Abstract
Mechanisms driving the consumption and transport of tidal marsh nutrients and energy by fishes are of key interest in the San Francisco Estuary, CA, USA. By combining multiple data sources (gill-net catches, gut contents, channel morphology, tides), we modeled spatial and temporal patterns of fish abundance and gut fullness across a tidal marsh elevation gradient. Channel depth, microhabitat, and tide were important predictors of fish abundance and gut fullness. Species, feeding guild, and season were also important to fish abundance but not to gut fullness, suggesting that abundance was more related to physical constraints of shallow water than to prey availability. Multiple feeding guilds overlapped in space and time at interaction hotspots in subtidal channel habitat near the marsh entrance. In contrast, fish use of shallow intertidal marsh channels was more variable and indicated tradeoffs between foraging and predation. Gut content analysis revealed moderate-to-high gut fullness for all feeding guilds and models predicted high gut fullness in subtidal reaches during tidal flooding, after which fish fed intensively throughout the marsh. While mysids, amphipods, and detritus were common prey among feeding guilds, variation in prey consumption was apparent. Overall, complex tidal marsh hydrogeomorphology driving land-water exchange and residence time may diversify and enhance benthic and pelagic food web pathways to fishes and invertebrates. Furthermore, these findings substantiate the notion that dynamic tidal marshes in this system can support robust secondary production, foraging by multiple feeding guilds, and trophic transfer by fishes to the estuarine mosaic.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Estuarine and coastal tidal marshes are among the most productive ecosystems in the world but are increasingly threatened by anthropogenic activities (Baker et al. 2020; Gilby et al. 2020). In the San Francisco Estuary, California (hereafter, “SFE”), widespread diking and draining during the nineteenth and twentieth centuries resulted in a 90–95% reduction in tidal marsh habitat area (Nichols et al. 1986). The transition from a marsh-dominated ecosystem to an open water-dominated ecosystem, combined with flow alterations and nonnative species introductions, likely disrupted food web processes (Cloern et al. 2016) and contributed to many native fishes becoming threatened, endangered, or extinct (Moyle et al. 2010). While evidence is accumulating that remnant tidal marshes provide important habitat for estuarine fish assemblages (Visintainer et al. 2006; Colombano et al. 2020a; Colombano et al. 2020b), the extent to which they provide foraging opportunities for multiple guilds and support secondary production transfers to the open estuary remains a key knowledge gap.
Mechanisms driving food web pathways to secondary consumers and production transfers to estuarine or coastal habitats have received much attention in the salt marsh literature, particularly along the southeastern and Gulf coasts, USA (Childers et al. 2000; Deegan et al. 2000; Kneib 2000; Smith et al. 2000; Zimmerman et al. 2000). The notion is that hydrogeomorphic complexity controls tidal connectivity between marsh plains and subtidal channels, which can influence the exchange and transport of benthic marsh nutrients, organisms, and detritus (Kneib 2000). While direct transport may occur during periods of tidal or fluvial exchange (Teal 1962; Nixon 1980; Odum 2000), indirect transport may occur when primary and secondary consumers (fishes and invertebrates) move out of marshes via tidal migration, juvenile recruitment to adult habitats, or consumption by transient predators (in a series of “trophic relays” (Kneib 2000)). Trophic relays have now been examined in numerous systems across the globe (Litvin and Weinstein 2004; Hollingsworth and Connolly 2006; Able et al. 2009; Nelson et al. 2012; Baker et al. 2013; Ziegler et al. 2019; Lesser et al. 2020; Nemerson and Able 2020), which have revealed substantial geographic variation in tidal marsh structure and function (Ziegler et al. 2021).
The contribution of tidal marshes to estuarine food webs has more recently become a topic of interest in the SFE (Herbold et al. 2014; Brown et al. 2016). Recent evidence suggests that benthic and pelagic food web pathways (e.g., marsh detritus and phytoplankton) may support diverse fish assemblages (Durand 2015; Schroeter et al. 2015; Young et al. 2020). Consumer resource use often varies among species, life histories, seasons, and marsh structural characteristics [e.g., channel order, amount of edge or vegetation; (Visintainer et al. 2006; Gewant and Bollens 2012; Whitley and Bollens 2014; Montgomery 2017)]. For transient fish species that move among habitats in the estuarine mosaic, tidal marsh area has been linked to foraging success (e.g., delta smelt [Hypomesus transpacificus]; (Hammock et al. 2019)) and nursery function (e.g., Sacramento splittail [Pogonichthys macrolepidotus], striped bass [Morone saxatilis]; Colombano et al. 2020a). Collectively, these studies suggest that tidal marshes in the SFE conform to established salt marsh concepts; however, targeted studies are needed to determine mechanisms driving these patterns.
Here, we conducted an integrative study on how tidal dynamics structure fish foraging patterns across a natural marsh habitat gradient in the SFE. First, we asked: How is the fish assemblage spatially and temporally distributed across the transition zone from low-elevation to high-elevation tidal marsh channels? We hypothesized that the distribution of fishes would be related to differences in structural characteristics such as channel depth or microhabitat type (e.g., confluence vs. edge; Kneib 2000). We also posited that changes in tide height, direction, and amplitude would play important roles in the spatial structure of fish feeding guilds because these factors govern access to tidal marsh edges (Colombano et al. 2020b). To address this question, we surveyed fish assemblages along the elevation gradient over a range of tidal conditions and then constructed spatially explicit generalized additive mixed models using soap-film smoothers, high-resolution bathymetry data, and continuous, high-frequency tidal time series data to make predictions about fish distribution and abundance.
Second, we asked: What prey items do fishes consume in tidal marsh? How is prey consumption by fish spatially and temporally distributed across the gradient from low-elevation to high-elevation tidal marsh channels? We expected filter-feeding planktivorous fishes to consume zooplankton associated with pelagic food webs and benthivorous fishes to consume macroinvertebrates associated with benthic food webs. But we also expected to find overlap among diets because many species present in the marsh are opportunists (Feyrer et al. 2003; O’Rear 2012; Schroeter et al. 2015). We hypothesized that gut fullness would increase with incoming tides, since flooding intertidal areas would expand surface area for feeding, and that piscivores would respond to movement of prey into the marsh. To address these questions, we conducted gut content analysis of fishes in three feeding guilds (planktivores, benthivores, piscivores) and implemented similar models used in the first part of the study to predict the percent gut fullness across the elevation gradient and a range of tidal conditions. Based on our findings and previous work in this system, we then developed a conceptual model of mechanisms driving benthic and pelagic food web pathways in tidal marshes of the SFE.
