Abstract
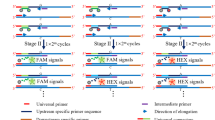
During diagnostic workflow when detecting sequence alterations, sometimes it is important to design an algorithm that includes screening and direct tests in combination. Normally the use of direct test, which is mainly sequencing, is limited. There is an increased need for effective screening tests, with “closed tube” during the whole process and therefore decreasing the risk of PCR product contamination. The aim of this study was to design such a closed tube, detection probe based screening assay to detect different kind of sequence alterations in the exon 11 of the human c-kit gene region. Inside this region there are variable possible deletions and single nucleotide changes. During assay setup, more probe chemistry formats were screened and tested. After some optimization steps the taqman probe format was selected.





Similar content being viewed by others
References
Antonescu CR, Sommer G, Sarran L, Tschernyavsky SJ, Riedel E, Woodruff JM, Robson M, Maki R, Brennan MF, Ladanyi M, DeMatteo RP, Besmer P (2003) Association of KIT exon 9 mutations with nongastric primary site and aggressive behavior: KIT mutation analysis and clinical correlates of 120 gastrointestinal stromal tumors. Clin Cancer Res 9(9):3329–3337
Kondiah K, Swanepoel R, Paweska JT, Burt FJ (2010) A simple-probe real-time PCR assay for genotyping reassorted and non-reassorted isolates of Crimean-Congo hemorrhagic fever virus in southern Africa. J Virol Methods 169(1):34–38
Grannemann S, Landt O, Breuer S, Blömeke B (2005) LightTyper assay with locked-nucleic-acid-modified oligomers for genotyping of the toll-like receptor 4 polymorphisms A896G and C1196T. Clin Chem 51(8):1523–1525
Frances F, Corella D, Sorli JV, Guillen M, Gonzalez JI, Portoles O (2005) Validating a rapid method for detecting common polymorphisms in the APOA5 gene by melting curve analysis using LightTyper. Clin Chem 51:1279–1282
Francés F, Portolés O, Sorlí JV, Guillén M, González JI, Corella D (2008) Single tube optimisation of APOE genotyping based on melting curve analysis. Clin Biochem 41(10–11):923–926
McGuigan FE, Ralston SH (2002) Single nucleotide polymorphism detection: allelic discrimination using TaqMan. Psychiatr Genet 12(3):133–136
Hui L, DelMonte T, Ranade K (2008) Genotyping using the TaqMan assay. Curr Protoc Hum Genet. doi:10.1002/0471142905.hg0210s56
Zhou L, Myers AN, Vandersteen JG, Wang L, Wittwer CT (2004) Closed-tube genotyping with unlabeled oligonucleotide probes and a saturating DNA dye. Clin Chem 50(8):1328–1335
Roche Applied Science (2004) LightCycler Manual, Technical Note No. LC 18/2004: Assay formats for use in real-time PCR. https://www.roche-applied-science.com/sis/rtpcr/lightcycler/lightcycler—docs/technical—notes/lc—18.pdf
Mackay J, Landt O (2007) Real-time PCR fluorescent chemistries. Methods Mol Biol 353:237–261
Korbie DJ, Mattick JS (2008) Touchdown PCR for increased specificity and sensitivity in PCR amplification. Nat Protoc 3(9):1452–1456
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Becságh, P., Szakács, O. Setting Up a Probe Based, Closed Tube Real-Time PCR Assay for Focused Detection of Variable Sequence Alterations. Pathol. Oncol. Res. 20, 813–818 (2014). https://doi.org/10.1007/s12253-014-9759-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-014-9759-9