Abstract
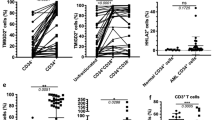
The NUP98::NSD1 fusion gene is associated with extremely poor prognosis in patients with acute myeloid leukemia (AML). NUP98::NSD1 induces self-renewal and blocks differentiation of hematopoietic stem cells, leading to development of leukemia. Despite its association with poor prognosis, targeted therapy for NUP98::NSD1-positive AML is lacking, as the details of NUP98::NSD1 function are unknown. Here, we generated 32D cells (a murine interleukin-3 (IL-3)-dependent myeloid progenitor cell line) expressing mouse Nup98::Nsd1 to explore the function of NUP98::NSD1 in AML, including comprehensive gene expression analysis. We identified two properties of Nup98::Nsd1 + 32D cells in vitro. First, Nup98::Nsd1 promoted blocking of AML cell differentiation, consistent with a previous report. Second, Nup98::Nsd1 increased dependence on IL-3 for cell proliferation, due to overexpression of the alpha subunit of the IL-3 receptor (IL3-RA, also known as CD123). Consistent with our in vitro data, IL3-RA was also upregulated in samples from patients with NUP98::NSD1-positive AML. These results highlight CD123 as a potential new therapeutic target in NUP98::NSD1-positive AML.







Similar content being viewed by others
Data availability
Original data presented in this manuscript are available upon request addressed to the corresponding author.
References
Gough SM, Slape CI, Aplan PD. NUP98 gene fusions and hematopoietic malignancies: common themes and new biologic insights. Blood. 2011;118(24):6247–57.
Wang GG, Cai L, Pasillas MP, Kamps MP. NUP98-NSD1 links H3K36 methylation to Hox-A gene activation and leukaemogenesis. Nat Cell Biol. 2007;9(7):804–12.
Xu H, Valerio DG, Eisold ME, Sinha A, Koche RP, Hu W, et al. NUP98 fusion proteins interact with the NSL and MLL1 complexes to drive Leukemogenesis. Cancer Cell. 2016;30(6):863–78.
Heikamp EB, Henrich JA, Perner F, Wong EM, Hatton C, Wen Y, et al. The menin-MLL1 interaction is a molecular dependency in NUP98-rearranged AML. Blood. 2022;139(6):894–906.
Cerveira N, Correia C, Dória S, Bizarro S, Rocha P, Gomes P, et al. Frequency of NUP98–NSD1 fusion transcript in childhood acute myeloid leukaemia. Leukemia. 2003;17(11):2244–7.
Struski S, Lagarde S, Bories P, Puiseux C, Prade N, Cuccuini W, et al. NUP98 is rearranged in 3.8% of pediatric AML forming a clinical and molecular homogenous group with a poor prognosis. Leukemia. 2017;31(3):565–72.
Ostronoff F, Othus M, Gerbing RB, Loken MR, Raimondi SC, Hirsch BA, et al. NUP98/NSD1 and FLT3/ITD coexpression is more prevalent in younger AML patients and leads to induction failure: a COG and SWOG report. Blood. 2014;124(15):2400–7.
Morita S, Kojima T, Kitamura T. Plat-E: an efficient and stable system for transient packaging of retroviruses. Gene Ther. 2000;7(12):1063–6.
McNiece IK, Bradley TR, Kriegler AB, Hodgson GS. A growth factor produced by WEHI-3 cells for murine high proliferative potential GM-progenitor colony forming cells. Cell Biol Int Rep. 1982;6(3):243–51.
Fujiki A, Imamura T, Sakamoto K, Kawashima S, Yoshida H, Hirashima Y, et al. All-trans retinoic acid combined with 5-Aza-2’-deoxycytidine induces C/EBPalpha expression and growth inhibition in MLL-AF9-positive leukemic cells. Biochem Biophys Res Commun. 2012;428(2):216–23.
Yoshida H, Imamura T, Fujiki A, Hirashima Y, Miyachi M, Inukai T, et al. Post-transcriptional modulation of C/EBPalpha prompts monocytic differentiation and apoptosis in acute myelomonocytic leukaemia cells. Leuk Res. 2012;36(6):735–41.
