Abstract
Introduction
The propagation of mechanochemical signals from the extracellular matrix to the cell nucleus has emerged as a central feature in regulating cellular differentiation and de-differentiation. This process of outside-in signaling and the associated mechanotransduction pathways have been well described in numerous developmental and pathological contexts. However, it remains less clear how mechanotransduction influences the activity of chromatin modifying enzymes that direct gene expression programs.
Objectives
The primary objective of this study was to explore how matrix mechanics and geometric confinement influence histone deacetylase (HDAC) activity in fibroblast culture.
Methods
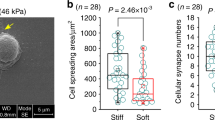
Polyacrylamide hydrogels were formed and functionalized with fibronectin patterns using soft lithography. Primary mouse embryonic fibroblasts (MEFs) were cultured on the islands until confluent, fixed, and immunolabeled for microscopy.
Results
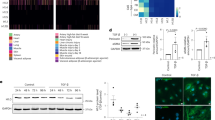
After 24 h MEFs cultured on soft hydrogels (0.5 kPa) show increased expression of class I HDACs relative to MEFs cultured on stiff hydrogels (100 kPa). A member of the class II family, HDAC4 shows a similar trend; however, there is a pronounced cytoplasmic localization on soft hydrogels suggesting a role in outside-in cytoplasmic signaling. Pharmacological inhibitor studies suggest that the opposing activities of extracellular related kinase 1/2 (ERK) and protein phosphatase 2a (PP2a) influence the localization of HDAC4. MEFs cultured on the soft hydrogels show enhanced expression of markers associated with a fibroblast–myofibroblast transition (Paxillin, αSMA).
Conclusions
We show that the phosphorylation state and cellular localization of HDAC4 is influenced by matrix mechanics, with evidence for a role in mechanotransduction and the regulation of gene expression associated with fibroblast–myofibroblast transitions. This work establishes a link between outside-in signaling and epigenetic regulation, which will assist efforts aimed at controlling gene regulation in engineered extracellular matrices.




Similar content being viewed by others
References
Abdeen, A. A., J. Lee, N. A. Bharadwaj, R. H. Ewoldt, and K. A. Kilian. Temporal modulation of stem cell activity using magnetoactive hydrogels. Adv. Healthc. Mater. 5(19):2536–2544, 2016.
Abdeen, A. A., J. Lee, Y. Li, and K. A. Kilian. Cytoskeletal priming of mesenchymal stem cells to a medicinal phenotype. Regen. Eng. Transl. Med. 3:5–14, 2017.
Abdeen, A. A., J. Lee, S. H. Mo, and K. A. Kilian. Spatially defined stem cell-laden hydrogel islands for directing endothelial tubulogenesis. J. Mater. Chem. B 3(40):7896–7898, 2015.
Abdeen, A. A., J. B. Weiss, J. Lee, and K. A. Kilian. Matrix composition and mechanics direct proangiogenic signaling from mesenchymal stem cells. Tissue Eng. Part A 20(19–20):2737–2745, 2014.
Alam, S. G., Q. Zhang, N. Prasad, Y. Li, S. Chamala, R. Kuchibhotla, B. Kc, V. Aggarwal, S. Shrestha, A. L. Jones, S. E. Levy, K. J. Roux, J. A. Nickerson, and T. P. Lele. The mammalian LINC complex regulates genome transcriptional responses to substrate rigidity. Sci. Rep. 6:38063, 2016.
Backs, J., K. Song, S. Bezprozvannaya, S. Chang, and E. N. Olson. CaM kinase II selectively signals to histone deacetylase 4 during cardiomyocyte hypertrophy. J. Clin. Invest. 116(7):1853–1864, 2006.
Banerjee, S., and M. C. Marchetti. Controlling cell-matrix traction forces by extracellular geometry. New J. Phys. 15(3):35015, 2013.
Baum, J., and H. S. Duffy. Fibroblasts and myofibroblasts: what are we talking about? J. Cardiovasc. Pharmacol. 57(4):376–379, 2011.
Cernotta, N., A. Clocchiatti, C. Florean, and C. Brancolini. Ubiquitin-dependent degradation of HDAC4, a new regulator of random cell motility. Mol. Biol. Cell 22(2):278–289, 2011.