Materials and methods
Study system
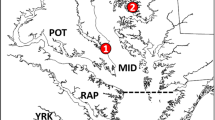
Suisun Marsh is a 470-km2 brackish tidal marsh located 80 km upstream from the Golden Gate Bridge (Whitcraft et al. 2011) at the geographic center of the northern SFE (Moyle et al. 2014). Suisun Marsh features a mosaic of habitat types including large and small tidal channels, fringing marsh, mudflats, seasonal wetlands, managed ponds, and upland transition zones (Manfree 2014). Suisun Slough is a large, sinuous channel that originates in Grizzly Bay and extends northward to Suisun City, and is a migratory corridor for fishes moving between bay and tidal marsh habitat. Ultimately draining into Suisun Slough, Spring Branch is a meandering dendritic tidal channel network lined with fringing marsh (dominated by Schoenoplectus and Typha spp.) and extensive tidal marsh plains (Fig. 1). This natural (undiked) tidal marsh provides temperature refugia when inundated by high-amplitude nighttime tides in spring and summer (Enright et al. 2013). It also features a gradual elevation gradient that shifts from low-elevation (subtidal) channels near the mouth to high-elevation (intertidal) channels toward the terminus. Midway along the longitudinal axis of the channel, there is a “tidal excursion boundary” marking a transition from higher tidal exchange (i.e., with other channel networks) in subtidal reaches with deeper, alternating point bars and cut banks to lower tidal exchange in relatively uniform and shallow intertidal reaches (Robinson et al. 2016; Stumpner et al. 2020). The terminus of the marsh connects to a seasonal freshwater creek draining the upland watershed.
Surveys
Fish sampling
In spring and summer 2015, we intensively surveyed fish assemblages along a tidal marsh elevation gradient in Spring Branch over nine separate days (Apr 29; May 5, 12, 17, 28; Jul 8, 15, 24; Aug 12) [see Supplement Table 1 for more detail on sampling effort]. We deployed 140 individual gill-net sets using experimental gill nets (27.4-m L and 2.4-m H with alternating panels of mesh ranging from 1.27 cm2 with 2.54-cm stretch to 7.62 cm2 with 15.24-cm stretch) from a 4.3-m-long jon boat. Nets were deployed for 10 to 30 minutes to nonselectively capture a range of fish species and sizes; depths per each set generally changed only a few centimeters. Random sampling locations were stratified by microhabitat along the longitudinal axis of the channel. Following designations in Kneib (2000), we targeted four microhabitat types: (1) the single large subtidal-open-water confluence (SOC) at the channel mouth; (2) subtidal-open-water areas (STO); (3) intertidal-subtidal confluences (ISC); and marsh surface edges (MSE; Fig. 2). On each sampling day during daylight hours, gill nets were deployed on incoming and outgoing tides to compare tidal movements of fishes. All captured fishes were identified and measured in standard length (“SL”; mm; Table 1) and returned to the water after recovering in an aerated water bath or stored on ice after blunt trauma euthanasia for future gut content analysis in the laboratory (IACUC protocol #18883). Abundant species with documented ontogenetic shifts in feeding habits (Moyle 2002) were split into size classes for analysis.
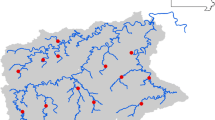
Spring Branch channel bathymetry measured by single-beam sonar and predicted using a soap-film smoother constrained by channel bank boundaries. Depths are expressed in relation to mean sea level (MSL=0; NAVD88 tidal datum). Gill-net sample locations are shown as symbols according to microhabitat category: subtidal-open-water confluence (SOC), subtidal-open-water (STO), intertidal-subtidal confluence (ISC), and marsh surface edges (MSE).
Gut content analysis
We examined guts of common fishes collected in Spring Branch gill-net samples. First, we measured SL (mm) and weighed (g) each whole fish, dissected the gut, and ranked percent gut fullness on a 0–4 scale: 0 (0%), 1 (1–25%), 2 (26–50%), 3 (51–75%), or 4 (76–100%). We then preserved gut contents in a 10% formalin solution for at least 1 month before transferring to a 70% ethanol solution. The contents of each sample were identified to the lowest practical taxonomic group under a dissecting microscope and then counted. Diagnostic bones such as cleithra were used to identify fish taxa in partially digested gut contents (Hansel et al. 1988). For quality control and assurance, approximately 20% of samples were randomly selected for a double-blind survey, the results of which were either averaged among the two observations or reconciled by a third person and then averaged among the three observations.
To characterize prey items found in the guts of fish species, we calculated the percent frequency of occurrence (%F) of each prey group, which is considered the most robust and interpretable measure of diet composition (Baker et al. 2014). Empty guts were excluded from the analysis. To compare the importance of prey items across fish species with diverse diets, we grouped invertebrates into broad taxonomic categories (e.g., “mysid” for opossum shrimp belonging to Mysidae).
Tide data collection
Water depth data were queried from the National Oceanographic and Atmospheric Administration National Estuarine Research Reserve System System-wide Monitoring Program (NOAA NERRS SWMP; NOAA 2019). All data collection and QA/QC methods followed national SWMP standards. A water quality datasonde (Yellow Springs International, Inc. 6600 instrument) in Spring Branch (known to the NERRS as “First Mallard Branch”) was affixed to a large wooden piling at the mouth of the channel and measured tide height (i.e., water depth [m]) from a non-vented sensor corrected for changes in barometric pressure on 15-minute intervals. Using established methods, we used the datasonde values as an input for the open-source software program TidalTrend (Donovan and Ayers 2019) to determine tide direction (incoming or outgoing tide) and spring-neap (high-amplitude “spring” tide or low-amplitude “neap” tide) categories and then matched them up to our gill-net sample dates and times.
