Abstract
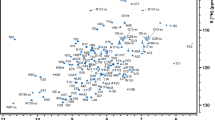
Ebola viral infections have resulted in several deadly epidemics in recent years in West and Central Africa. Because only one of the seven proteins encoded by the viral genome possesses enzymatic activity, disruption of protein–protein interactions is a promising route for antiviral drug development. We carried out a screening campaign to identify small, drug-like compounds that bind to the C-terminal region of the multifunctional Ebola nucleoprotein (eNP) with the objective of discovering ones that disrupt its binding to other Ebola proteins or to the single-stranded RNA genome. In the course of this effort we assigned the backbone 1H, 15N, and 13C resonances of residues 600‒739, the region that contains the critical eVP30 binding region 600‒615 targeted by host factors, and used the assigned chemical shifts to predict secondary structural features and peptide dynamics. This work supports and extends the previous X-ray crystal structures and NMR studies of residues 641‒739. We found that the 600‒739 domain consists of separate regions that are largely disordered and ordered.


Similar content being viewed by others
References
Batra J, Hultquist JF, Liu D, Shtanko O, Von Dollen J, Satkamp L, Jang GM, Luthra P, Schwarz TM, Small GI et al (2018) Protein interaction mapping identifies RBBP6 as a negative regulator of Ebola virus replication. Cell 175(1917–1930):e1913
Berjanskii MV, Wishart DS (2005) A simple method to predict protein flexibility using secondary chemical shifts. J Am Chem Soc 127:14970–14971. https://doi.org/10.1021/ja054842f
Delaglio F, Grzesiek S, Vuister G, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293. https://doi.org/10.1007/BF00197809
Dziubańska PJ, Derewenda U, Ellena JF, Engel DA, Derewenda ZS (2014) The structure of the C-terminal domain of the Zaire ebolavirus nucleoprotein. Acta Crystallogr D 70:2420–2429. https://doi.org/10.1107/S1399004714014710
Eghbalnia HR, Wang L, Bahrami A, Assadi A, Markley JL (2005) Protein energetic conformational analysis from NMR chemical shifts (PECAN) and its use in determining secondary structural elements. J Biomol NMR 32:71–81. https://doi.org/10.1007/s10858-005-5705-1
Lee W, Markley JL (2018) PINE-SPARKY.2 for automated NMR-based protein structure research. Bioinformatics 34:1586–1588. https://doi.org/10.1093/bioinformatics/btx785
Lee W, Westler WM, Bahrami A, Eghbalnia HR, Markley JL (2009) PINE-SPARKY: graphical interface for evaluating automated probabilistic peak assignments in protein NMR spectroscopy. Bioinformatics 25:2085–2087. https://doi.org/10.1093/bioinformatics/btp345
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327. https://doi.org/10.1093/bioinformatics/btu830
Lee W, Cornilescu G, Dashti H, Eghbalnia HR, Tonelli M, Westler WM, Butcher SE, Henzler-Wildman KA, Markley JL (2016) Integrative NMR for biomolecular research. J Biomol NMR 64:307–332. https://doi.org/10.1007/s10858-016-0029-x
Messaoudi I, Amarasinghe GK, Basler CF (2015) Filovirus pathogenesis and immune evasion: insights from Ebola virus and Marburg virus. Nat Rev Microbiol 13:663–676. https://doi.org/10.1038/nrmicro3524
Shen Y, Bax A (2013) Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J Biomol NMR 56:227–241. https://doi.org/10.1007/s10858-013-9741-y
Su Z, Wu C, Shi L, Luthra P, Pintilie GD, Johnson B, Porter JR, Ge P, Chen M, Liu G, Frederick TE, Binning JM, Bowman GR, Zhou ZH, Basler CF, Gross ML, Leung DW, Chiu W, Amarasinghe GK (2018) Electron cryo-microscopy structure of Ebola virus nucleoprotein reveals a mechanism for nucleocapsid-like assembly. Cell 172:966–978.e12. https://doi.org/10.1016/j.cell.2018.02.009
Wang L, Eghbalnia HR, Bahrami A, Markley JL (2005) Linear analysis of carbon-13 chemical shift differences and its application to the detection and correction of errors in referencing and spin system identifications. J Biomol NMR 32:13–22. https://doi.org/10.1007/s10858-005-1717-0
Xu W, Luthra P, Wu C, Batra J, Leung DW, Basler CF, Amarasinghe GK (2017) Ebola virus VP30 and nucleoprotein interactions modulate viral RNA synthesis. Nat Commun 8:15576. https://doi.org/10.1038/ncomms15576
Ying J, Delaglio F, Torchia DA, Bax A (2017) Sparse multidimensional iterative lineshape-enhanced (SMILE) reconstruction of both non-uniformly sampled and conventional NMR data. J Biomol NMR 68:101–118. https://doi.org/10.1007/s10858-016-0072-7
Acknowledgements
WL, MT, and JLM were supported by NIH Grant P41GM103399 (NIGMS), DA, CW, and GKA were supported by NIH Grant R01AI123926 and P01AI120943. This study made use of the National Magnetic Resonance Facility at Madison, which is supported by NIH Grant P41GM103399 (NIGMS). Equipment at NMRFAM was purchased with funds from the University of Wisconsin-Madison, NIH Grants P41GM103399, S10RR02781, S10RR08438, S10RR023438, S10RR025062, and S10RR029220), and NSF Grants DMB-8415048, OIA-9977486, and BIR-9214394.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lee, W., Tonelli, M., Wu, C. et al. Backbone resonance assignments and secondary structure of Ebola nucleoprotein 600–739 construct. Biomol NMR Assign 13, 315–319 (2019). https://doi.org/10.1007/s12104-019-09898-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-019-09898-7