Chambliss, A. B., P.-H. Wu, W.-C. Chen, S. X. Sun, and D. Wirtz. Simultaneously defining cell phenotypes, cell cycle, and chromatin modifications at single-cell resolution. FASEB J. 27(7):2667–2676, 2013.
Chancellor, T. J., J. Lee, C. K. Thodeti, and T. Lele. Actomyosin tension exerted on the nucleus through nesprin-1 connections influences endothelial cell adhesion, migration, and cyclic strain-induced reorientation. Biophys. J. 99(1):115–123, 2010.
Dahl, K. N., A. J. S. Ribeiro, and J. Lammerding. Nuclear shape, mechanics, and mechanotransduction. Circ. Res. 102(11):1307–1318, 2008.
Discher, D. E., D. J. Mooney, and P. W. Zandstra. Growth factors, matrices, and forces combine and control stem cells. Science 324(5935):1673–1677, 2009.
Dupont, S., L. Morsut, M. Aragona, E. Enzo, S. Giulitti, M. Cordenonsi, F. Zanconato, J. Le Digabel, M. Forcato, S. Bicciato, N. Elvassore, and S. Piccolo. Role of YAP/TAZ in mechanotransduction. Nature 474(7350):179–183, 2011.
Evans, N. D., C. Minelli, E. Gentleman, V. LaPointe, S. N. Patankar, M. Kallivretaki, X. Chen, C. J. Roberts, and M. M. Stevens. Substrate stiffness affects early differentiation events in embryonic stem cells. Eur. Cells Mater. 18:1–13, 2009.
Farge, E. Mechanical induction of Twist in the Drosophila foregut/stomodeal primordium. Curr. Biol. 13(16):1365–1377, 2003.
Glenisson, W., V. Castronovo, and D. Waltregny. Histone deacetylase 4 is required for TGF beta 1-induced myofibroblastic differentiation. Biochim. Biophys. Acta 1773(10):1572–1582, 2007.
Grozinger, C. M., and S. L. Schreiber. Regulation of histone deacetylase 4 and 5 and transcriptional activity by 14-3- 3-dependent cellular localization. Proc. Natl. Acad. Sci. USA. 97(14):7835–7840, 2000.
Guo, W., B. Shan, R. C. Klingsberg, X. Qin, and J. A. Lasky. Abrogation of TGF-beta1-induced fibroblast-myofibroblast differentiation by histone deacetylase inhibition. Am. J. Physiol. Lung Cell. Mol. Physiol. 297(5):L864–L870, 2009.
Haberland, M., R. L. Montgomery, and E. N. Olson. The many roles of histone deacetylases in development and physiology: implications for disease and therapy. Nat. Rev. Genet. 10(1):32–42, 2009.
Hebbes, T. R., A. W. Thorne, and C. Crane-Robinson. A direct link between core histone acetylation and transcriptionally active chromatin. EMBO J. 7(5):1395–1402, 1988.
Heo, S.-J., T. P. Driscoll, S. D. Thorpe, N. L. Nerurkar, B. M. Baker, M. T. Yang, C. S. Chen, D. A. Lee, and R. L. Mauck. Differentiation alters stem cell nuclear architecture, mechanics, and mechano-sensitivity. Elife, 5:e18207, 2016
Heo, S.-J., S. D. Thorpe, T. P. Driscoll, R. L. Duncan, D. A. Lee, and R. L. Mauck. Biophysical regulation of chromatin architecture instills a mechanical memory in mesenchymal stem cells. Sci. Rep. 5(1):16895, 2015.
Huang, X., N. Yang, V. F. Fiore, T. H. Barker, Y. Sun, S. W. Morris, Q. Ding, V. J. Thannickal, and Y. Zhou. Matrix stiffness-induced myofibroblast differentiation is mediated by intrinsic mechanotransduction. Am. J. Respir. Cell Mol. Biol. 47(3):340–348, 2012.
Ingber, D. E. Cellular mechanotransduction: putting all the pieces together again. FASEB J. 20(7):27, 2006.
Jagielska, A., A. L. Lowe, E. Makhija, L. Wroblewska, J. Guck, R. J. M. Franklin, G. V. Shivashankar, and K. J. Van Vliet. Mechanical strain promotes oligodendrocyte differentiation by global changes of gene expression. Front. Cell. Neurosci. 11:93, 2017.