Bathymetry data collection
Bathymetric data used in this study were gathered in January and February 2015 using single-beam sonar supplemented with side-scan imagery following established methods (Handley 2015). Single-beam sonar is a cost-effective, accurate, and robust method for collecting data in shallow water and is recommended for surveys in waters with an average depth less than 5 m (USACE 2013). Surveys were conducted from a 4.3-m-long jon boat using a Humminbird XTM 9 HDSI 180 T (20° cone) commercial side-scan fishfinder attached to the bottom of a 1.2-m galvanized steel pipe with a Topcon HiPer V Real-Time Kinematic GPS (RTK) rover attached to the top. An RTK base station was set up near the survey location to broadcast corrections, and a Topcon Tesla handheld tablet was used to record the RTK position data at 1 Hz during the survey. Sonar data were recorded on the fishfinder receiver and merged with RTK positioning during post-processing. The jon boat was steered back and forth from bank to bank in the study channel network, including connected navigable intertidal channels, to collect transverse transect data with a final longitudinal transect conducted as a cross-check [see Supplement Fig. S1]. Depth profiles were first processed using Leraand Engineering SonarTRX-SI Pro software to remove artifacts and submerged aquatic vegetation, then digitized and converted to GIS vector data, and finally clipped to remove points beyond shoreline banks from a local DEM (USGS 2015). Finite area smoothing was used to model channel bathymetry with soap-film methods using the packages, “raster,” “sf,” and “mgcv” in Program R (Wood et al. 2008; Simpson 2016; Wood 2017; Pebesma 2018; Hijmans and Van Etten 2019; Team 2019) [see Supplement for bathymetry modeling].
Gut fullness and feeding guild modeling
To measure habitat selection by different feeding guilds, we evaluated the relative effects of season, channel depth (from the bathymetry survey), microhabitat, tide height and direction, and spring-neap tidal cycles on fish abundance. We constructed GAMMs using a negative binomial distribution with a log link function, specified smoothed varying-intercept and varying-slope terms, and included a soap-film smoother and a 1-m resolution grid of knots to constrain model fitting to x,y coordinates within channel bank boundaries. At each x,y coordinate representing a gill-net sample, we extracted predicted depths from the bathymetry raster at the gill net’s midpoint and used the resulting data as a predictor variable. Gill-net minutes per sample were log-transformed and included as an offset variable to adjust for differences in sampling effort. Following established methods for the “mgcv” package, we fit all models using the “gam” function (Wood 2017; Pedersen et al. 2019). Using a stepwise approach, we included each predictor variable of interest and then added interactions to test for conditional effects (e.g., to test whether the effect of tide height on fish abundance varied by fish species). We evaluated the results using the “summary.gam,” “gam.check,” and “gam.vcomp” functions, which produce model coefficients and diagnostic information about the fitting procedure, visualizations of residuals, and estimates of the random effect variances. According to established methods (Wood 2017), models were first estimated with maximum likelihood (ML) and compared with corrected Akaike information criterion (AICc) scores, effective degrees of freedom (edf), and percent (%) deviance explained. The selected model, with the lowest AICc and highest % deviance explained, was then estimated using restricted maximum likelihood (REML) to reduce bias of the ML procedure.
We used a similar approach to evaluate the relative effects of season, channel depth, microhabitat, tide height, tide direction, and spring-neap tidal cycles on percent gut fullness (i.e., the proportion of a fish gut filled with prey items), as a measure of foraging success among guilds. To do so, we specified a beta distribution with a logit link function, which is typically implemented for response variables that have values distributed between 0 and 1 (Wood 2017).
Using top-ranked models for fish abundance and gut fullness, we generated predictions via the “predict.gam” function (Wood 2017) along the axis of the channel using 1-m resolution grids of x,y coordinates and corresponding predicted depth values from the bathymetry raster. Specifically, we summarized median predicted fish abundance and gut fullness to evaluate our hypotheses about feeding guilds and to visualize the effects of channel depth, tide height, and tide direction. To achieve this, we visualized distributions as heat maps using color gradients to represent the range of predicted values and cross-referenced the predictions with fitted relationships from the GAMMs (Figs. S2 and S3).
Results
Catch
We captured 348 fish in our gill-net surveys in Spring Branch. Nine species from three feeding guilds represented 99% of the total survey catch (Tables 1 and 2). Other species caught were Chinook salmon (Oncorhynchus tshawytscha; n = 2) and yellowfin goby (Acanthogobius flavimanus; n = 1). The two most abundant species captured by the gill nets were striped bass (n = 82) and Sacramento splittail (n = 162). In general, the gill net more effectively sampled fish with standard lengths greater than 70 mm, so young-of-the-year fish were not well represented in our catches (Table 1).
Gut fullness and diets
We collected 286 fish from six species for gut content analysis (Table 1). Three fish had empty guts (fullness score of 0) and the remaining 283 fish examined had a range of gut fullness scores (Fig. 3). The highest proportion of guts that were full (score of 4) occurred in striped bass, splittail, and American shad (Alosa sapidissima), while the highest proportion of tule perch (Hysterocarpus traski) scored 3 and the highest proportion of threadfin shad (Dorosoma petenense) scored 1. Notably, Sacramento pikeminnow (Ptychocheilus grandis; n = 4) all had relatively low proportions of contents (score of 1).
According to the %F analysis, dominant prey taxa across fish species were mysids (Neomysis kadiakensis, Alienacanthomysis macropsis, and Hyperacanthomysis longirostris) and amphipods (Eogammarus confervicolus, Gammarus daiberi, Corophiidae, and Talitridae) (Table 2). Detritus was consistently present in gut contents of all species and was the only documented item in Sacramento pikeminnow and threadfin shad. American shad and striped bass ≤ 100 mm both had diets dominated by mysids but also consumed detritus and unidentified fish. In addition, the %F of amphipods ranged from 7% in Sacramento splittail > 100 mm to 30% in tule perch, 8% in striped bass ≤ 100 mm, and 20% in striped bass > 100 mm. Tule perch also consumed isopods (Isopoda; 30%). Both size classes of splittail consumed nematodes (Nematoda), and splittail > 100 mm also consumed clams (Corbulidae). Striped bass > 100 mm consumed the most diverse prey groups of those identified (i.e., fish, invertebrates, detritus) among all fish species analyzed; their diets were dominated by mysids (35%), followed by three-spined stickleback (Gasterosteus aculeatus; 10%), shrimp (Palaemonidae; 7%), amphipods (20%), and unidentified fish (6%); other fish species found in striped bass > 100 mm included tule perch, common carp (Cyprinus carpio), gobies (Gobiidae), sculpins (Cottidae), and Mississippi silverside (Menidia audens). Harpacticoida was also recorded in a single striped bass stomach, representing the only copepod in the entire gut content analysis.
Spatial distribution of gut fullness
The best fitting gut fullness model included channel depth and tide direction (Tables 3 and 4). Notably, species, fish guild, and season were poor predictors of gut fullness. Median predicted gut fullness values ranged from 0.55 to 0.80, which coincide with gut fullness scores of 3 and 4 (Fig. 4). Predicted gut fullness hotspots were distributed across the elevation gradient but were concentrated in two main areas: (1) in subtidal channel habitat downstream of the tidal excursion boundary and especially near the channel mouth, during incoming tides, and (2) across the entire elevation gradient, but particularly in shallow reaches, during outgoing tides. Factor-smooth interactions yielded a relatively stronger effect of incoming tides in deeper water on gut fullness compared to outgoing tides in shallower water (Fig. S2).
