Abstract
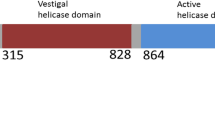
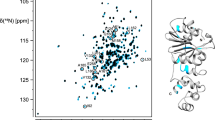
DNA ligase D (LigD), consisting of polymerase, ligase and phosphoesterase domains, is the essential catalyst of the bacterial non-homologous end-joining pathway of DNA double-strand break repair. The phosphoesterase (PE) module performs manganese-dependent 3′-phosphomonoesterase and 3′-ribonucleoside resection reactions that heal broken ends in preparation for sealing. LigD PE exemplifies a structurally and mechanistically unique class of DNA end-processing enzymes. Here, we present the resonance assignments of the PE domain of Pseudomonas aeruginosa LigD comprising the N-terminal 177 residues.


Similar content being viewed by others
References
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Johnson BA (2004) Using NMRView to visualize and analyze the NMR spectra of macromolecules. Methods Mol Biol 278:313–352
Marsh JA, Singh VK, Jia Z, Forman-Kay JD (2006) Sensitivity of secondary structure propensities to sequence differences between alpha- and gamma-synuclein: implications for fibrillation. Protein Sci 15:2795–2804
Nair PA, Smith P, Shuman S (2010) Structure of bacterial LigD 3′-phosphoesterase unveils a DNA repair superfamily. Proc Natl Acad Sci USA 107:12822–12827
Schubert M, Labudde D, Oschkinat H, Schmieder P (2002) A software tool for the prediction of Xaa-Pro peptide bond conformations in proteins based on 13C chemical shift statistics. J Biomol NMR 24:149–154
Shen Y, Bax A (2009) Prediction of Xaa-Pro peptide bond conformation from sequence and chemical shifts. J Biomol NMR 46:199–204
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–223
Shuman S, Glickman MS (2007) Bacterial DNA repair by non-homologous end joining. Nat Rev Microbiol 5:852–861
Venters RA, Farmer BT 2nd, Fierke CA, Spicer LD (1996) Characterizing the use of perdeuteration in NMR studies of large proteins: 13C, 15N and 1H assignments of human carbonic anhydrase II. J Mol Biol 264:1101–1116
Zhu H, Shuman S (2005) Novel 3′-ribonuclease and 3′-phosphatase activities of the bacterial non-homologous end-joining protein, DNA ligase D. J Biol Chem 280:25973–25981
Zhu H, Wang LK, Shuman S (2005) Essential constituents of the 3′-phosphoesterase domain of bacterial DNA ligase D, a nonhomologous end-joining enzyme. J Biol Chem 280:33707–33715
Acknowledgments
This work was supported by the following grants: NSF MCB 083141 (RG); NIH GM63611 (SS) and NIH 5G12 RR03060 (partial support of the core facilities at CCNY). SS is an American Cancer Society Research Professor. RG is a member of the New York Structural Biology Center, a STAR center supported by the New York State Office for Science, Technology and Academic Research.
Author information
Authors and Affiliations
Corresponding author
Additional information
Kaushik Dutta and Aswin Natarajan contributed equally to this work.
Rights and permissions
About this article
Cite this article
Dutta, K., Natarajan, A., Nair, P.A. et al. Sequence-specific 1H, 13C and 15N assignments of the phosphoesterase (PE) domain of Pseudomonas aeruginosa DNA ligase D (LigD). Biomol NMR Assign 5, 151–155 (2011). https://doi.org/10.1007/s12104-010-9289-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-010-9289-7