Jain, N., K. V. Iyer, A. Kumar, and G. V. Shivashankar. Cell geometric constraints induce modular gene-expression patterns via redistribution of HDAC3 regulated by actomyosin contractility. Proc. Natl. Acad. Sci. USA 110(28):11349–11354, 2013.
Karamichos, D., R. A. Brown, and V. Mudera. Collagen stiffness regulates cellular contraction and matrix remodeling gene expression. J. Biomed. Mater. Res. A 83(3):887–894, 2007.
Kocgozlu, L., P. Lavalle, G. Koenig, B. Senger, Y. Haikel, P. Schaaf, J.-C. Voegel, H. Tenenbaum, and D. Vautier. Selective and uncoupled role of substrate elasticity in the regulation of replication and transcription in epithelial cells. J. Cell Sci. 123(Pt 1):29–39, 2010.
Kumar, A., and G. V. Shivashankar. Dynamic interaction between actin and nesprin2 maintain the cell nucleus in a prestressed state. Methods Appl. Fluoresc. 4(4):44008, 2016.
Lee, J., A. A. Abdeen, T. H. Huang, and K. A. Kilian. Controlling cell geometry on substrates of variable stiffness can tune the degree of osteogenesis in human mesenchymal stem cells. J. Mech. Behav. Biomed. Mater. 38:209–218, 2014.
Lee, J., A. A. Abdeen, A. S. Kim, and K. A. Kilian. Influence of biophysical parameters on maintaining the mesenchymal stem cell phenotype. ACS Biomater. Sci. Eng. 1(4):218–226, 2015.
Lee, J., A. A. Abdeen, K. L. Wycislo, T. M. Fan, and K. A. Kilian. Interfacial geometry dictates cancer cell tumorigenicity. Nat Mater. 15(8):856–862, 2016.
Lee, C., A. Grodzinsky, and M. Spector. The effects of cross-linking of collagen-glycosaminoglycan scaffolds on compressive stiffness, chondrocyte-mediated contraction, proliferation and biosynthesis. Biomaterials 22(23):3145–3154, 2001.
Li, J., J. Chen, C. L. Ricupero, R. P. Hart, M. S. Schwartz, A. Kusnecov, and K. Herrup. Nuclear accumulation of HDAC4 in ATM deficiency promotes neurodegeneration in ataxia telangiectasia. Nat. Med. 18(5):783–790, 2012.
Li, Y., J. S. Chu, K. Kurpinski, X. Li, D. M. Bautista, L. Yang, K.-L. P. Sung, and S. Li. Biophysical regulation of histone acetylation in mesenchymal stem cells. Biophys. J. 100(8):1902–1909, 2011.
Li, Y., D. Lovett, Q. Zhang, S. Neelam, R. A. Kuchibhotla, R. Zhu, G. G. Gundersen, T. P. Lele, and R. B. Dickinson. Moving cell boundaries drive nuclear shaping during cell spreading. Biophys. J. 109(4):670–686, 2015.
Lovett, D. B., N. Shekhar, J. A. Nickerson, K. J. Roux, and T. P. Lele. Modulation of nuclear shape by substrate rigidity. Cell. Mol. Bioeng. 6(2):230–238, 2013.
Low, B. C., C. Q. Pan, G. V. Shivashankar, A. Bershadsky, M. Sudol, and M. Sheetz. YAP/TAZ as mechanosensors and mechanotransducers in regulating organ size and tumor growth. FEBS Lett. 588(16):2663–2670, 2014.
Maniotis, A. J., C. S. Chen, and D. E. Ingber. Demonstration of mechanical connections between integrins, cytoskeletal filaments, and nucleoplasm that stabilize nuclear structure. Proc. Natl. Acad. Sci. 94(3):849–854, 1997.
Mao, A. S., J.-W. Shin, and D. J. Mooney. Effects of substrate stiffness and cell-cell contact on mesenchymal stem cell differentiation. Biomaterials 98:184–191, 2016.
Mih, J. D., A. Marinkovic, F. Liu, A. S. Sharif, and D. J. Tschumperlin. Matrix stiffness reverses the effect of actomyosin tension on cell proliferation. J. Cell Sci. 125(Pt 24):5974–5983, 2012.
Park, J. S., J. S. Chu, A. D. Tsou, R. Diop, Z. Tang, A. Wang, and S. Li. The effect of matrix stiffness on the differentiation of mesenchymal stem cells in response to TGF-β. Biomaterials 32:3921–3930, 2011.