Spatial distribution of feeding guilds
The best fitting fish count model selected included the variables season, species, channel depth, tide height, tide direction, and gill-net minutes (Tables 3 and 5). Two categorical variables (season and species) and two continuous variables (channel depth and tide height) had the strongest effects on fish abundance along the elevation gradient. Notably, the factor-smoothed interactions, channel depth by species and tide height by species, had strong effects on abundance (P < 0.001). Likewise, microhabitat, which covaried with channel depth, also had an effect: subtidal-open-water confluence (SOC), subtidal-open-water sites (STO), and intertidal-subtidal confluence (ISC) were all significant (P < 0.05); in contrast, marsh surface edge (MSE) was not. Feeding guild was also an important predictor of fish abundance according to the AICc comparison (Table 5). Model predictions indicated distinct patterns in the distribution and abundance of feeding guilds as a function of the elevation gradient (i.e., channel depth), tide height, and tide direction, as shown by heat map visualizations (Fig. 5a–c) and corresponding species-specific model fits (Fig. S3).
Distribution of (A) planktivores, (B) benthivores, and (C) piscivores represented as median predicted fish catch per unit effort (CPUE) according to tide direction and tidal height (lower=1.2 m; higher=1.8 m) from the selected model. Smoothed relationships between CPUE and x,y coordinates were achieved using a soap-film smoother constrained by channel bank boundaries.
Model results indicated that planktivores had the lowest median predicted catches among feeding guilds, ranging from 2 to 12.5% CPUE (Fig. 5a). In addition, models predicted a broad spatial distribution along the elevation gradient, with higher predicted median CPUE concentrated in deeper waters at lower outgoing tides and highly variable predicted median CPUE above the tidal excursion boundary. Species driving the observed differences included American shad, which were generally more abundant, larger in size, and captured in deeper areas, whereas threadfin shad and striped bass ≤ 100 mm were smaller and more commonly captured in shallower areas (Fig. S3). Overall, planktivore catches were predicted to be higher during lower-than-average tide heights in both tide directions. Consistent hotspots across all tidal conditions occurred near the main channel mouth where pockets of deeper water were more evenly distributed.
Model results indicated that benthivores had median predicted CPUE ranging from 7 to 22%, with slightly higher catches occurring during incoming tides (Fig. 5b). The predicted distribution along the elevation gradient generally decreased toward the marsh terminus; however, several species showed no relationship to channel depth (e.g., Sacramento splittail > 100 mm, common carp; Fig. S3). Model results suggested that piscivores had a similar range in median predicted CPUE as benthivores (5–25%). However, catches of larger piscivores (i.e., Sacramento pikeminnow and striped bass) were generally concentrated near the deeper channel mouth during outgoing tides, particularly during higher-than-average tide heights (Figs. 5c, S3a, b).
Discussion
Hydrogeomorphic complexity structured fish foraging patterns in the natural tidal marsh channel network, suggesting that fishes were adapted to dynamic, rapidly changing conditions and opportunities to access food and refuge. Overall, larger piscivores occurred more often in subtidal channel habitat, reflecting an association with deeper water at the channel mouth; benthivores used the entire elevation gradient but were also more common in subtidal channel habitat; and planktivores were more broadly distributed across the elevation gradient during lower incoming tides. Interaction hotspots, where high concentrations of multiple feeding guilds overlap in space and time (Kneib 2000), occurred in subtidal reaches of the main channel and especially at the subtidal-open-water confluence (i.e., the channel mouth). In contrast, habitat partitioning occurred in shallow intertidal reaches above the tidal excursion boundary. Similar to other systems, small-sized fish likely had to balance the potential benefits of prey availability and predator avoidance in shallow reaches with the potential risks of predation in deeper reaches (Rypel et al. 2007; Whitfield 2017; Boswell et al. 2019; Jones et al. 2020). Collectively, the results of this study reflect a healthy tidal marsh food web that supports fish foraging and secondary production transfers as described in the conceptual model.
Temporal habitat partitioning
Our modeling results indicate that temporal habitat partitioning occurred among feeding guilds and size ranges. Planktivores had the highest predicted catches in subtidal reaches during lower outgoing tides, which may have reflected avoidance of larger piscivores that were concentrated in deeper water at the channel mouth, a phenomenon seen at a larger scale in an Atlantic estuary (Nemerson and Able 2004). Lower catches of planktivores during higher tides likely reflected their movement into flooded areas to feed, similar to bay anchovy (Anchoa mitchilli) in Delaware Bay (Nemerson and Able 2020). Large benthivores were more abundant when channels were sufficiently flooded, especially on incoming tides, likely due to inundation of soft-bottom sediments and marsh surfaces and margins that provide substrate for benthic macroinvertebrates. The larger benthivore’s feeding habits, higher tolerance of shallow water, and lower risk of predation by piscivores likely explain their high overlap with other guilds.
Gut fullness models predicted fullness hotspots in subtidal channel habitat below the tidal excursion boundary during incoming tides, suggesting that prey availability attracted fishes into the marsh. Fish distribution models showed corresponding hotspots among fish guilds near the channel mouth. The sparser distribution of fish, especially larger piscivores, in shallow areas above the tidal excursion boundary was likely due to physiological limitations on large-bodied fish (Paterson and Whitfield 2000). Larger piscivores likely only opportunistically use shallow channels, which shift rapidly in depth, temperature, and dissolved oxygen according to tidal and diurnal cycles (Rountree and Able 1997), to pursue small fish not adequately sampled by gill nets. Gut fullness models also predicted a broad distribution of high gut fullness along the elevation gradient during outgoing tides, suggesting that prey acquisition increased toward the marsh terminus as a function of tidal inundation duration.
Fish feeding guild habits
Gut content analyses supported our use of conventional feeding guilds. Striped bass, which is typically considered the dominant piscivore in SFE tidal marshes (Moyle 2002), consumed many benthic fishes and invertebrates. For striped bass > 100 mm, three-spined stickleback, common carp, tule perch, gobies/sculpins, amphipods, isopods, crabs, crayfish, and shrimp represented 50% of total %F in diets (Table 2). The occurrence of detritus (8%) and chironomids (2%) provides further evidence of a terrestrial-derived component. Importantly, the occurrence of mysids was also high (35%), demonstrating the importance of tidal marshes in providing high-quality macrozooplankton prey (Feyrer et al. 2003). In contrast, Sacramento pikeminnow guts contained detritus and were nearly empty (score of 1), likely reflecting their propensity to evacuate their guts upon capture in gill nets (Moyle, UCD, pers. comm.). However, recent evidence suggests high dietary overlap and possible functional equivalence of striped bass and Sacramento pikeminnow (Stompe et al. 2020), which may translate into higher predation pressure (i.e., exerted by both species) than was characterized in this study. The absence of microzooplankton (e.g., copepods) from gut contents may have been an artifact of the size selectivity of gill nets for larger planktivores.