Lavau C, Szilvassy SJ, Slany R, Cleary ML. Immortalization and leukemic transformation of a myelomonocytic precursor by retrovirally transduced HRX-ENL. EMBO J. 1997;16(14):4226–37.
Lin S, Luo RT, Ptasinska A, Kerry J, Assi SA, Wunderlich M, et al. Instructive role of MLL-fusion proteins revealed by a model of t(4;11) Pro-B acute lymphoblastic Leukemia. Cancer Cell. 2016;30(5):737–49.
Michmerhuizen NL, Klco JM, Mullighan CG. Mechanistic insights and potential therapeutic approaches for NUP98-rearranged hematologic malignancies. Blood. 2020;136(20):2275–89.
Deshpande AJ, Deshpande A, Sinha AU, Chen L, Chang J, Cihan A, et al. AF10 regulates progressive H3K79 methylation and HOX gene expression in diverse AML subtypes. Cancer Cell. 2014;26(6):896–908.
Mizuki M, Fenski R, Halfter H, Matsumura I, Schmidt R, Müller C, et al. Flt3 mutations from patients with acute myeloid leukemia induce transformation of 32D cells mediated by the Ras and STAT5 pathways. Blood. 2000;96(12):3907–14.
Peiris MN, Meyer AN, Nelson KN, Bisom-Rapp EW, Donoghue DJ. Oncogenic fusion protein BCR-FGFR1 requires the breakpoint cluster region-mediated oligomerization and chaperonin Hsp90 for activation. Haematologica. 2020;105(5):1262–73.
Testa U, Pelosi E, Frankel A. CD 123 is a membrane biomarker and a therapeutic target in hematologic malignancies. Biomark Res. 2014;2(1):4.
Shiba N, Yoshida K, Hara Y, Yamato G, Shiraishi Y, Matsuo H, et al. Transcriptome analysis offers a comprehensive illustration of the genetic background of pediatric acute myeloid leukemia. Blood Adv. 2019;3(20):3157–69.
Thanasopoulou A, Tzankov A, Schwaller J. Potent co-operation between the NUP98-NSD1 fusion and the FLT3-ITD mutation in acute myeloid leukemia induction. Haematologica. 2014;99(9):1465–71.
Hollink IH, van den Heuvel-Eibrink MM, Arentsen-Peters ST, Pratcorona M, Abbas S, Kuipers JE, et al. NUP98/NSD1 characterizes a novel poor prognostic group in acute myeloid leukemia with a distinct HOX gene expression pattern. Blood. 2011;118(13):3645–56.
Schmoellerl J, Barbosa IAM, Eder T, Brandstoetter T, Schmidt L, Maurer B, et al. CDK6 is an essential direct target of NUP98 fusion proteins in acute myeloid leukemia. Blood. 2020;136(4):387–400.
Jin L, Lee EM, Ramshaw HS, Busfield SJ, Peoppl AG, Wilkinson L, et al. Monoclonal antibody-mediated targeting of CD123, IL-3 receptor alpha chain, eliminates human acute myeloid leukemic stem cells. Cell Stem Cell. 2009;5(1):31–42.
El Achi H, Dupont E, Paul S, Khoury JD. CD123 as a biomarker in hematolymphoid malignancies: principles of detection and targeted therapies. Cancers (Basel). 2020;12(11):3087.
Jen EY, Gao X, Li L, Zhuang L, Simpson NE, Aryal B, et al. FDA approval summary: Tagraxofusp-erzs For treatment of blastic plasmacytoid dendritic cell neoplasm. Clin Cancer Res. 2020;26(3):532–6.
Cangini D, Silimbani P, Cafaro A, Giannini MB, Masini C, et al. Tagraxofusp and anti-CD123 in blastic plasmacytoid dendritic cell neoplasm: a new hope. Minerva Med. 2020;111(5):467–77.
