Abstract
This study sought to reveal the proteomic profiling of methicillin-resistant Staphylococcus aureus (MRSA)-derived extracellular vesicles (EVs) after exposure to imipenem. The advanced isobaric tags for relative and absolute quantitation (iTRAQ®) proteomic approach were used to analyze the alterations in MRSA-derived EV protein patterns upon exposure to imipenem. A total of 1260 EV proteins were identified and quantified. Among these, 861 differentially expressed exosome proteins (P < 0.05) were found. Multivariate analysis, Gene Ontology (GO) annotation, and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis were used to analyze the identified proteins. Enrichment analysis of GO annotations indicated that imipenem primarily regulated the metabolic processes in MRSA. The metabolism of differentially expressed proteins was found to be the most significant in the combined analysis of the KEGG pathway analysis. Based on the results from the STRING analysis, 50S ribosomal protein L16 (RplP) and 30S ribosomal protein S8 (RpsH) were involved in the imipenem-induced MRSA-derived EVs. These results provide vital information on MRSA-derived EVs, increasing our knowledge of the proteome level changes in EVs upon exposure to imipenem. Moreover, these results pave the way for developing novel MRSA treatments.






Similar content being viewed by others
References
Admyre C, Grunewald J, Thyberg J, Gripenbäck S, Tornling G, Eklund A, Scheynius A, Gabrielsson S (2003) Exosomes with major histocompatibility complex class II and co-stimulatory molecules are present in human BAL fluid. Eur Respir J 22:578–583. https://doi.org/10.1183/09031936.03.00041703
Alves NJ, Turner KB, Daniele MA, Oh E, Medintz IL, Walper SA (2015) Bacterial nanobioreactors-directing enzyme packaging into bacterial outer membrane vesicles. ACS Appl Mater Interfaces 7:24963–24972. https://doi.org/10.1021/acsami.5b08811
Armogida SA, Yannaras NM, Melton AL, Srivastava MD (2004) Identification and quantification of innate immune system mediators in human breast milk. Allergy Asthma Proc 25:297–304
Beresin GA, Wright JM, Rice GE, Jagai JS (2017) Swine exposure and methicillin-resistant Staphylococcus aureus infection among hospitalized patients with skin and soft tissue infections in Illinois: a ZIP code-level analysis. Environ Res 159:46–60. https://doi.org/10.1016/j.envres.2017.07.037
Brown L, Wolf JM, Prados-Rosales R, Casadevall A (2015) Through the wall: extracellular vesicles in Gram-positive bacteria, mycobacteria and fungi. Nat Rev Microbiol 13:620–630. https://doi.org/10.1038/nrmicro3480
Chan DS et al (2007) Interaction between trichosanthin, a ribosome-inactivating protein, and the ribosomal stalk protein P2 by chemical shift perturbation and mutagenesis analyses. Nucleic Acids Res 35:1660–1672. https://doi.org/10.1093/nar/gkm065
Chiou JC, Li XP, Remacha M, Ballesta JP, Tumer NE (2008) The ribosomal stalk is required for ribosome binding, depurination of the rRNA and cytotoxicity of ricin A chain in Saccharomyces cerevisiae. Mol Microbiol 70:1441–1452. https://doi.org/10.1111/j.1365-2958.2008.06492.x
Clissold SP, Todd PA, Campoli-Richards DM (1987) Imipenem/cilastatin. A review of its antibacterial activity, pharmacokinetic properties and therapeutic efficacy. Drugs 33:183–241. https://doi.org/10.2165/00003495-198733030-00001
Cortes T, Cox RA (2015) Transcription and translation of the rpsJ, rplN and rRNA operons of the tubercle bacillus. Microbiology 161:719–728. https://doi.org/10.1099/mic.0.000037
Dorward DW, Garon CF (1990) DNA is packaged within membrane-derived vesicles of gram-negative but not gram-positive bacteria. Appl Environ Microbiol 56:1960–1962
