Abstract
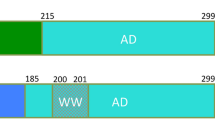
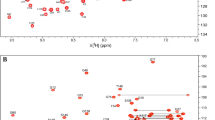
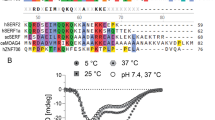
The human MDM2 protein regulates the tumor suppressor protein p53 by restricting its transcriptional activity and by promoting p53 degradation. MDM2 is ubiquitously expressed, with its overexpression implicated in many forms of cancer. The inhibitory effects of MDM2 on p53 have been shown to involve its N-terminal p53-binding domain and its C-terminal RING domain. The presence of an intact central acidic domain of MDM2 has also been shown to regulate p53 ubiquitination, with this domain shown to directly interact with the p53 DNA-binding domain to regulate the DNA binding activity of p53. To date, little structural information has been obtained for the MDM2 acidic domain. Thus, to gain insight into the structure and function relationship of this region, we have applied solution-state NMR spectroscopy to characterize the segment of MDM2 spanning residues 215–300. These boundaries for the acidic domain were determined on the basis of consensus observed in multiple sequence alignment. Here, we report the 1H, 15N and 13C backbone and 13Cβ chemical shift assignments and steady-state {1H}-15N heteronuclear NOE enhancement factors as a function of residue for the acidic domain of MDM2. We show that this domain exhibits the hallmarks of being a disordered protein, on the basis both of assigned chemical shifts and residue-level backbone dynamics, with localized variation in secondary structure propensity inferred from chemical shift analysis.





Similar content being viewed by others
Data Availability
The assignments for the backbone resonances and the {1H}-15N heteronuclear NOE enhancement factors have been deposited at the Biological Magnetic Resonance Data Bank under the Accession Number 51334. The datasets and materials generated during and/or analyzed during the current study are available from the corresponding authors on reasonable request.
References
Bálint É, Vousden KH (2001) Activation and activities of the p53 tumour suppressor protein. Br J Cancer 85:1813–1823. https://doi.org/10.1054/bjoc.2001.2128
Camilloni C, De Simone A, Vranken WF, Vendruscolo M (2012) Determination of secondary structure populations in disordered states of proteins using nuclear magnetic resonance chemical shifts. Biochemistry 51:2224–2231. https://doi.org/10.1021/bi3001825
Cross B, Chen L, Cheng Q, Li B, Yuan ZM, Chen J (2011) Inhibition of p53 DNA binding function by the MDM2 protein acidic domain. J Biol Chem 286:16018–16029. https://doi.org/10.1074/jbc.M111.228981
Dai MS, Lu H (2004) Inhibition of MDM2-mediated p53 ubiquitination and degradation by ribosomal protein L5. J Biol Chem 279:44475–44482. https://doi.org/10.1074/jbc.M403722200
Dai MS, Zeng SX, Jin Y, Sun XX, David L, Lu H (2004) Ribosomal protein L23 activates p53 by inhibiting MDM2 function in response to ribosomal perturbation but not to translation inhibition. Mol Cell Biol 24:7654–7668. https://doi.org/10.1128/mcb.24.17.7654-7668.2004
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293. https://doi.org/10.1007/bf00197809
Dyson HJ, Wright PE (2021) NMR illuminates intrinsic disorder. Curr Opin Struct Biol 70:44–52. https://doi.org/10.1016/j.sbi.2021.03.015
Fang S, Jensen JP, Ludwig RL, Vousden KH, Weissman AM (2000) Mdm2 is a RING finger-dependent ubiquitin protein ligase for itself and p53. J Biol Chem 275:8945–8951. https://doi.org/10.1074/jbc.275.12.8945
Farrow NA, Muhandiram R, Singer AU, Pascal SM, Kay CM, Gish G, Shoelson SE, Pawson T, Forman-Kay JD, Kay LE (1994) Backbone dynamics of a free and phosphopeptide-complexed Src homology 2 domain studied by 15 N NMR relaxation. Biochemistry 33:5984–6003. https://doi.org/10.1021/bi00185a040
