Abstract
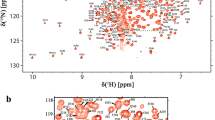
Ubiquitination of proteins is required to regulate several cellular mechanisms in cells. Skp1-Cullin-1-F-box (SCF), the largest family of the RING E3 ligases, recognizes and carries out the poly-ubiquitination of many substrate proteins. SCF E3 ligase is a multi-component protein complex, and the human S-phase kinase-associated protein 1 (Skp1) is the adapter protein, which binds and presents the substrate binding protein F-box (FBP) to the rest of the E3 ligase. Several crystallographic studies have solved the partial structure of Skp1 in complex with various FBPs, but there is no structure of standalone Skp1. Understanding the conformational and structural properties of Skp1 with and without FBPs is required to understand the complete mechanism of poly-ubiquitination. Here, we report ~90 % backbone and 64 % side chain 1H, 13C, 15N assignments of Skp1 protein using various double and triple resonance NMR experiments.



Similar content being viewed by others
References
Bielskienė K, Bagdonienė L, Mozūraitienė J, Kazbarienė B, Janulionis E (2015) E3 ubiquitin ligases as drug targets and prognostic biomarkers in melanoma. Medicina 51:1–9. doi:10.1016/j.medici.2015.01.007
Cardozo T, Pagano M (2004) The SCF ubiquitin ligase: insights into a molecular machine. Nat Rev Mol Cell Biol 5:739–751
Chen ZJJ, Sun LJJ (2009) Nonproteolytic functions of ubiquitin in cell signaling. Mol Cell 33:275–286. doi:10.1016/j.molcel.2009.01.014
Ciechanover A (2005) Intracellular protein degradation: from a vague idea thru the lysosome and the ubiquitin-proteasome system and onto human diseases and drug targeting. Cell Death Differ 12:1178–1190
Dantu SC, Kachariya NN, Kumar A (2016) Molecular dynamics simulations elucidate the mode of protein recognition by Skp1 and the F-box domain in the SCF complex. Proteins 84:159–171. doi:10.1002/prot.24963
Deshaies RJ, Joazeiro CAP (2009) RING domain E3 ubiquitin ligases. Annu Rev Biochem 78:399–434. doi:10.1146/annurev.biochem.78.101807.093809
Hao B, Zheng N, Schulman BA, Wu G, Miller JJ, Pagano M, Pavletich NP (2005) Structural basis of the Cks1-dependent recognition of p27Kip1 by the SCFSkp2 ubiquitin ligase. Mol Cell 20:9–19. doi:10.1016/j.molcel.2005.09.003
Hao B, Oehlmann S, Sowa ME, Harper JW, Pavletich NP (2007) Structure of a Fbw7-Skp1-Cyclin E complex: multisite-phosphorylated substrate recognition by SCF ubiquitin ligases. Mol Cell 26:131–143. doi:10.1016/j.molcel.2007.02.022
Hershko A (1983) Ubiquitin: roles in protein modification and breakdown. Cell 34:11–12. doi:10.1016/0092-8674(83)90131-9
Jia L, Sun Y (2011) SCF E3 ubiquitin ligases as anticancer targets. Curr Cancer Drug Targets 11:347–356. doi:10.2174/156800911794519734
Komander D (2009) The emerging complexity of protein ubiquitination. Biochem Soc Trans 37:937–953
Kumanomidou T et al (2015) The structural differences between a glycoprotein specific F-Box protein Fbs1 and its homologous protein FBG3. PLoS One 10:e0140366. doi:10.1371/journal.pone.0140366
Mandel SA, Fishman-Jacob T, Youdim MBH (2012) Genetic reduction of the E3 ubiquitin ligase element, SKP1A and environmental manipulation to emulate cardinal features of Parkinson’s disease. Parkinsonism Relat Disord 18(Supplement 1):S177–S179. doi:10.1016/S1353-8020(11)70055-4
Mizushima T, Yoshida Y, Kumanomidou T, Hasegawa Y, Suzuki A, Yamane T, Tanaka K (2007) Structural basis for the selection of glycosylated substrates by SCFFbs1 ubiquitin ligase. Proc Natl Acad Sci U S A 104:5777–5781. doi:10.1073/pnas.0610312104
Molochnikov L et al (2012) A molecular signature in blood identifies early Parkinson’s disease. MolNeurodegener 7:1–10. doi:10.1186/1750-1326-7-26
Schulman BA et al (2000) Insights into SCF ubiquitin ligases from the structure of the Skp1–Skp2 complex. Nature 408:381–386. http://www.nature.com/nature/journal/v408/n6810/suppinfo/408381a0_S1.html
Vranken WF et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696. doi:10.1002/prot.20449
Wang J, Maldonado MA (2006) The ubiquitin-proteasome system and its role in inflammatory and autoimmune diseases. Cell Mol Immunol 3:255–261
Wishart DS, Sykes BD (1994) The 13C Chemical-Shift Index: a simple method for the identification of protein secondary structure using 13C chemical-shift data. J Biomol NMR 4:171–180. doi:10.1007/bf00175245
Wishart DS, Sykes BD, Richards FM (1992) The chemical shift index: a fast and simple method for the assignment of protein secondary structure through NMR spectroscopy. Biochemistry 31:1647–1651. doi:10.1021/bi00121a010
Wishart DS, Bigam CG, Holm A, Hodges RS, Sykes BD (1995) 1H, 13C and 15N random coil NMR chemical shifts of the common amino acids. I. Investigations of nearest-neighbor effects. J Biomol NMR 5:67–81. doi:10.1007/bf00227471
Wu G, Xu G, Schulman BA, Jeffrey PD, Harper JW, Pavletich NP (2003) Structure of a β-TrCP1-Skp1-β-Catenin complex: destruction motif binding and lysine specificity of the SCFβ-TrCP1 ubiquitin ligase. Mol Cell 11:1445–1456. doi:10.1016/S1097-2765(03)00234-X
Zheng N et al (2002) Structure of the Cul1-Rbx1-Skp1-F boxSkp2 SCF ubiquitin ligase complex. Nature 416:703–709. http://www.nature.com/nature/journal/v416/n6882/suppinfo/416703a_S1.html
Acknowledgments
NNK was funded by the Department of Science and Technology (DST: Grant Number: SR/SO/BB/022/2012), Govt. of India. SCD was funded by the Ramalingaswamy fellowship (BT/RLF/Re-entry/22/2010) awarded to Ashutosh Kumar. Research was funded by DST (Grant Number: SR/SO/BB/022/2012, India) and Council of Scientific and Industrial Research (Grant Number: 37(1509)/11/EMRII). We are very grateful to Research Infrastructure Funding Committee and Industrial Research & Consultancy Centre for providing 750 MHz NMR facility and MALDI-TOFF facility at IIT Bombay. We are also thankful to Tata Institute of Fundamental Research, Mumbai, India for 800 MHz NMR facility. We also acknowledge the help received from Dr. Vaibhav Kumar Shukla, Dr. Sheeja Vasudevan, and Priyatosh Ranjan in analysing the data.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kachariya, N.N., Dantu, S.C. & Kumar, A. Backbone and side chain assignments of human cell cycle regulatory protein S-phase kinase-associated protein 1. Biomol NMR Assign 10, 351–355 (2016). https://doi.org/10.1007/s12104-016-9699-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-016-9699-2