Abstract
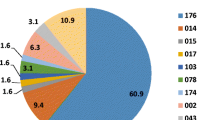
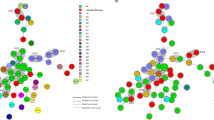
The purpose of this study is to analyze isolates of Clostridium difficile from patients with nosocomial acquired infection in respect to their molecular type and antimicrobial susceptibility. Fifty-nine randomly selected clinical isolates were characterized. Molecular typing was performed by rep-PCR (DiversiLab). Isolates were tested by disk diffusion towards 11 different antibiotics. All isolates were susceptible to metronidazole and vancomycin. Fifty five (93 %) isolates were resistant to erythromycin and fifty six (95 %) exhibited resistance to both clindamycin and moxifloxacin. Twenty rep-PCR types were identified, but most clinical isolates formed four major rep-PCR clusters (A1 24/59, 40 %; A2 20/59, 33 %; A3 5/59, 8 %; A4 3/59, 5 %). These results show high genetic variability, which demonstrate clearly the complexity of the strains of C. difficile and also show an increasing rate of resistance to fluoroquinolones in our region emphasizing the importance of implementing surveillance programs in order to prevent further spread of resistance in C. difficile.


Similar content being viewed by others
References
Ackermann G, Thomalla S, Ackermann F, Schaumann R, Rodloff AC, Ruf BR (2005) Prevalence and characteristics of bacteria and host factors in an outbreak situation of antibiotic-associated diarrhoea. J Med Microbiol 54:149–153
Adriaenssen N, Coenen S, Versporten A, Muller A, Minalu G, Faes C et al (2011) European Surveillance of Antimicrobial Consumption (ESAC): outpatient quinolone use in Europe (1997–2009). J Antimicrob Chemother 66:47–56
Akerlund T, Svenungsson B, Lagergren A, Burman LG (2006) Correlation of disease severity with fecal toxin levels in patients with Clostridium difficile-associated diarrhea and distribution of PCVR ribotypes and toxin yields in vitro of corresponding isolates. J Clin Microbiol 44:353–358
Alfa MJ, Kabani A, Lyerly D, Moncrief S, Neville LM, Al-Barrak A et al (2000) Characterization of a toxin A-negative, toxin B-positive strain of Clostridium difficile responsible for a nosocomial outbreak of Clostridium difficile-associated diarrhea. J Clin Microbiol 38:2706–2714
Aspevalla O, Lundberg A, Burman LG, Akerlund T, Svenungsson B (2006) Antimicrobial susceptibility pattern of Clostridium difficile and its relation to PCR ribotypes in a Swedish University hospital. Antimicrob Agents Chemother 50:1890–1892
Barbut F, Mastrantonio P, Delmee M, Brazier J, Kuijper E, Poxton I (2007) Prospective study of Clostridium difficile infections in Europe with phenotypic and genotypic characterisation of the isolates. European Study Group on Clostridium difficile. Clin Microbiol Infect 13:1048–1057
Bauer MP, Notermans DW, van Benthem BH, Brazier JS, Wilcox MH, Rupnik M et al (2011) Clostridium difficile infection in Europe: a hospital-based survey. Lancet 377:63–73
Blossom DB, McDonald LC (2007) The challenges posed by reemerging Clostridium difficile infection. Clin Infect Dis 45:222–227
Brazier JS, Fawley W, Freeman J, Wilcox MH (2001) Reduced susceptibility of Clostridium difficile to metronidazole. J Antimicrob Chemother 48:741–742
Climo MW, Israel DS, Wong ES, Williams D, Coudron P, Markowitz SM (1998) Hospital-wide restriction of clindamycin: effect on the incidence of Clostridium difficile-associated diarrhea and cost. Ann Intern Med 128:989–995
Di Bella S, Musso M, Cataldo MA, Meleandri M, Bordi E, Capozzi D et al. (2013) Clostridium difficile infection in Italian urban hospitals. Data from 2006 through 2011. BMC Infect Dis, in press. doi:10.1186/1471-2334-13-146
Erikstrup LT, Danielsen TKL, Hall V, Olsen KEP, Kristensen B, Kahlmeter G et al (2012) Antimicrobial susceptibility testing of Clostridium difficile using EUCAST epidemiological cut-off values and disk diffusion correlates. Clin Microbiol Infect 18:266–272
Freeman J, Bauer MP, Baines SD, Corver J, Fawley WN, Goorhuis B et al (2010) The changing epidemiology of Clostridium difficile infections. Clin Microbiol Rev 23:259–549
Kelly CP, LaMont JT (2008) Clostridium difficile—more difficult than ever. N Engl J Med 359:1932–1940
Killgore G, Thompson A, Johnson S, Brazier J, Kuijper E, Pepin J et al (2008) Comparison of seven techniques for typing international epidemic strains of Clostridium difficile: restriction endonuclease analysis, pulsed field gel electrophoresis, PCR-ribotyping, multilocus variable number tandem-repeat analysis, amplified fragment length polymorphism, and surface layer protein a gene sequence typing. J Clin Microbiol 46:431–437
