, Devaraj A, Steinemann D, Szuhai K, Stein U, Cantz T, Schambach A, Baum C, Izsvak Z, Sarkadi B, Ivics Z (2013) Sleeping beauty transposon-based system for cellular reprogramming and targeted gene insertion in induced pluripotent stem cells. Nucleic Acids Res 41(3):1829–1847. doi:10.1093/nar/gks1305
Sebe A, Ivics Z (2016) Reprogramming of human fibroblasts to induced pluripotent stem cells with sleeping beauty transposon-based stable gene delivery. Methods Mol Biol 1400:419–427. doi:10.1007/978-1-4939-3372-3_26
Wang D, Chang PS, Wang Z, Sutherland L, Richardson JA, Small E, Krieg PA, Olson EN (2001) Activation of cardiac gene expression by myocardin, a transcriptional cofactor for serum response factor. Cell 105(7):851–862
Hamar P, Liu S, Viklicky O, Szabo A, Muller V, Heemann U (2000) Cyclosporine a and azathioprine are equipotent in chronic kidney allograft rejection. Transplantation 69(7):1290–1295
Hamar P, Liptak P, Heemann U, Ivanyi B (2005) Ultrastructural analysis of the fisher to Lewis rat model of chronic allograft nephropathy. Transpl Int : Official journal of the European Society for Organ Transplantation 18(7):863–870. doi:10.1111/j.1432-2277.2005.00146.x
Sanderson N, Factor V, Nagy P, Kopp J, Kondaiah P, Wakefield L, Roberts AB, Sporn MB, Thorgeirsson SS (1995) Hepatic expression of mature transforming growth factor beta 1 in transgenic mice results in multiple tissue lesions. Proc Natl Acad Sci U S A 92(7):2572–2576
Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, Szallasi Z (2010) An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat 123(3):725–731. doi:10.1007/s10549-009-0674-9
Gyorffy B, Surowiak P, Budczies J, Lanczky A (2013) Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS One 8(12):e82241. doi:10.1371/journal.pone.0082241
Gyorffy B, Molnar B, Lage H, Szallasi Z, Eklund AC (2009) Evaluation of microarray preprocessing algorithms based on concordance with RT-PCR in clinical samples. PLoS One 4(5):e5645. doi:10.1371/journal.pone.0005645
Mihaly Z, Kormos M, Lanczky A, Dank M, Budczies J, Szasz MA, Gyorffy B (2013) A meta-analysis of gene expression-based biomarkers predicting outcome after tamoxifen treatment in breast cancer. Breast Cancer Res Treat 140(2):219–232. doi:10.1007/s10549-013-2622-y
Li X, Zhuang S (2014) Recent advances in renal interstitial fibrosis and tubular atrophy after kidney transplantation. Fibrogenesis Tissue Repair 7:15. doi:10.1186/1755-1536-7-15
Tullius SG, Nieminen M, Bechstein WO, Jonas S, Steinmuller T, Qun Y, Pratschke J, Graser E, Sinha P, Volk HD, Neuhaus P, Tilney NL (1998) Contribution of early acute rejection episodes to chronic rejection in a rat kidney retransplantation model. Kidney Int 53(2):465–472. doi:10.1046/j.1523-1755.1998.00757.x
Grande MT, Sanchez-Laorden B, Lopez-Blau C, De Frutos CA, Boutet A, Arevalo M, Rowe RG, Weiss SJ, Lopez-Novoa JM, Nieto MA (2015) Snail1-induced partial epithelial-to-mesenchymal transition drives renal fibrosis in mice and can be targeted to reverse established disease. Nat Med 21(9):989–997. doi:10.1038/nm.3901
Lin L LD, Liang H, Xue L, Su C, Liu M. (2015) MiR-1228 promotes breast cancer cell growth and metastasis through targeting SCAI protein. Int J Clin Exp Pathol 1 (8(6)):6646–6655
Lobry C, Oh P, Aifantis I (2011) Oncogenic and tumor suppressor functions of Notch in cancer: it's NOTCH what you think. J Exp Med 208(10):1931–1935. doi:10.1084/jem.20111855
Nozawa RS, Nagao K, Masuda HT, Iwasaki O, Hirota T, Nozaki N, Kimura H, Obuse C (2010) Human POGZ modulates dissociation of HP1alpha from mitotic chromosome arms through aurora B activation. Nat Cell Biol 12(7):719–727. doi:10.1038/ncb2075
Brauchle M, Yao Z, Arora R, Thigale S, Clay I, Inverardi B, Fletcher J, Taslimi P, Acker MG, Gerrits B, Voshol J, Bauer A, Schubeler D, Bouwmeester T, Ruffner H (2013) Protein complex interactor analysis and differential activity of KDM3 subfamily members towards H3K9 methylation. PLoS One 8(4):e60549. doi:10.1371/journal.pone.0060549
Moldovan GL, Pfander B, Jentsch S (2007) PCNA, the maestro of the replication fork. Cell 129(4):665–679. doi:10.1016/j.cell.2007.05.003
Mailand N, Gibbs-Seymour I, Bekker-Jensen S (2013) Regulation of PCNA-protein interactions for genome stability. Nat Rev Mol Cell Biol 14(5):269–282. doi:10.1038/nrm3562
Milutinovic S, Zhuang Q, Szyf M (2002) Proliferating cell nuclear antigen associates with histone deacetylase activity, integrating DNA replication and chromatin modification. J Biol Chem 277(23):20974–20978. doi:10.1074/jbc.M202504200
Murzina N, Verreault A, Laue E, Stillman B (1999) Heterochromatin dynamics in mouse cells. Mol Cell 4(4):529–540. doi:10.1016/s1097-2765(00)80204-x
Hansen RK, Mund A, Poulsen SL, Sandoval M, Klement K, Tsouroula K, Tollenaere MA, Raschle M, Soria R, Offermanns S, Worzfeld T, Grosse R, Brandt DT, Rozell B, Mann M, Cole F, Soutoglou E, Goodarzi AA, Daniel JA, Mailand N, Bekker-Jensen S (2016) SCAI promotes DNA double-strand break repair in distinct chromosomal contexts. Nat Cell Biol. doi:10.1038/ncb3436
Isobe SY, Nagao K, Nozaki N, Kimura H, Obuse C (2017) Inhibition of RIF1 by SCAI allows BRCA1-mediated repair. Cell Rep 20(2):297–307. doi:10.1016/j.celrep.2017.06.056
Bai F, Chan HL, Scott A, Smith MD, Fan C, Herschkowitz JI, Perou CM, Livingstone AS, Robbins DJ, Capobianco AJ, Pei XH (2014) BRCA1 suppresses epithelial-to-mesenchymal transition and stem cell dedifferentiation during mammary and tumor development. Cancer Res 74(21):6161–6172. doi:10.1158/0008-5472.CAN-14-1119
Acknowledgments
We thank Dr. Marcell A. Szász for valuable discussions, and Erika Sklánitzné Samodai for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declare no conflicts of interest.
Rights and permissions
About this article
Cite this article
Gasparics, Á., Kökény, G., Fintha, A. et al. Alterations in SCAI Expression during Cell Plasticity, Fibrosis and Cancer. Pathol. Oncol. Res. 24, 641–651 (2018). https://doi.org/10.1007/s12253-017-0293-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-017-0293-4