Elkon KB, Parnassa AP, Foster CL (1985) Lupus autoantibodies target ribosomal P proteins. J Exp Med 162:459–471. https://doi.org/10.1084/jem.162.2.459
Gonin-Giraud S, Mathieu AL, Diocou S, Tomkowiak M, Delorme G, Marvel J (2002) Decreased glycolytic metabolism contributes to but is not the inducer of apoptosis following IL-3-starvation. Cell Death Differ 9:1147–1157. https://doi.org/10.1038/sj.cdd.4401079
Gui MJ, Dashper SG, Slakeski N, Chen YY, Reynolds EC (2016) Spheres of influence: Porphyromonas gingivalis outer membrane vesicles. Mol Oral Microbiol 31:365–378. https://doi.org/10.1111/omi.12134
Haddadin RN, Saleh S, Al-Adham IS, Buultjens TE, Collier PJ (2010) The effect of subminimal inhibitory concentrations of antibiotics on virulence factors expressed by Staphylococcus aureus biofilms. J Appl Microbiol 108:1281–1291. https://doi.org/10.1111/j.1365-2672.2009.04529.x
Hall BG (2004) Predicting the evolution of antibiotic resistance genes. Nat Rev Microbiol 2:430–435. https://doi.org/10.1038/nrmicro888
Hiramatsu K, Cui L, Kuroda M, Ito T (2001) The emergence and evolution of methicillin-resistant Staphylococcus aureus. Trends Microbiol 9:486–493. https://doi.org/10.1016/s0966-842x(01)02175-8
Hu FP, Guo Y, Zhu DM, Wang F, Jiang XF, Xu YC, Zhang XJ, Zhang CX, Ji P, Xie Y, Kang M, Wang CQ, Wang AM, Xu YH, Shen JL, Sun ZY, Chen ZJ, Ni YX, Sun JY, Chu YZ, Tian SF, Hu ZD, Li J, Yu YS, Lin J, Shan B, du Y, Han Y, Guo S, Wei LH, Wu L, Zhang H, Kong J, Hu YJ, Ai XM, Zhuo C, Su DH, Yang Q, Jia B, Huang W (2016) Resistance trends among clinical isolates in China reported from CHINET surveillance of bacterial resistance, 2005-2014. Clin Microbiol Infect 22(Suppl 1):S9–S14. https://doi.org/10.1016/j.cmi.2016.01.001
Im H, Lee S, Soper SA, Mitchell RJ (2017) Staphylococcus aureus extracellular vesicles (EVs): surface-binding antagonists of biofilm formation. Mol BioSyst 13:2704–2714. https://doi.org/10.1039/c7mb00365j
James EL, Michalek RD, Pitiyage GN, de Castro AM, Vignola KS, Jones J, Mohney RP, Karoly ED, Prime SS, Parkinson EK (2015) Senescent human fibroblasts show increased glycolysis and redox homeostasis with extracellular metabolomes that overlap with those of irreparable DNA damage, aging, and disease. J Proteome Res 14:1854–1871. https://doi.org/10.1021/pr501221g
Ji X, Liu X, Peng Y, Zhan R, Xu H, Ge X (2017) Comparative analysis of methicillin-sensitive and resistant Staphylococcus aureus exposed to emodin based on proteomic profiling. Biochem Biophys Res Commun 494:318–324. https://doi.org/10.1016/j.bbrc.2017.10.033
Jones M, Nielson C, Gupta K, Khader K, Evans M (2015) Collateral benefit of screening patients for methicillin-resistant Staphylococcus aureus at hospital admission: isolation of patients with multidrug-resistant gram-negative bacteria. Am J Infect Control 43:31–34. https://doi.org/10.1016/j.ajic.2014.09.016
Kemmer D, McHardy LM, Hoon S, Rebérioux D, Giaever G, Nislow C, Roskelley CD, Roberge M (2009) Combining chemical genomics screens in yeast to reveal spectrum of effects of chemical inhibition of sphingolipid biosynthesis. BMC Microbiol 9:9. https://doi.org/10.1186/1471-2180-9-9
Klein EY, Mojica N, Jiang W, Cosgrove SE, Septimus E, Morgan DJ, Laxminarayan R (2017) Trends in methicillin-resistant Staphylococcus aureus hospitalizations in the United States, 2010-2014. Clin Infect Dis 65:1921–1923. https://doi.org/10.1093/cid/cix640
Kuroda M, Ohta T, Hayashi H (1995) Isolation and the gene cloning of an alkaline shock protein in methicillin resistant Staphylococcus aureus. Biochem Biophys Res Commun 207:978–984. https://doi.org/10.1006/bbrc.1995.1281