Paroni, G., N. Cernotta, C. Dello Russo, P. Gallinari, M. Pallaoro, C. Foti, F. Talamo, L. Orsatti, C. Steinkuhler, and C. Brancolini. PP2A regulates HDAC4 nuclear import. Mol. Biol. Cell 19(2):655–667, 2007.
Provenzano, P. P., D. R. Inman, K. W. Eliceiri, and P. J. Keely. Matrix density-induced mechanoregulation of breast cell phenotype, signaling and gene expression through a FAK-ERK linkage. Oncogene 28(49):4326–4343, 2009.
Provenzano, P. P., and P. J. Keely. Mechanical signaling through the cytoskeleton regulates cell proliferation by coordinated focal adhesion and Rho GTPase signaling. J. Cell Sci. 124(Pt 8):1195–1205, 2011.
Ralphs, J. R., A. D. Waggett, and M. Benjamin. Actin stress fibres and cell-cell adhesion molecules in tendons: organisation in vivo and response to mechanical loading of tendon cells in vitro. Matrix Biol. 21(1):67–74, 2002.
Ren, K., T. Crouzier, C. Roy, and C. Picart. Polyelectrolyte multilayer films of controlled stiffness modulate myoblast cells differentiation. Adv. Funct. Mater. 18(9):1378–1389, 2008.
Rozwadowska, N., T. Kolanowski, E. Wiland, M. Siatkowski, P. Pawlak, A. Malcher, T. Mietkiewski, M. Olszewska, and M. Kurpisz. Characterisation of nuclear architectural alterations during in vitro differentiation of human stem cells of myogenic origin. PLoS ONE 8(9):e73231, 2013.
Ruthenburg, A. J., H. Li, D. J. Patel, and C. D. Allis. Multivalent engagement of chromatin modifications by linked binding modules. Nat. Rev. Mol. Cell Biol. 8(12):983–994, 2007.
Santiago, J.-J., A. L. Dangerfield, S. G. Rattan, K. L. Bathe, R. H. Cunnington, J. E. Raizman, K. M. Bedosky, D. H. Freed, E. Kardami, and I. M. C. Dixon. Cardiac fibroblast to myofibroblast differentiation in vivo and in vitro: expression of focal adhesion components in neonatal and adult rat ventricular myofibroblasts. Dev. Dyn. 239(6):1573–1584, 2010.
Serebryannyy, L. A., C. M. Cruz, P. de Lanerolle, R. D. Mullins, and P. D. McCrea. A role for nuclear actin in HDAC 1 and 2 regulation. Sci. Rep. 6(1):28460, 2016.
Shih, Y.-R. V., K.-F. Tseng, H.-Y. Lai, C.-H. Lin, and O. K. Lee. Matrix stiffness regulation of integrin-mediated mechanotransduction during osteogenic differentiation of human mesenchymal stem cells. J. Bone Miner. Res. 26(4):730–738, 2011.
Swift, J., I. L. Ivanovska, A. Buxboim, T. Harada, P. C. D. P. Dingal, J. Pinter, J. D. Pajerowski, K. R. Spinler, J.-W. Shin, M. Tewari, F. Rehfeldt, D. W. Speicher, and D. E. Discher. Nuclear lamin-A scales with tissue stiffness and enhances matrix-directed differentiation. Science 341(6149):1240104, 2013.
Tan, P. S., and S. H. Teoh. Effect of stiffness of polycaprolactone (PCL) membrane on cell proliferation. Mater. Sci. Eng. C 27(2):304–308, 2007.
Tse, J. R., and A. J. Engler. Preparation of hydrogel substrates with tunable mechanical properties. Curr. Protoc. Cell Biol. 47:10.16:10.16.1–10.16.16, 2010
Vega, R. B., K. Matsuda, J. Oh, A. C. Barbosa, X. Yang, E. Meadows, J. McAnally, C. Pomajzl, J. M. Shelton, J. A. Richardson, G. Karsenty, and E. N. Olson. Histone deacetylase 4 controls chondrocyte hypertrophy during skeletogenesis. Cell 119(4):555–566, 2004.
Veith, C., L. M. Marsh, M. Wygrecka, K. Rutschmann, W. Seeger, N. Weissmann, and G. Kwapiszewska. Paxillin regulates pulmonary arterial smooth muscle cell function in pulmonary hypertension. Am. J. Pathol. 181(5):1621–1633, 2012.