The dominance of macrozooplankton (mysids) and benthic macroinvertebrates (especially amphipods) across feeding guilds highlights the potential of tidal marshes to support robust benthic and pelagic food webs (Durand 2015; Schroeter et al. 2015; Young et al. 2020). The consumption of gammarid, corophiid, and talitrid amphipods is noteworthy because they are associated with complex benthic and epibenthic habitats such as soft-bottom sediments, root wads, and submerged aquatic vegetation such as Stuckenia spp. (Kelley 1966; de Szalay and Resh 1996). As the tide goes out, these organisms may be more evenly distributed across the marsh and thus support dispersed feeding by fishes, as shown in Fig. 4. The low incidence of empty guts suggests that tidal marshes can provide abundant food resources for all feeding guilds.
Support for established salt marsh concepts
Established salt marsh ecology concepts such as the trophic relay hypothesis are generally supported by the findings of this study. Different feeding guilds moved asynchronously along the elevation gradient with the tides, presumably to maximize foraging in the marsh and/or minimize predation risk at the channel mouth (Kneib 2000). The small contribution of resident marsh-platform fish species in this system is a notable caveat, however (Brown 2003). Recent research on geographic variation in feeding habits in salt marshes across the eastern US shows that tidal amplitude regulates the strength of trophic transfers from marsh-platform species (Ziegler et al. 2019). Given the relatively low percentage of high tides that flood over channel banks in Suisun Marsh (~15%; Enright et al. 2013), it is perhaps not surprising that three-spined stickleback, the only small resident fish species known to use the flooded marsh surface in this system (Moyle 2002), was one of the many prey items found in larger striped bass, an ambush predator that waits for prey to flush off of the marsh plain in its native range (e.g., Delaware Bay; Nemerson and Able 2003). Consumption of marsh-platform fish appears to be opportunistic (i.e., similar to an infrequently flooded salt marsh in a subtropical Australian estuary; Hollingsworth and Connolly 2006), whereas consumption of in-channel fish and invertebrates appears to be more reliable. Despite these differences, tidal regulation of predator-prey interactions and subsequent trophic transfers by fishes are consistent with Kneib (2000).
Conceptual model of benthic and pelagic food web coupling
Based on the findings of our study and others (Durand 2015; Schroeter et al. 2015; Robinson et al. 2016; Montgomery 2017; Young et al. 2020), we developed a conceptual model highlighting how complex hydrogeomorphology, which drives land-water exchange and residence time, may couple benthic (i.e., detritus-based) and pelagic (i.e., phytoplankton-based) food webs in tidal marshes (Fig. 6). An intact tidal marsh food web includes planktivores that feed on zooplankton in the open water column, benthivores that feed on macroinvertebrates along soft-bottom sediments and marsh surfaces and edges, and piscivores that effectively capture benthic and pelagic prey moving into and out of tidal marsh channels. Not only does this arrangement allow fishes of many trophic levels to forage on a broad range of prey groups within the marsh, it provides robust food web pathways to fishes that can accumulate and transport marsh nutrients and energy to the estuarine mosaic in a series of trophic relays (sensu Kneib 2000).
Proposed conceptual model of benthic and pelagic pathways to secondary consumers (fishes and invertebrates) in San Francisco Estuary tidal marshes. Complex hydrogeomorphology enhances benthic-pelagic coupling, or the intersection of detritus-based food webs (e.g., emergent vegetation) resulting from increased exchange across the land-water boundary (left) and phytoplankton-based food webs resulting from in situ production (e.g., algae) in the water column and mixed residence time of water (right). Fishes can transport tidal marsh nutrients and energy to the estuarine mosaic in a series of trophic relays (Kneib 2000).
Secondary production as a management target
Conserving, restoring, and managing healthy tidal marshes in the SFE require consideration of the complex suite of hydrogeomorphic features that drive secondary production and foraging success. Secondary production of fish and invertebrates in nearshore habitats indicates process-based ecosystem function and health and thus serves as a composite metric for evaluating the performance of remnant and restored marshes (Weinstein et al. 2014; Weinstein and Litvin 2016; Layman and Rypel 2020). Assessing tidal marsh ecosystem function and health using secondary production is becoming increasingly important given the potentially dire effects of climate change and sea level rise on tidal marsh habitats, food webs, and fisheries support (Colombano et al. 2021; Baker et al. 2020).
References
Able, K.W., K.M.M. Jones, and D.A. Fox. 2009. Large nektonic fishes in Marsh Creek habitats in the Delaware Bay estuary. Northeastern Naturalist 16 (1): 27–44. https://doi.org/10.1656/045.016.0103.
Baker, R., B. Fry, L.P. Rozas, and T.J. Minello. 2013. Hydrodynamic regulation of salt marsh contributions to aquatic food webs. Marine Ecology Progress Series 490: 37–52. https://doi.org/10.3354/meps10442.
Baker, R., A. Buckland, and M. Sheaves. 2014. Fish gut content analysis: robust measures of diet composition. Fish and Fisheries 15 (1): 170–177. https://doi.org/10.1111/faf.12026.
Baker, R., M.D. Taylor, K.W. Able, M.W. Beck, J. Cebrian, D.D. Colombano, R.M. Connolly, C. Currin, et al. 2020. Fisheries rely on threatened salt marshes. Science 370: 670–671. https://doi.org/10.1126/science.abe9332.
Boswell, K.M., M.E. Kimball, G. Rieucau, J.G.A. Martin, D.A. Jacques, D. Correa, and D.M. Allen. 2019. Tidal stage mediates periodic asynchrony between predator and prey nekton in salt marsh creeks. Estuaries and Coasts 42 (5): 1342–1352. https://doi.org/10.1007/s12237-019-00553-x.
Brown, L. R. 2003. Will tidal wetland restoration enhance populations of native fishes? San Francisco Estuary and Watershed Science 1. https://doi.org/10.15447/sfews.2003v1iss1art2.