Li SM, Zhou YF, Li L, Fang LX, Duan JH, Liu FR, Liang HQ, Wu YT, Gu WQ, Liao XP, Sun J, Xiong YQ, Liu YH (2018) Characterization of the multi-drug resistance gene cfr in methicillin-resistant Staphylococcus aureus (MRSA) strains isolated from animals and humans in China. Front Microbiol 9:2925. https://doi.org/10.3389/fmicb.2018.02925
Liao EC, Hsu YT, Chuah QY, Lee YJ, Hu JY, Huang TC, Yang PM, Chiu SJ (2014) Radiation induces senescence and a bystander effect through metabolic alterations. Cell Death Dis 5:e1255. https://doi.org/10.1038/cddis.2014.220
McCallum N, Spehar G, Bischoff M, Berger-Bachi B (2006) Strain dependence of the cell wall-damage induced stimulon in Staphylococcus aureus. Biochim Biophys Acta 1760:1475–1481. https://doi.org/10.1016/j.bbagen.2006.06.008
McCluskey AJ, Poon GM, Bolewska-Pedyczak E, Srikumar T, Jeram SM, Raught B, Gariepy J (2008) The catalytic subunit of Shiga-like toxin 1 interacts with ribosomal stalk proteins and is inhibited by their conserved C-terminal domain. J Mol Biol 378:375–386. https://doi.org/10.1016/j.jmb.2008.02.014
McDougal LK, Thornsberry C (1986) The role of beta-lactamase in staphylococcal resistance to penicillinase-resistant penicillins and cephalosporins. J Clin Microbiol 23:832–839
McNicholas PM, Mann PA, Najarian DJ, Miesel L, Hare RS, Black TA (2001) Effects of mutations in ribosomal protein L16 on susceptibility and accumulation of evernimicin. Antimicrob Agents Chemother 45:79–83. https://doi.org/10.1128/AAC.45.1.79-83.2001
Monnappa AK, Dwidar M, Seo JK, Hur JH, Mitchell RJ (2014) Bdellovibrio bacteriovorus inhibits Staphylococcus aureus biofilm formation and invasion into human epithelial cells. Sci Rep 4:3811. https://doi.org/10.1038/srep03811
Muller M et al (2014) Deletion of membrane-associated Asp23 leads to upregulation of cell wall stress genes in Staphylococcus aureus. Mol Microbiol 93:1259–1268. https://doi.org/10.1111/mmi.12733
Niccoli AA, Artesi AL, Candio F, Ceccarelli S, Cozzali R, Ferraro L, Fiumana D, Mencacci M, Morlupo M, Pazzelli P, Rossi L, Toscano M, Drago L (2014) Preliminary results on clinical effects of probiotic Lactobacillus salivarius LS01 in children affected by atopic dermatitis. J Clin Gastroenterol 48(Suppl 1):S34–S36. https://doi.org/10.1097/mcg.0000000000000233
Perez-Cruz C, Carrion O, Delgado L, Martinez G, Lopez-Iglesias C, Mercade E (2013) New type of outer membrane vesicle produced by the Gram-negative bacterium Shewanella vesiculosa M7T: implications for DNA content. Appl Environ Microbiol 79:1874–1881. https://doi.org/10.1128/aem.03657-12
Perez-Cruz C, Delgado L, Lopez-Iglesias C, Mercade E (2015) Outer-inner membrane vesicles naturally secreted by gram-negative pathogenic bacteria. PLoS One 10:e0116896. https://doi.org/10.1371/journal.pone.0116896
Pradelli LA, Villa E, Zunino B, Marchetti S, Ricci JE (2014) Glucose metabolism is inhibited by caspases upon the induction of apoptosis. Cell Death Dis 5:e1406. https://doi.org/10.1038/cddis.2014.371
Raimondo F, Morosi L, Chinello C, Magni F, Pitto M (2011) Advances in membranous vesicle and exosome proteomics improving biological understanding and biomarker discovery. Proteomics 11:709–720. https://doi.org/10.1002/pmic.201000422
Schooling SR, Hubley A, Beveridge TJ (2009) Interactions of DNA with biofilm-derived membrane vesicles. J Bacteriol 191:4097–4102. https://doi.org/10.1128/jb.00717-08
Sirichoat A, Lulitanond A, Kanlaya R, Tavichakorntrakool R, Chanawong A, Wongthong S, Thongboonkerd V (2016) Phenotypic characteristics and comparative proteomics of Staphylococcus aureus strains with different vancomycin-resistance levels. Diagn Microbiol Infect Dis 86:340–344. https://doi.org/10.1016/j.diagmicrobio.2016.09.011