Kim J, Kang JO, Kim H, Seo MR, Choi TY, Pai H et al (2012) Epidemiology of Clostridium difficile infections in a tertiary-care hospital in Korea. Clin Microbiol Infect. doi:10.1111/j.1469-0691.2012.03910.x
Klaassen CH, van Haren HA, Horrevorts AM (2002) Molecular fingerprinting of Clostridium difficile isolates: pulsed-field electrophoresis versus amplified fragment length polymorphism. J Clin Microbiol 40:101–104
Lin YC, Huang YT, Tsai PJ, Lee TF, Lee NY, Liao CH et al (2011) Antimicrobial susceptibilities and molecular epidemiology of Clostridium difficile in Taiwan. Antimicrob Agents Chemother 55:1701–1705
McDonald LC, Killgore GE, Thompson A, Owens RC Jr, Kazakova SV, Sambol SP et al (2005) An epidemic, toxin gene-variant strain of Clostridium difficile. N Engl J Med 353:2433–2441
McDonald LC, Owings M, Jernigan DB (2006) Clostridium difficile infection among patients discharged from US short-stay hospitals, 1996–2003. Emerg Infect Dis 12:409–415
Merrigan M, Venugopal A, Mallozzi M, Roxas B, Viswanathan VH, Johnson S et al (2010) Human hypervirulent Clostridium difficile strains exhibit increased sporulation as well as robust toxin production. J Bacteriol 192:4904–4911
O’Connor JR, Johnson S, Gerding DN (2009) Clostridium difficile infection caused by the epidemic BI/NAP1/027 strain. Gastroenterology 136:1913–1924
Owens RC Jr, Donskey CJ, Gaynes RP, Loo VG, Muto CA (2008) Antimicrobial associated risk factors for Clostridium difficile infection. Clin Infect Dis 46(Suppl 1):S19–S31
Pasanen T, Kotila SM, Horsma J, Virolainen A, Jalava J, Ibrahem S et al (2011) Comparison of repetitive extragenic palindromic sequence-based PCR with PCR ribotyping and pulsed field gel electrophoresis in studying the clonality of Clostridium difficile. Clin Microbiol Infect 7:166–175
Pear SM, Williamson TH, Bettin KM, Gerding DN, Galgiani JN (1994) Decrease in nosocomial Clostridium difficile-associated diarrhea by restricting clindamycin use. Ann Intern Med 120:272–277
Pelaéz T, Alcala L, Alonso R, Martin Lopez A, Garcia-Arias V, Marin M et al (2005) In vitro activity of ramoplanin against Clostridium difficile, including strains with reduced susceptibility to vancomycin or with resistance to metronidazole. Antimicrob Agents Chemother 49:1157–1159
Pepin J, Valiquette L, Alary ME (2004) Clostridium difficile-associated diarrhea in a region of Quebec from 1991 to 2003: a changing pattern of disease severity. CMAJ 171:466–472
Poilane I, Bert F, Cruad P, Nicolas-Chanoine MH, Collignon A (2007) Interest of the disk diffusion method for screening of Clostridium difficile isolates with decreased susceptibility to antibiotics. Pathol Biol 55:429–433
Poutanen SM, Simor AE (2004) Clostridium difficile-associated diarrhea in adults. CMAJ 171:51–58
Redelings MD, Sorvillo F, Mascola L (2007) Increase in Clostridium difficile-related mortality rates, Unites States, 1999–2004. Emerg Infect Dis 13:1417–1419
Rousseau C, Lemée L, Le Monnier A, Poilane I, Pons JL, Collignon A (2011) Prevalence and diversity of Clostridium difficile strains in infants. J Med Microbiol 60:1112–1118
Russello G, Russo A, Sisto F, Scaltrito MM, Farina C (2012) Laboratory diagnosis of Clostridium difficile associated diarrhoea and molecular characterization of clinical isolates. New Microbiol 35:307–316
Sansone S, Aschbacher R, Staffler, Bombonato M, Girardi F, Larcher C et al (2009) Nosocomial diarrhea in adult medical patients: the role of Clostridium difficile in a North Italian acute care teaching hospital. J Prev Med Hyg 50:117–120
Spigaglia P, Barbanti F, Dionisi AM, Mastrantonio P (2010) Clostridium difficile isolates resistant to fluoroquinolones in Italy: emergence of PCR ribotype 018. J Clin Microbiol 48:2892–2896
Stabler RA, Dawson LF, Phua LT, Wren BW (2008) Comparative analysis of BI/NAP1/027 hypervirulent strains reveals novel toxin B-encoding gene (tcdB) sequences. J Med Microbiol 57:771–775
Acknowledgments
We would like to thank Surace Antonella for revising the text.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Silvia Corbellini and Giorgio Piccinelli contributed equally to this work.
Rights and permissions
About this article
Cite this article
Corbellini, S., Piccinelli, G., De Francesco, M.A. et al. Molecular epidemiology of Clostridium difficile strains from nosocomial-acquired infections. Folia Microbiol 59, 173–179 (2014). https://doi.org/10.1007/s12223-013-0281-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-013-0281-3