Brown, L. R., W. Kimmerer, J. L. Conrad, S. Lesmeister, and A. Mueller-Solger. 2016. Food webs of the Delta, Suisun Bay, and Suisun Marsh: an update on current understanding and possibilities for management. San Francisco Estuary and Watershed Science 14. https://doi.org/10.15447/sfews.2016v14iss3art4.
CalAtlas. 2012. California Geospatial Clearinghouse: highways, railroads, California state boundary. State of California. (June 2012).
Childers, D.L., J.W. Day, and H.N. Mckellar. 2000. Twenty more years of marsh and estuarine flux studies: revisiting Nixon (1980). In Concepts and Controversies in Tidal Marsh Ecology, ed. M.P. Weinstein and D.A. Kreeger, 391–423. Dordrecht: Springer Netherlands. https://doi.org/10.1007/0-306-47534-0_18.
Cloern, J.E., A. Robinson, A. Richey, L. Grenier, R. Grossinger, K.E. Boyer, J. Burau, E.A. Canuel, et al. 2016. Primary Production in the Delta: Then and Now. Primary production in the delta: then and now. San Francisco Estuary and Watershed Science 14 (3): 14. https://doi.org/10.15447/sfews.2016v14iss3art1.
Colombano, D., S.Y. Litvin, S.B. Alford, R. Baker, M.A. Barbeau, J. Cebrian, R.M. Connolly, C.A. Currin, et al. 2021. Climate change implications for tidal marshes and food web linkages to estuarine and coastal nekton. Estuaries and Coasts. https://doi.org/10.1007/s12237-020-00891-1.
Colombano, D.D., A.D. Manfree, T.A. O’Rear, J.R. Durand, and P.B. Moyle. 2020a. Estuarine-terrestrial habitat gradients enhance nursery function for resident and transient fishes in the San Francisco Estuary. Marine Ecology Progress Series 637: 141–157. https://doi.org/10.3354/meps13238.
Colombano, D.D., J.M. Donovan, D.E. Ayers, T.A. O’Rear, and P.B. Moyle. 2020b. Tidal effects on marsh habitat use by three fishes in the San Francisco Estuary. Environmental Biology of Fishes 103: 605–623. https://doi.org/10.1007/s10641-020-00973-w.
Deegan, L.A., J.E. Hughes, and R.A. Rountree. 2000. Salt Marsh Ecosystem Support of Marine Transient Species. In Concepts and Controversies in Tidal Marsh Ecology, ed. M.P. Weinstein and D.A. Kreeger, 333–365. Dordrecht: Springer Netherlands. https://doi.org/10.1007/0-306-47534-0_16.
Donovan, J.M., and D.E. Ayers. 2019. TidalTrend software technical report. Dept. of Interior: US Geological Survey.
Durand, J.R. 2015. A Conceptual Model of the Aquatic Food Web of the Upper San Francisco Estuary. A conceptual model of the aquatic food web of the upper San Francisco Estuary. San Francisco Estuary and Watershed Science 13. https://doi.org/10.15447/sfews.2015v13iss3art5.
DWR. 2007. Suisun LIDAR dataset. Department of Water Resources. (January 2012).
Enright, C., S.D. Culberson, and J.R. Burau. 2013. Broad Timescale Forcing and Geomorphic Mediation of Tidal Marsh Flow and Temperature Dynamics. Estuaries and Coasts 36 (6): 1319–1339. https://doi.org/10.1007/s12237-013-9639-7.
Feyrer, F., B. Herbold, S.A. Matern, and P.B. Moyle. 2003. Dietary shifts in a stressed fish assemblage: Consequences of a bivalve invasion in the San Francisco Estuary. Environmental Biology of Fishes 67 (3): 277–288. https://doi.org/10.1023/A:1025839132274.
Gesch, D., M. Oimoen, S. Greenlee, C. Nelson, M. Steuck, and D. Tyler. 2002. The national elevation dataset. Photogrammetric Engineering and Remote Sensing 68: 5–32.
Gewant, D., and S.M. Bollens. 2012. Fish assemblages of interior tidal marsh channels in relation to environmental variables in the upper San Francisco Estuary. Environmental Biology of Fishes 94 (2): 483–499. https://doi.org/10.1007/s10641-011-9963-3.
Gilby, B.L., M.P. Weinstein, R. Baker, J. Cebrian, S.B. Alford, A. Chelsky, D. Colombano, R.M. Connolly, C.A. Currin, I.C. Feller, A. Frank, J.A. Goeke, L.A. Goodridge Gaines, F.E. Hardcastle, C.J. Henderson, C.W. Martin, A.E. McDonald, B.H. Morrison, A.D. Olds, J.S. Rehage, N.J. Waltham, and S.L. Ziegler. 2020. Human actions alter tidal marsh seascapes and the provision of ecosystem services. Estuaries and Coasts. https://doi.org/10.1007/s12237-020-00830-0.
Handley, T. B. 2015. Cost-effective methods for accurately measuring shallow water bathymetry with single-beam sonar. M.S., United States -- California: University of California, Davis.
Hansel, H.C., S.D. Duke, P.T. Lofy, and G.A. Gray. 1988. Use of Diagnostic Bones to Identify and Estimate Original Lengths of Ingested Prey Fishes. Transactions of the American Fisheries Society 117 (1): 55–62. https://doi.org/10.1577/1548-8659(1988)117<0055:UODBTI>2.3.CO;2.
Kelley, D.W. 1966. Zoobenthos of the Sacramento-San Joaquin Delta. In J. L. Turner and D. W. Kelley, 133:113–139, ed. Ecological Studies of the Sacramento-San Joaquin Estuary, part 1: zooplankton, zoobenthos, and fishes of San Pablo and Suisun bays, zooplankton and zoobenthos of the Delta. California Department of Fish and Game Bulletin.
Herbold, B., D. M. Baltz, L. Brown, R. Grossinger, W. Kimmerer, P. Lehman, C. (Si) Simenstad, C. Wilcox, et al. 2014. The Role of Tidal Marsh Restoration in Fish Management in the San Francisco Estuary. San Francisco Estuary and Watershed Science. https://doi.org/10.15447/sfews.2014v12iss1art112, 1.
Hijmans, R.J., and J. Van Etten. 2019. raster: Geographic data analysis and modeling. (version R Program 3.0-2). Vienna, Austria: The R Foundation.
Hollingsworth, A., and R.M. Connolly. 2006. Feeding by fish visiting inundated subtropical saltmarsh. Journal of Experimental Marine Biology and Ecology 336 (1): 88–98. https://doi.org/10.1016/j.jembe.2006.04.008.
