Abstract
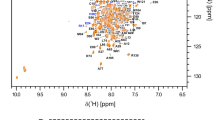
The Endosomal Sorting Complex Required for Transport (ESCRT) pathway, through inverse topology membrane remodeling, is involved in many biological functions, such as ubiquitinated membrane receptor trafficking and degradation, multivesicular bodies (MVB) formation and cytokinesis. Dysfunctions in ESCRT pathway have been associated to several human pathologies, such as cancers and neurodegenerative diseases. The ESCRT machinery is also hijacked by many enveloped viruses to bud away from the plasma membrane of infected cells. Human tumor susceptibility gene 101 (Tsg101) protein is an important ESCRT-I complex component. The structure of the N-terminal ubiquitin E2 variant (UEV) domain of Tsg101 (Tsg101-UEV) comprises an ubiquitin binding pocket next to a late domain [P(S/T)AP] binding groove. These two binding sites have been shown to be involved both in the physiological roles of ESCRT-I and in the release of the viral particles, and thus are attractive targets for antivirals. The structure of the Tsg101-UEV domain has been characterized, using X-ray crystallography or NMR spectroscopy, either in its apo-state or bound to ubiquitin or late domains. In this study, we report the backbone NMR resonance assignments, including the proline signals, of the apo human Tsg101-UEV domain, that so far was not publicly available. These data, that are in good agreement with the crystallographic structure of Tsg101-UEV domain, can therefore be used for further NMR studies, including protein-protein interaction studies and drug discovery.



Similar content being viewed by others
Data availability
All the chemical shift values of apo human Tsg101-UEV domain were deposited in the Biological Magnetic Resonance Data Bank (BMRB) under accession code 50765.
References
Bouamr F, Melillo JA, Wang MQ et al (2003) PPPYEPTAP Motif is the late domain of human T-Cell leukemia virus type 1 gag and mediates its Functional Interaction with Cellular Proteins Nedd4 and Tsg101. J Virol 77:11882–11895. https://doi.org/10.1128/JVI.77.22.11882-11895.2003
Boura E, Różycki B, Herrick DZ et al (2011) Solution structure of the ESCRT-I complex by small-angle X-ray scattering, EPR, and FRET spectroscopy. Proc Natl Acad Sci 108:9437–9442. https://doi.org/10.1073/pnas.1101763108
Dolnik O, Kolesnikova L, Welsch S et al (2014) Interaction with Tsg101 is necessary for the efficient transport and release of Nucleocapsids in Marburg Virus-Infected cells. PLoS Pathog 10:e1004463. https://doi.org/10.1371/journal.ppat.1004463
Ferraiuolo R-M, Manthey KC, Stanton MJ et al (2020) The multifaceted roles of the Tumor susceptibility gene 101 (TSG101) in Normal Development and Disease. Cancers 12:450. https://doi.org/10.3390/cancers12020450
Flower TG, Takahashi Y, Hudait A et al (2020) A helical assembly of human ESCRT-I scaffolds reverse-topology membrane scission. Nat Struct Mol Biol 27:570–580. https://doi.org/10.1038/s41594-020-0426-4
Freed EO (2002) Viral late domains. J Virol 76:4679–4687. https://doi.org/10.1128/JVI.76.10.4679-4687.2002
Garrus JE, von Schwedler UK, Pornillos OW et al (2001) Tsg101 and the Vacuolar protein sorting pathway are essential for HIV-1 budding. Cell 107:55–65. https://doi.org/10.1016/S0092-8674(01)00506-2
Hafsa NE, Arndt D, Wishart DS (2015) CSI 3.0: a web server for identifying secondary and super-secondary structure in proteins using NMR chemical shifts. Nucleic Acids Res 43:W370–W377. https://doi.org/10.1093/nar/gkv494
Henne WM, Buchkovich NJ, Emr SD (2011) The ESCRT pathway. Dev Cell 21:77–91. https://doi.org/10.1016/j.devcel.2011.05.015
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinforma Oxf Engl 31:1325–1327. https://doi.org/10.1093/bioinformatics/btu830
Lee W, Bahrami A, Dashti HT et al (2019) I-PINE web server: an integrative probabilistic NMR assignment system for proteins. J Biomol NMR 73:213–222. https://doi.org/10.1007/s10858-019-00255-3
Leis J, Luan C-H, Audia JE et al (2021) Ilaprazole and other Novel Prazole-Based Compounds that bind Tsg101 inhibit viral budding of herpes simplex virus 1 and 2 and human immunodeficiency virus from cells. J Virol 95:e00190–e00121. https://doi.org/10.1128/JVI.00190-21
Lennard KR, Gardner RM, Doigneaux C et al (2019) Development of a cyclic peptide inhibitor of the p6/UEV protein–protein Interaction. ACS Chem Biol 14:1874–1878. https://doi.org/10.1021/acschembio.9b00627