Fuchs SY, Adler V, Buschmann T, Wu X, Ronai Z (1998) Mdm2 association with p53 targets its ubiquitination. Oncogene 17:2543–2547. https://doi.org/10.1038/sj.onc.1202200
Honda R, Tanaka H, Yasuda H (1997) Oncoprotein MDM2 is a ubiquitin ligase E3 for tumor suppressor p53. FEBS Lett 420:25–27. https://doi.org/10.1016/s0014-5793(97)01480-4
Honda R, Yasuda H (2000) Activity of MDM2, a ubiquitin ligase, toward p53 or itself is dependent on the RING finger domain of the ligase. Oncogene 19:1473–1476. https://doi.org/10.1038/sj.onc.1203464
Hornbeck PV, Zhang B, Murray B, Kornhauser JM, Latham V, Skrzypek E (2015) PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Res 43:D512-520. https://doi.org/10.1093/nar/gku1267
Kay LE, Torchia DA, Bax A (1989) Backbone dynamics of proteins as studied by 15N inverse detected heteronuclear NMR spectroscopy: application to staphylococcal nuclease. Biochemistry 28:8972–8979. https://doi.org/10.1021/bi00449a003
Kruiswijk F, Labuschagne CF, Vousden KH (2015) p53 in survival, death and metabolic health: a lifeguard with a licence to kill. Nat Rev Mol Cell Biol 16:393–405. https://doi.org/10.1038/nrm4007
Kubbutat MH, Jones SN, Vousden KH (1997) Regulation of p53 stability by Mdm2. Nature 387:299–303. https://doi.org/10.1038/387299a0
Kussie PH, Gorina S, Marechal V, Elenbaas B, Moreau J, Levine AJ, Pavletich NP (1996) Structure of the MDM2 oncoprotein bound to the p53 tumor suppressor transactivation domain. Science 274:948–953. https://doi.org/10.1126/science.274.5289.948
Linke K, Mace PD, Smith CA, Vaux DL, Silke J, Day CL (2008) Structure of the MDM2/MDMX RING domain heterodimer reveals dimerization is required for their ubiquitylation in trans. Cell Death Differ 15:841–848. https://doi.org/10.1038/sj.cdd.4402309
Maciejewski MW, Schuyler AD, Gryk MR, Moraru II, Romero PR, Ulrich EL, Eghbalnia HR, Livny M, Delaglio F, Hoch JC (2017) NMRbox: A resource for biomolecular NMR computation. Biophys J 112:1529–1534. https://doi.org/10.1016/j.bpj.2017.03.011
Magnussen HM, Ahmed SF, Sibbet GJ, Hristova VA, Nomura K, Hock AK, Archibald LJ, Jamieson AG, Fushman D, Vousden KH, Weissman AM, Huang DT (2020) Structural basis for DNA damage-induced phosphoregulation of MDM2 RING domain. Nat Commun 11:2094. https://doi.org/10.1038/s41467-020-15783-y
Markley JL, Bax A, Arata Y, Hilbers CW, Kaptein R, Sykes BD, Wright PE, Wuthrich K (1998) Recommendations for the presentation of NMR structures of proteins and nucleic acids - IUPAC-IUBMB-IUPAB Inter-Union Task Group on the standardization of data bases of protein and nucleic acid structures determined by NMR spectroscopy. Eur J Biochem 256:1–15. https://doi.org/10.1046/j.1432-1327.1998.2560001.x
Momand J, Jung D, Wilczynski S, Niland J (1998) The MDM2 gene amplification database. Nucleic Acids Res 26:3453–3459. https://doi.org/10.1093/nar/26.15.3453
Roth J, Dobbelstein M, Freedman DA, Shenk T, Levine AJ (1998) Nucleo-cytoplasmic shuttling of the hdm2 oncoprotein regulates the levels of the p53 protein via a pathway used by the human immunodeficiency virus rev protein. EMBO J 17:554–564. https://doi.org/10.1093/emboj/17.2.554
Sattler M, Schleucher J, Griesinger C (1999) Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog Nucl Magn Reson Spectrosc 34:93–158. https://doi.org/10.1016/S0079-6565(98)00025-9
Shaikh MF, Morano WF, Lee J, Gleeson E, Babcock BD, Michl J, Sarafraz-Yazdi E, Pincus MR, Bowne WB (2016) Emerging role of MDM2 as target for anti-cancer therapy: a review. Ann Clin Lab Sci 46:627–634
Sievers F, Higgins DG (2018) Clustal Omega for making accurate alignments of many protein sequences. Protein Sci 27:135–145. https://doi.org/10.1002/pro.3290
Song Q, Liu XQ, Rainey JK (2022) 1H, 15N and 13C backbone resonance assignments of the acidic domain of the human MDMX protein. Biomol NMR Assign 16:171–178. https://doi.org/10.1007/s12104-022-10081-8
