Abstract
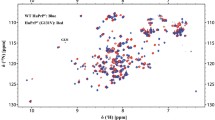
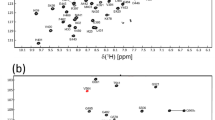
Human small EDRK-rich factor protein SERF2 is a cellular driver of protein amyloid formation, a process that has been linked to neurodegenerative diseases including Alzheimer’s and Parkinson’s disease. SERF2 is a 59 amino acid protein, highly charged, and well conserved whose structure and physiological function is unclear. SERF family proteins including human SERF2 have shown a tendency to form fuzzy complexes with misfolded proteins such as α-Synuclein which has been linked to Parkinson’s disease. SERF family proteins have been recently identified to bind nucleic acids, but the binding mechanism(s) remain enigmatic. Here, using multidimensional solution NMR, we report the 1H, 15N, and 13C chemical shift assignments (~ 86% of backbone resonance assignments) for human SERF2. TALOS-N predicted secondary structure of SERF2 showed three very short helices (3–4 residues long) in the N-terminal region of the protein and a long helix in the C-terminal region spanning residues 37–46 which is consistent with the helical content indicated by circular dichroism spectroscopy. Paramagnetic relaxation enhancement NMR analysis revealed that a short C-terminal region E53-K55 is in the proximity of the N-terminus. Having the backbone assignment of SERF2 allowed us to probe its interaction with α-Synuclein and to identify the residues in SERF2 binding interfaces that likely promote α-Synuclein aggregation.



Similar content being viewed by others
Data availability
The 1H, 13C, and 15N chemical shifts of human SERF2 has been deposited to the BioMagResBank (www.bmrb.wisc.edu) under the accession number BMRB-52141.
References
Cleverley K, Lee WC, Mumford P et al (2021) A novel knockout mouse for the small EDRK-rich factor 2 (Serf2) showing developmental and other deficits. Mamm Genome 32:94–103. https://doi.org/10.1007/s00335-021-09864-6
Falsone SF, Meyer NH, Schrank E et al (2012) SERF protein is a direct modifier of amyloid Fiber assembly. Cell Rep 2:358–371. https://doi.org/10.1016/j.celrep.2012.06.012
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31. https://doi.org/10.1093/bioinformatics/btu830
Liu Y, Wang C, Jin Y et al (2022) Backbone resonance assignments and dynamics of S. Cerevisiae SERF. Biomol NMR Assign. https://doi.org/10.1007/s12104-022-10077-4. 16:
Meinen BA, Gadkari VV, Stull F et al (2019) SERF engages in a fuzzy complex that accelerates primary nucleation of amyloid proteins. 116:23040–23049. https://doi.org/10.6084/m9.figshare.c.4684733.v1
Meyer NH, Dellago H, Tam-Amersdorfer C et al (2020) Structural fuzziness of the RNA-Organizing protein SERF determines a toxic gain-of-interaction. J Mol Biol 432:930–951. https://doi.org/10.1016/j.jmb.2019.11.014
Micsonai A, Wien F, Bulyáki É et al (2018) BeStSel: a web server for accurate protein secondary structure prediction and fold recognition from the circular dichroism spectra. Nucleic Acids Res 46. https://doi.org/10.1093/nar/gky497
Pras A, Houben B, Aprile FA et al (2021) The cellular modifier MOAG-4/SERF drives amyloid formation through charge complementation. EMBO J 40. https://doi.org/10.15252/embj.2020107568
Sahoo BR, Bardwell JCA (2022) SERF, a family of tiny highly conserved, highly charged proteins with enigmatic functions. FEBS J. https://doi.org/10.1111/febs.16555
Sahoo BR, Deng X, Wong EL et al (2023) SERF2, an RNA G-quadruplex binding protein, promotes stress granule formation. https://doi.org/10.1101/2023.10.09.561572. bioRxiv
Sahoo BR, Kocman V, Clark N et al (2023a) Effects of protein G-quadruplex interactions on phase transitions and protein aggregation. https://doi.org/10.1101/2023.09.21.558871. bioRxiv
Shen Y, Bax A (2015) Protein structural information derived from NMR chemical shift with the neural network program TALOS-N. Methods Mol Biol 1260. https://doi.org/10.1007/978-1-4939-2239-0_2
Stefanis L (2012) α-Synuclein in Parkinson’s disease. Cold Spring Harb Perspect Med. https://doi.org/10.1101/cshperspect.a009399. 2:
Stroo E, Janssen L, Sin O et al (2023) Deletion of SERF2 in mice delays embryonic development and alters amyloid deposit structure in the brain. Life Sci Alliance 6. https://doi.org/10.26508/lsa.202201730
van Ham TJ, Holmberg MA, van der Goot AT et al (2010) Identification of MOAG-4/SERF as a regulator of age-related proteotoxicity. Cell 142:601–612. https://doi.org/10.1016/j.cell.2010.07.020
Yamaguchi T, Matsuzaki K, Hoshino M (2011) Transient formation of intermediate conformational states of amyloid-β peptide revealed by heteronuclear magnetic resonance spectroscopy. FEBS Lett 585:1097–1102. https://doi.org/10.1016/j.febslet.2011.03.014
Yoshimura Y, Holmberg MA, Kukic P et al (2017) MOAG-4 promotes the aggregation of α-synuclein by competing with self-protective electrostatic interactions. J Biol Chem 292:8269–8278. https://doi.org/10.1074/jbc.M116.764886
Acknowledgements
J.C.A.B. is a Howard Hughes Medical Institute (HHMI) Investigator and this study was supported by funds from HHMI.
Funding
This study was supported by funds from Howard Hughes Medical Institute to J.C.A.B.
Author information
Authors and Affiliations
Contributions
B.S. and J.C.A.B. planned the study and designed experiments. B.S. performed experiments. B.S. and V.S. interpreted the results. B.S. and J.C.A.B. wrote the manuscript.
Corresponding author
Ethics declarations
Ethical approval
The authors declare that the experiments described in this publication were done in compliance with the ethical standards of the country in which they were performed.
Consent for publication
Yes.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sahoo, B.R., Subramanian, V. & Bardwell, J.C. Backbone 1H, 13C, and 15N chemical shift assignments for human SERF2. Biomol NMR Assign (2024). https://doi.org/10.1007/s12104-024-10167-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12104-024-10167-5