Lu Q, Hope LW, Brasch M et al (2003) TSG101 interaction with HRS mediates endosomal trafficking and receptor down-regulation. Proc Natl Acad Sci 100:7626–7631. https://doi.org/10.1073/pnas.0932599100
Marsh JA, Singh VK, Jia Z, Forman-Kay JD (2006) Sensitivity of secondary structure propensities to sequence differences between α- and γ-synuclein: implications for fibrillation. Protein Sci 15:2795–2804. https://doi.org/10.1110/ps.062465306
Martin-Serrano J, Zang T, Bieniasz PD (2001) HIV-1 and Ebola virus encode small peptide motifs that recruit Tsg101 to sites of particle assembly to facilitate egress. Nat Med 7:1313–1319. https://doi.org/10.1038/nm1201-1313
McDonald B, Martin-Serrano J (2008) Regulation of Tsg101 expression by the steadiness box: a role of Tsg101-associated ligase. Mol Biol Cell 19:754–763. https://doi.org/10.1091/mbc.e07-09-0957
Nagashima S, Takahashi M, Jirintai S et al (2011) Tumour susceptibility gene 101 and the vacuolar protein sorting pathway are required for the release of hepatitis E virions. J Gen Virol 92:2838–2848. https://doi.org/10.1099/vir.0.035378-0
Palencia A, Martinez JC, Mateo PL et al (2006) Structure of human TSG101 UEV domain. Acta Crystallogr D Biol Crystallogr 62:458–464. https://doi.org/10.1107/S0907444906005221
Pornillos O, Alam SL, Davis DR, Sundquist WI (2002a) Structure of the Tsg101 UEV domain in complex with the PTAP motif of the HIV-1 p6 protein. Nat Struct Mol Biol 9:812–817. https://doi.org/10.1038/nsb856
Pornillos O, Alam SL, Rich RL et al (2002b) Structure and functional interactions of the Tsg101 UEV domain. EMBO J 21:2397–2406. https://doi.org/10.1093/emboj/21.10.2397
Schmidt O, Teis D (2012) The ESCRT machinery. Curr Biol 22:R116–R120. https://doi.org/10.1016/j.cub.2012.01.028
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–223. https://doi.org/10.1007/s10858-009-9333-z
Siarot L, Chutiwitoonchai N, Sato H et al (2018) Identification of human immunodeficiency virus type-1 Gag-TSG101 interaction inhibitors by high-throughput screening. Biochem Biophys Res Commun 503:2970–2976. https://doi.org/10.1016/j.bbrc.2018.08.079
Strickland M, Ehrlich LS, Watanabe S et al (2017) Tsg101 chaperone function revealed by HIV-1 assembly inhibitors. Nat Commun 8:1391. https://doi.org/10.1038/s41467-017-01426-2
Tavassoli A, Lu Q, Gam J et al (2008) Inhibition of HIV budding by a genetically selected cyclic peptide targeting the gag – TSG101 Interaction. ACS Chem Biol 3:757–764. https://doi.org/10.1021/cb800193n
VerPlank L, Bouamr F, LaGrassa TJ et al (2001) Tsg101, a homologue of ubiquitin-conjugating (E2) enzymes, binds the L domain in HIV type 1 Pr55Gag. Proc Natl Acad Sci 98:7724–7729. https://doi.org/10.1073/pnas.131059198
Vietri M, Radulovic M, Stenmark H (2020) The many functions of ESCRTs. Nat Rev Mol Cell Biol 21:25–42. https://doi.org/10.1038/s41580-019-0177-4
Williams RL, Urbé S (2007) The emerging shape of the ESCRT machinery. Nat Rev Mol Cell Biol 8:355–368. https://doi.org/10.1038/nrm2162
Acknowledgements
The NMR facilities were funded by the Nord Region Council, CNRS, Institut Pasteur de Lille, European Union (FEDER), French Research Ministry and University of Lille. Financial support from the NMR division of Infranalytics (FR 2054 CNRS) is gratefully acknowledged.
Funding
This study was supported by the French National Agency for Research on AIDS and Viral Hepatitis (ANRS) Grant ECTZ101316, and a PhD fellowship from ANRS to D.M. (ECTZ103422).
Author information
Authors and Affiliations
Contributions
D.M. and X.H performed protein expression and purification. F-X.C. and E.B. collected the NMR data. D.M. analysed the data and wrote the first draft of the manuscript. D.M. prepared Figs. 1, 2 and 3. All authors commented on previous versions of the manuscript. All authors reviewed the manuscript. Funding acquisition and supervision were done by X.H.
Corresponding author
Ethics declarations
The authors declare no competing interests.
Ethics approval and consent to participate
Not applicable.
Consent for publication
All authors have agreed to the publication of the manuscript.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Moschidi, D., Cantrelle, FX., Boll, E. et al. Backbone NMR resonance assignment of the apo human Tsg101-UEV domain. Biomol NMR Assign 17, 49–54 (2023). https://doi.org/10.1007/s12104-023-10119-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-023-10119-5