Wang, L.-S., J. Boulaire, P. P. Y. Chan, J. E. Chung, and M. Kurisawa. The role of stiffness of gelatin-hydroxyphenylpropionic acid hydrogels formed by enzyme-mediated crosslinking on the differentiation of human mesenchymal stem cell. Biomaterials 31(33):8608–8616, 2010.
Wang, N., J. P. Butler, and D. E. Ingber. Mechanotransduction across the cell surface and through the cytoskeleton. Science 260(5111):1124–1127, 1993.
Wang, A. H., M. J. Kruhlak, J. Wu, N. R. Bertos, M. Vezmar, B. I. Posner, D. P. Bazett-Jones, and X. J. Yang. Regulation of histone deacetylase 4 by binding of 14-3-3 proteins. Mol. Cell. Biol. 20(18):6904–6912, 2000.
Wang, A., S. K. Kurdistani, and M. Grunstein. Requirement of Hos2 histone deacetylase for gene activity in yeast. Science 298(5597):1412–1414, 2002.
Wang, Z., G. Qin, and T. C. Zhao. HDAC4: mechanism of regulation and biological functions. Epigenomics 6(1):139–150, 2014.
Wang, A. H., and X. J. Yang. Histone deacetylase 4 possesses intrinsic nuclear import and export signals. Mol. Cell. Biol. 21(17):5992–6005, 2001.
Yeung, T., P. C. Georges, L. A. Flanagan, B. Marg, M. Ortiz, M. Funaki, N. Zahir, W. Ming, V. Weaver, and P. A. Janmey. Effects of substrate stiffness on cell morphology, cytoskeletal structure, and adhesion. Cell Motil. Cytoskelet. 60(1):24–34, 2005.
Yuan, H., K. Denton, L. Liu, X. J. Li, S. Benashski, L. McCullough, and J. Li. Nuclear translocation of histone deacetylase 4 induces neuronal death in stroke. Neurobiol. Dis. 91:182–193, 2016.
Zhou, X., V. M. Richon, A. H. Wang, X.-J. Yang, R. A. Rifkind, and P. A. Marks. Histone deacetylase 4 associates with extracellular signal-regulated kinases 1 and 2, and its cellular localization is regulated by oncogenic Ras. Proc. Natl. Acad. Sci. 97(26):14329–14333, 2000.
Acknowledgments
The authors would like to thank Dr. Quanxi Li in the Department of Comparative Biosciences at the University of Illinois for the generous isolation and donation of MEFs and Dr. Karin Jensen in the Department of Bioengineering at the University of Illinois for guidance with western blots.
Conflict of interest
Yanfen Li, Claire B. Tang, and Kristopher A. Kilian declare that they have no conflicts of interest.
Funding
Funding was provided by the National Science Foundation Grant No. 1454616 CAR. Y.L. was supported by the National Science Foundation Graduate Research Fellowship Program under Grant No. DGE – 1144245.
Research Involved in Human and Animal Rights
No humans or animals were employed in this research by the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
Associate Editor Alyssa Panitch oversaw the review of this article.
Kristopher A. Kilian
received B.S. and M.S. degrees in Chemistry from the University of Washington in 1999 and 2003 respectively. He worked for Merck Research Labs in the Methods Development group from 2000 to 2004 before travelling to Sydney, Australia to do his PhD with Justin Gooding at the University of New South Wales. In 2007, he joined the laboratory of Milan Mrksich at the University of Chicago as a postdoctoral fellow to investigate new methods for directing the differentiation of stem cells. Kris joined the faculty of the University of Illinois at Urbana-Champaign in 2011 with appointments in the Departments of Materials Science and Engineering, and Bioengineering. Kris is a 2008 recipient of the NIH Ruth L. Kirchstein National Research Service Award, a 2014 Kavli Fellow of the 19th German-American Frontiers of Science, and a 2015 recipient of the National Science Foundation’s CAREER award. His research interests include the design and development of model extracellular matrices for cell engineering and fundamental studies in cell biology.

This article is part of the 2017 CMBE Young Innovators special issue.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, Y., Tang, C.B. & Kilian, K.A. Matrix Mechanics Influence Fibroblast–Myofibroblast Transition by Directing the Localization of Histone Deacetylase 4. Cel. Mol. Bioeng. 10, 405–415 (2017). https://doi.org/10.1007/s12195-017-0493-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12195-017-0493-8