Tal M, Silberstein A, Nusser E (1985) Why does Coomassie Brilliant Blue R interact differently with different proteins? A partial answer. J Biol Chem 260:9976–9980
The UniProt Consortium (2021) UniProt: The universal protein knowledgebase in 2021. Nucleic Acids Res 49:D480–D489. https://doi.org/10.1093/nar/gkaa1100
Tjandra N, Kuboniwa H, Ren H, Bax A (1995) Rotational dynamics of calcium-free calmodulin studied by 15N NMR relaxation measurements. Eur J Biochem 230:1014–1024. https://doi.org/10.1111/j.1432-1033.1995.tb20650.x
Uhrinova S, Uhrin D, Powers H, Watt K, Zheleva D, Fischer P, McInnes C, Barlow PN (2005) Structure of free MDM2 N-terminal domain reveals conformational adjustments that accompany p53-binding. J Mol Biol 350:587–598. https://doi.org/10.1016/j.jmb.2005.05.010
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: Development of a software pipeline. Proteins 59:687–696. https://doi.org/10.1002/prot.20449
Wang B, Wang Y, Wishart DS (2010) A probabilistic approach for validating protein NMR chemical shift assignments. J Biomol NMR 47:85–99. https://doi.org/10.1007/s10858-010-9407-y
Waterhouse AM, Procter JB, Martin DM, Clamp M, Barton GJ (2009) Jalview Version 2 - A multiple sequence alignment editor and analysis workbench. Bioinformatics 25:1189–1191. https://doi.org/10.1093/bioinformatics/btp033
Wishart DS, Bigam CG, Yao J, Abildgaard F, Dyson HJ, Oldfield E, Markley JL, Sykes BD (1995) 1H, 13C and 15N chemical shift referencing in biomolecular NMR. J Biomol NMR 6:135–140. https://doi.org/10.1007/BF00211777
Yu GW, Allen MD, Andreeva A, Fersht AR, Bycroft M (2006) Solution structure of the C4 zinc finger domain of HDM2. Protein Sci 15:384–389. https://doi.org/10.1110/ps.051927306
Zheng J, Lang Y, Zhang Q, Cui D, Sun H, Jiang L, Chen Z, Zhang R, Gao Y, Tian W, Wu W, Tang J, Chen Z (2015) Structure of human MDM2 complexed with RPL11 reveals the molecular basis of p53 activation. Genes Dev 29:1524–1534. https://doi.org/10.1101/gad.261792.115
Acknowledgements
J.K.R. is grateful to Dr. Michael W. Gray for generous donation of the FPLC system. NMR experiments were acquired at QANUC and we thank Dr. Tara Sprules, NMR facility manager at QANUC, for her expert NMR data acquisition and technical support. We are also grateful for access to the NMRbox resource for data processing and analysis.
Funding
This work was supported by Discovery Grants from the Natural Sciences and Engineering Research Council of Canada (NSERC, RGPIN/05907 − 2017 to J.K.R. and RGPIN/06083 − 2020 to X.-Q.L.) Key equipment was provided by NSERC Research Tools and Instruments Grants (to J.K.R. and X.-Q.L.) and Dalhousie Medical Research Foundation Capital Equipment Grants (to J.K.R. and X.-Q.L.) The Québec/Eastern Canada High Field NMR Facility (QANUC) is supported by the Canada Foundation for Innovation and the McGill University Faculty of Science and Department of Chemistry. NMRbox: National Center for Biomolecular NMR Data Processing and Analysis NMRbox is a Biomedical Technology Research Resource (BTRR), which is supported by NIH grant P41GM111135 (NIGMS).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. QS: material preparation was performed. QS: data processing and analysis were performed primarily working with JKR. X-QL and JKR: Funding for this work was obtained. QS: The manuscript, including figure preparation, was drafted and all authors edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, Q., Liu, XQ. & Rainey, J.K. 1H, 15N and 13C backbone resonance assignments of the acidic domain of the human MDM2 protein. Biomol NMR Assign 17, 9–16 (2023). https://doi.org/10.1007/s12104-022-10112-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-022-10112-4