Feng L, Xu X, Zhao K. NFYB potentiates STK33 activation to promote cisplatin resistance in diffuse large B-cell lymphoma. Leuk Res. 2021;111: 106708.
Babij C, Zhang Y, Kurzeja RJ, Munzli A, Shehabeldin A, Fernando M, et al. STK33 kinase activity is nonessential in KRAS-dependent cancer cells. Cancer Res. 2011;71(17):5818–26.
Acknowledgements
This study was supported by grants-in-aid for scientific research from the Japanese Ministry of Education, Culture, Sports, Science and Technology (17K10124 and 21K07759).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12185_2023_3612_MOESM1_ESM.pdf
Supplementary file1 Fig. 1 Homology between human and murine NUP98::NSD1 amino acid sequences. Comparison between human and murine aa sequences; aa 518 of human NUP98 (Pro) is equivalent to aa 535 (Pro) in the murine Nup98, and aa 1166 of NSD1 (Ser) is equivalent to aa 1167 (Ser) in murine Nsd1. Human and murine NUP98 aa sequences fused to NSD1 exhibited approximately 98% homology, while human and murine NSD1 aa sequences fused to NUP98 had around 85% identity. Pro, Proline; Ser, Serine. Fig. 2 RT-PCR revealed expression of Nup98::Nsd1 mRNA in transduced 32D cells. Fig. 3 Western blotting did not show constitutive phosphorylation of Jak2 and Stat5 in Nup98::Nsd1+ 32D cells. Fig. 4 (a) qPCR revealed that the expression level of Meis1 was not upregulated in Nup98::Nsd1+ 32D cells. RQ, relative quantification. (b) RT-PCR did not reveal expression of Hoxa9 mRNA in either Nup98::Nsd1+ 32D or mock-transduced 32D cells. Fig. 5 Effects of Nup98::Nsd1 on colony formation. (a), (b) Neither C57BL/6 nor BALB/c Lin- bone marrow cells were immortalized by Nup98::Nsd1. KMT2A::Aff1 served as a positive control, which we previously confirmed has immortalization ability. (c), (d) RT-PCR revealed expression of Nup98::Nsd1 mRNA in murine Lin- BM cells (we used primers designed to straddle the fusion point; therefore, the size of the PCR product is 468 bp), indicating successful transduction of Nup98::Nsd1; however, Nup98::Nsd1 did not exhibit cell transformation activity. RT-PCR of Gapdh was performed using forward (5’-CATCACTGCCACCCAGAAGACTG-3’) and reverse (5’- ATGCCAGTGAGCTTCCCGTTCAG-3’) primers; therefore, the size of the PCR product is 153 bp. BM, bone marrow; PC, positive control; NTC, no template control. Fig. 6 The sensitivity of Nup98::Nsd1 to cytarabine and daunorubicin. Nup98::Nsd1+ and mock-transduced 32D cells were cultured in IL-3 containing medium at 1 × 105 cells per well with 1, 10, 100 and 1,000 nM cytarabine or daunorubicin. Viable cell numbers were measured at 24, 48, 72 and 96 hours. (a), (b) Cytarabine did not attenuate cell proliferation of Nup98::Nsd1+ 32D cells. (c), (d) Daunorubicin did not visibly attenuate cell proliferation of Nup98::Nsd1+ 32D cells. (e), (f) STK33 inhibitor had no effect to suppress the proliferation of Nup98::Nsd1+ 32D cells, because the vehicle control (indicated in green) was at the bottom. (PDF 691 KB)
12185_2023_3612_MOESM2_ESM.xlsx
Supplementary file2 Table 1 Lists of the top 10 up- and down-regulated genes in Nup98::Nsd1+ 32D cells, Table 2 Lists of the top 10 oncogenic signature gene sets (C6) enriched in Nup98::Nsd1+ 32D cells. (XLSX 15 KB)
About this article
Cite this article
Okamoto, K., Imamura, T., Tanaka, S. et al. The Nup98::Nsd1 fusion gene induces CD123 expression in 32D cells. Int J Hematol 118, 277–287 (2023). https://doi.org/10.1007/s12185-023-03612-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12185-023-03612-z