Soleymanzadeh-Moghadam S, Azimi L, Amani L, Rastegar Lari A, Alinejad F, Rastegar Lari A (2015) Analysis of antibiotic consumption in burn patients. GMS Hyg Infect Control 10:Doc09. https://doi.org/10.3205/dgkh000252
Tran F, Boedicker JQ (2017) Genetic cargo and bacterial species set the rate of vesicle-mediated horizontal gene transfer. Sci Rep 7:8813. https://doi.org/10.1038/s41598-017-07447-7
Wang J, Duan M, Erdemutu PS, Wei C, Han Y, Hospital A (2017) Effects of sub-inhibitory concentration of imipenem on proliferation in vitro and mRNA expression levels of MRSA virulence related genes. J Jilin Univ 43:479–484
Wool IG, Chan YL, Gluck A (1995) Structure and evolution of mammalian ribosomal proteins. Biochem Cell Biol 73:933–947. https://doi.org/10.1139/o95-101
Yang Y, Zheng N, Zhao X, Zhang Y, Han R, Ma L, Zhao S, Li S, Guo T, Wang J (2015) Proteomic characterization and comparison of mammalian milk fat globule proteomes by iTRAQ analysis. J Proteome 116:34–43. https://doi.org/10.1016/j.jprot.2014.12.017
Yang M, Song D, Cao X, Wu R, Liu B, Ye W, Wu J, Yue X (2017) Comparative proteomic analysis of milk-derived exosomes in human and bovine colostrum and mature milk samples by iTRAQ-coupled LC-MS/MS. Food Res Int 92:17–25. https://doi.org/10.1016/j.foodres.2016.11.041
Zakharzhevskaya NB, Tsvetkov VB, Vanyushkina AA, Varizhuk AM, Rakitina DV, Podgorsky VV, Vishnyakov IE, Kharlampieva DD, Manuvera VA, Lisitsyn FV, Gushina EA, Lazarev VN, Govorun VM (2017) Interaction of Bacteroides fragilis toxin with outer membrane vesicles reveals new mechanism of its secretion and delivery. Front Cell Infect Microbiol 7:2. https://doi.org/10.3389/fcimb.2017.00002
Funding
The study was funded by the National Natural Science Foundation of China (grant number 81660352).
Author information
Authors and Affiliations
Contributions
In vitro proliferation assay: WJC; data analysis and manuscript writing: WJC and LEM; iTRAQ and MRM data analysis: WJR and RL; protein extraction: WYY and ZCX; bioinformatics analysis: SP; transmission electron microscopy (TEM) and scanning electron microscopy (SEM) analysis: ZXH.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research involving human participants and/or animals
Not applicable.
Informed consent
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Online Resource 1
Global proteomic data in MRSA-derived EVs (XLSX 125 kb)
Online Resource 2
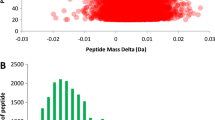
Differentially EVs expressed proteins, volcano plot (A) and histograms (B), between imipenem treated S23 and S23 only (XLSX 163 kb)
Online Resource 3
Gene ontology analysis, including cellular component (A), biological process (B), and molecular function (C), in EVs between imipenem treated S23 and S23 only (XLSX 1442 kb)
Online Resource 4
The KEGG pathway analysis in EVs between imipenem treated S23 and S23 only (XLSX 22 kb)
Online Resource 5
MRM analysis in EVs between imipenem treated S23 and S23 only (XLSX 15 kb)
Online Resource 6
Correlation between MRM and iTRAQ® analysis in EVs between imipenem treated S23 and S23 only (XLSX 17 kb)
Rights and permissions
About this article
Cite this article
Wang, J., Wang, J., Wang, Y. et al. iTRAQ®-based quantitative proteomics reveals the proteomic profiling of methicillin-resistant Staphylococcus aureus-derived extracellular vesicles after exposure to imipenem. Folia Microbiol 66, 221–230 (2021). https://doi.org/10.1007/s12223-020-00836-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-020-00836-y