Abstract
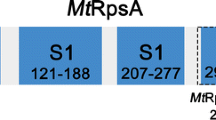
RpsA, also known as ribosomal protein S1, is an essential protein required for translation initiation of mRNAs when their Shine-Dalgarno sequence is degenerated (Sorensen et al. 1998). In addition, RpsA of Mycobacterium tuberculosis (M. tb) is involved in trans-translation, which is an effective system mediated by tmRNA-SmpB to release stalled ribosomes from mRNA in the presence of rare codons (Keiler 2008). Shi et al. found that POA binds to RpsA of Mtb and disrupts the formation of RpsA–tmRNA complex (Shi et al. 2011) and mutations at the C-terminus of RpsA confer PZA resistance. The previous work reported the pyrazinoic acid-binding domain of RpsA (Yang et al. Mol Microbiol 95:791–803, 2015). However, the HSQC spectra of the isolated S1 domain does not overlap with that of MtRpsA280-438, suggesting that substantial interactions occur between the flexible C-terminus and the S1 domain in MtRpsA .To further study the PZA resistance and how substantial interactions influence/affect protein structure, using heteronuclear NMR spectroscopy, we have completed backbone and side-chain 1H, 15N, 13C chemical shift assignments of MtRpsA280-438 which contains S1 domain and the flexible C-terminus. These NMR resonance assignments provide the framework for detailed characterization of the solution-state protein structure determination, dynamic studies of this domain, as well as NMR-based drug discovery efforts.



Similar content being viewed by others
References
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax AD (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol NMR 6:277–293
Huang B, Fu J, Guo C et al (2016) 1H, 15N, 13C resonance assignments for pyrazinoic acid binding domain of ribosomal protein S1 from Mycobacterium tuberculosis. Biomol NMR Assign 10:321
Salah P, Bisaglia M, Aliprandi P, Uzan M, Sizun C, Bontems F (2009) Probing the relationship between Gram-negative and Gram-positive S1 proteins by sequence analysis. Nucleic Acids Res 37:5578–5588
Shi et al (2011) Pyrazinamide inhibits trans-translation in Mycobacterium tuberculosis. Science 333:1630–1632
Takada K et al (2007) In vitro trans-translation of Thermus thermophilus: ribosomal protein S1 is not required for the early stage of trans-translation. Rna-a Publ Rna Society 13(4):503–510
Vranken WF, Boucher W, Stevens TJ et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins Struct Funct Bioinf 59(4):687–696
Wishart DS, Sykes BD (1994). Chemical shifts as a tool for structure determination. Methods Enzymol 239:363–392
Yang JL, Bi Y, Cai J, Liao Q, Li X, Guo W, Zhang C, Lin Q, Zhao T, Wang Y, Liu H, Zhang J, Lin X, D (2015) Structural basis for targeting the ribosomal protein S1 of Mycobacterium tuberculosis by pyrazinamide. Mol Microbiol 95:791–803
Zhi L, Xu Y, Shuai Y, et al (2006) Sequence-specific assignment of aromatic resonances of uniformly 13C,15N-labeled proteins by using 13C- and 15N-edited NOESY spectra. Angew Chem 45(12):1960
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (No. 31270777) and National Key Research and Development Project of China (No. 2016YFA0500600). We thank Jin Zhang for his help with data processing and analysis.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Rights and permissions
About this article
Cite this article
Fu, J., Huang, B., Lin, D. et al. Chemical shift assignments of Ribosomal protein S1 from Mycobacterium tuberculosis . Biomol NMR Assign 11, 133–136 (2017). https://doi.org/10.1007/s12104-017-9734-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-017-9734-y