Jones, T.R., C.J. Henderson, A.D. Olds, R.M. Connolly, T.A. Schlacher, B.J. Hourigan, L.A. Goodridge Gaines, and B.L. Gilby. 2020. The Mouths of Estuaries Are Key Transition Zones that Concentrate the Ecological Effects of Predators. Estuaries and Coasts. https://doi.org/10.1007/s12237-020-00862-6.
Kneib, R.T. 2000. Salt Marsh Ecoscapes and Production Transfers by Estuarine Nekton in the Southeastern United States. In Concepts and Controversies in Tidal Marsh Ecology, ed. M.P. Weinstein and D.A. Kreeger, 267–291. Dordrecht: Springer Netherlands. https://doi.org/10.1007/0-306-47534-0_13.
Layman, C.A., and A.L. Rypel. 2020. Secondary production is an underutilized metric to assess restoration initiatives. Food Webs 25: e00174. https://doi.org/10.1016/j.fooweb.2020.e00174.
Lesser, J.S., C.A. Bechtold, L.A. Deegan, and J.A. Nelson. 2020. Habitat decoupling via saltmarsh creek geomorphology alters connection between spatially-coupled food webs. Estuarine, Coastal and Shelf Science 241: 106825. https://doi.org/10.1016/j.ecss.2020.106825.
Litvin, S.Y., and M.P. Weinstein. 2004. Multivariate analysis of stable-isotope ratios to infer movements and utilization of estuarine organic matter by juvenile weakfish (Cynoscion regalis). Canadian Journal of Fisheries and Aquatic Sciences 61 (10): 1851–1861. https://doi.org/10.1139/f04-121.
Manfree, A. D. 2014. Landscape change in Suisun Marsh. Ph.D., United States -- California: University of California, Davis.
Montgomery, J. R. 2017. Foodweb dynamics in shallow tidal sloughs of the San Francisco Estuary. M.S., United States -- California: University of California, Davis.
Moyle, P.B. 2002. Inland Fishes of California: Revised and Expanded. University of California Press.
Moyle, P.B., J.R. Lund, W.A. Bennett, and W.E. Fleenor. 2010. Habitat variability and complexity in the upper San Francisco Estuary. San Francisco Estuary and Watershed Science 8. https://doi.org/10.15447/sfews.2010v8iss3art1.
Nelson, J., R. Wilson, F. Coleman, C. Koenig, D. DeVries, C. Gardner, and J. Chanton. 2012. Flux by fin: fish-mediated carbon and nutrient flux in the northeastern Gulf of Mexico. Marine Biology 159: 365–372. https://doi.org/10.1007/s00227-011-1814-4.
Nemerson, D.M., and K.W. Able. 2003. Spatial and temporal patterns in the distribution and feeding habits of Morone saxatilis in marsh creeks of Delaware Bay, USA. Fisheries Management and Ecology 10 (5): 337–348. https://doi.org/10.1046/j.1365-2400.2003.00371.x.
Nemerson, D.M., and K.W. Able. 2004. Spatial patterns in diet and distribution of juveniles of four fish species in Delaware Bay marsh creeks: factors influencing fish abundance. Marine Ecology Progress Series 276: 249–262. https://doi.org/10.3354/meps276249.
Nemerson, D.M., and K.W. Able. 2020. Diel and tidal influences on the abundance and food habits of four young-of-the-year fish in Delaware Bay, USA, marsh creeks. Environmental Biology of Fishes 103 (3): 251–268. https://doi.org/10.1007/s10641-020-00956-x.
Nixon, S.W. 1980. Between Coastal Marshes and Coastal Waters — A Review of Twenty Years of Speculation and Research on the Role of Salt Marshes in Estuarine Productivity and Water Chemistry. In Estuarine and Wetland Processes: With Emphasis on Modeling, ed. P. Hamilton and K. B. Macdonald, 437–525. Marine Science. Boston, MA: Springer US. https://doi.org/10.1007/978-1-4757-5177-2_20.
NOAA. 2019. Water depth dataset. National Oceanographic and Atmospheric Administration Centralized Data Management Office. August 2019.
Odum, E.P. 2000. Tidal Marshes as Outwelling/Pulsing Systems. In Concepts and Controversies in Tidal Marsh Ecology, ed. M.P. Weinstein and D.A. Kreeger, 3–7. Dordrecht: Springer Netherlands. https://doi.org/10.1007/0-306-47534-0_1.
O’Rear, T. A. 2012. Diet of an Introduced Estuarine Population of White Catfish in California. M.S., United States -- California: University of California, Davis.
Paterson, A.W., and A.K. Whitfield. 2000. Do Shallow-water Habitats Function as Refugia for Juvenile Fishes? Estuarine, Coastal and Shelf Science 51 (3): 359–364. https://doi.org/10.1006/ecss.2000.0640.
Pebesma, E. 2018. Simple Features for R: Standardized Support for Spatial Vector Data. The R Journal 10: 439. 10.32614/RJ-2018-009.
Pedersen, E.J., D.L. Miller, G.L. Simpson, and N. Ross. 2019. Hierarchical generalized additive models in ecology: an introduction with mgcv. PeerJ 7. PeerJ Inc.: e6876. https://doi.org/10.7717/peerj.6876.
Robinson, A., A. Richey, J. E. Cloern, K. E. Boyer, J. R. Burau, E. A. Canuel, J. F. DeGeorge, E. R. Howe, et al. 2016. Primary production in the Sacramento-San Joaquin Delta: A science strategy to quantify change and identify future potential. 781. Richmond, CA: San Francisco Estuary Institute - Aquatic Science Center.
Rountree, R.A., and K.W. Able. 1997. Nocturnal Fish Use of New Jersey Marsh Creek and Adjacent Bay Shoal Habitats. Estuarine, Coastal and Shelf Science 44 (6): 703–711. https://doi.org/10.1006/ecss.1996.0134.
Rypel, A.L., C.A. Layman, and D.A. Arrington. 2007. Water depth modifies relative predation risk for a motile fish taxon in Bahamian tidal creeks. Estuaries and Coasts 30 (3): 518–525. https://doi.org/10.1007/BF03036517.
San Francisco Estuary Institute (SFEI) 2012. Bay Area EcoAtlas Modern baylands. June 2012.
Schroeter, R.E., T.A. O’Rear, M.J. Young, and P.B. Moyle. 2015. The Aquatic Trophic Ecology of Suisun Marsh, San Francisco Estuary, California, During Autumn in a Wet Year. San Francisco Estuary and Watershed Science 13. https://doi.org/10.15447/sfews.2015v13iss3art6.
Simpson, G. 2016. Soap-film smoothers & lake bathymetries. From the Bottom of the Heap.
Smith, K.J., G.L. Taghon, and K.W. Able. 2000. Trophic Linkages in Marshes: Ontogenetic Changes in Diet for Young-of-the-Year Mummichog, Fundulus Heteroclitus. In Concepts and Controversies in Tidal Marsh Ecology, ed. M.P. Weinstein and D.A. Kreeger, 121–237. Dordrecht: Springer Netherlands. https://doi.org/10.1007/0-306-47534-0_11.
Stompe, D. K., J. D. Roberts, C. A. Estrada, D. M. Keller, N. M. Balfour, and A. I. Banet. 2020. Sacramento River Predator Diet Analysis: A Comparative Study. San Francisco Estuary and Watershed Science 18. 10.15447/sfews.2020v18iss1art4.
Stumpner, P.R., J.R. Burau, and A.L. Forrest. 2020. A Lagrangian-to-Eulerian Metric to Identify Estuarine Pelagic Habitats. Estuaries and Coasts. https://doi.org/10.1007/s12237-020-00861-7.
de Szalay, F. A., and V. H. Resh. 1996. Spatial and temporal variability of trophic relationships among aquatic macroinvertebrates in a seasonal marsh. Wetlands 16. Springer: 458–466.
Teal, J. M. 1962. Energy flow in the salt marsh ecosystem of Georgia. Ecology 43: 614–624. https://doi.org/10.2307/1933451.
Team, R.C. 2019. R Project for Statistical Computing (2019). Vienna: Austria.
USACE. 2013. Hydrographic surveying. US Army Corps of Engineers.
USGS. 2004. National hydrography dataset. US Geological Survey. (June 2012).
USGS. 2015. EarthExplorer orthophoto dataset. US Geological Survey. (September 2015).
Visintainer, T., S. Bollens, and C. Simenstad. 2006. Community composition and diet of fishes as a function of tidal channel geomorphology. Marine Ecology Progress Series 321: 227–243. https://doi.org/10.3354/meps321227.
Weinstein, M.P., and S.Y. Litvin. 2016. Macro-restoration of tidal wetlands: a whole estuary approach. Ecological Restoration 34: 27–38.https://doi.org/10.3368/er.34.1.27.
Weinstein, M.P., S.Y. Litvin, and J.M. Krebs. 2014. Restoration ecology: ecological fidelity, restoration metrics, and a systems perspective. Ecological Engineering 65: 71–87. https://doi.org/10.1016/j.ecoleng.2013.03.001.
Whitcraft, C.R., B.J. Grewell, and P.R. Baye. 2011. Estuarine Vegetation at Rush Ranch Open Space Preserve, San Franciso Bay National Estuarine Research Reserve, California. San Francisco Estuary and Watershed Science 9. https://doi.org/10.15447/sfews.2011v9iss3art6.
Whitfield, A.K. 2017. The role of seagrass meadows, mangrove forests, salt marshes and reed beds as nursery areas and food sources for fishes in estuaries. Reviews in Fish Biology and Fisheries 27 (1): 75–110. https://doi.org/10.1007/s11160-016-9454-x.
Whitley, S. N., and S. M. Bollens. 2014. Fish assemblages across a vegetation gradient in a restoring tidal freshwater wetland: diets and potential for resource competition. Environmental Biology of Fishes 97: 659–674. https://doi.org/10.1007/s10641-013-0168-9.
Wood, S.N. 2017. Generalized Additive Models: An Introduction with R. Second Edition: CRC Press.
Wood, S.N., M.V. Bravington, and S.L. Hedley. 2008. Soap film smoothing. Journal of the Royal Statistical Society: Series B (Statistical Methodology) 70 (5): 931–955. https://doi.org/10.1111/j.1467-9868.2008.00665.x.
Young, M., E. Howe, T. O’Rear, K. Berridge, and P. Moyle. 2020. Food Web Fuel Differs Across Habitats and Seasons of a Tidal Freshwater Estuary. Estuaries and Coasts. 44 (1): 286–301. https://doi.org/10.1007/s12237-020-00762-9.
Ziegler, S.L., R. Baker, S.C. Crosby, M.A. Barbeau, J. Cebrian, D. Mallick, C.W. Martin, J.A. Nelson, et al. 2021. Geographic variation in salt marsh structure and function for nekton: a guide to finding commonality across multiple scales. Estuaries and Coasts. https://doi.org/10.1007/s12237-020-00894-y.
Ziegler, S.L., K.W. Able, and F.J. Fodrie. 2019. Dietary shifts across biogeographic scales alter spatial subsidy dynamics. Ecosphere 10 (12): e02980. https://doi.org/10.1002/ecs2.2980.
Zimmerman, R.J., T.J. Minello, and L.P. Rozas. 2000. Salt Marsh Linkages to Productivity of Penaeid Shrimps and Blue Crabs in the Northern Gulf of Mexico. In Concepts and Controversies in Tidal Marsh Ecology, ed. M.P. Weinstein and D.A. Kreeger, 293–314. Dordrecht: Springer Netherlands. https://doi.org/10.1007/0-306-47534-0_14.
Acknowledgments
We thank R. Baker, M. Ferner, S. Ziegler, and anonymous reviewers for helpful comments on the manuscript.
Funding
Funding for data collection was provided by the California Department of Fish and Wildlife Ecosystem Restoration Program [E1183013], the California Department of Water Resources [4600011551], the California Department of Fish and Wildlife Proposition 1 [P1696010], and the S.D. Bechtel Jr. Foundation. Additional funding for D. Colombano was provided by the Delta Science Fellowship administered by the Delta Stewardship Council and California Sea Grant.
Author information
Authors and Affiliations
Contributions
B. Williamson, E. Davidson, E. Loughridge, and N. Ekasumara assisted in data collection; J. Donovan and D. Ayers provided TidalTrend support; and A. Manfree created the site map.
Corresponding author
Ethics declarations
Ethics
This study adhered to Animal Care and Use standards [18883].
Additional information
Communicated by Henrique Cabral
Supplementary Information
ESM 1
(DOCX 939 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Colombano, D.D., Handley, T.B., O’Rear, T.A. et al. Complex Tidal Marsh Dynamics Structure Fish Foraging Patterns in the San Francisco Estuary. Estuaries and Coasts 44, 1604–1618 (2021). https://doi.org/10.1007/s12237-021-00896-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12237-021-00896-4