Abstract
Background
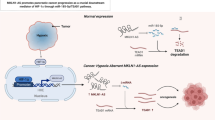
Renal cancer is one of the common malignant tumors of the urinary tract, prone to distant metastasis and drug resistance, with a poor clinical prognosis. SLC14A1 belongs to the solute transporter family, which plays a role in urinary concentration and urea nitrogen recycling in the renal, and is closely associated with the development of a variety of tumors.
Methods
Transcription data for renal clear cell carcinoma (KIRC) were obtained from the public databases Gene Expression Omnibus database (GEO) and The Cancer Genome Atlas (TCGA), and we investigated the differences in SLC14A1 expression in cancerous and normal tissues of renal cancer, its correlation with the clinicopathological features of renal cancer patients. Then, we verified the expression levels of SLC14A1 in renal cancer tissues and their Paracancerous tissues using RT-PCR, Western-blotting and immunohistochemistry. Finally, we used renal endothelial cell line HEK-293 and renal cancer cell lines 786-O and ACHN to explore the effects of SLC14A1 on the biological behaviors of renal cancer cell proliferation, invasion and metastasis using EDU, MTT proliferation assay, Transwell invasion assay and scratch healing assay.
Results
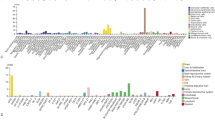
SLC14A1 was lowly expressed in renal cancer tissues and this was further validated by RT-PCR, Western blotting, and immunohistochemistry in our clinical samples. Analysis of KIRC single-cell data suggested that SLC14A1 was mainly expressed in endothelial cells. Survival analysis showed that low levels of SLC14A1 expression were associated with a better clinical prognosis. In biological behavioral studies, we found that upregulation of SLC14A1 expression levels inhibited the proliferation, invasion, and metastatic ability of renal cancer cells.
Conclusion
SLC14A1 plays an important role in the progression of renal cancer and has the potential to become a new biomarker for renal cancer.






Similar content being viewed by others
Data availability
The datasets generated and/or analyzed during the current study are available in the public databases Gene Expression Omnibus database (GEO) (https://www.ncbi.nlm.nih.gov/geo/)and The Cancer Genome Atlas (TCGA) (https://tcga-data.nci.nih.gov/tcga/). Additional data can be requested from the corresponding author.
References
Wang J, Ren Y, Guo X, Cheng H, Ye Y, Qi J, et al. Alterations in enhancer of zeste homolog 2, matrix metalloproteinase-2 and tissue inhibitor of metalloproteinase-2 expression are associated with ex vivo and in vitro bone metastasis in renal cell carcinoma. Mol Med Rep. 2015;11(5):3585–92. https://doi.org/10.3892/mmr.2015.3164.
Capitanio U, Montorsi F. Renal cancer. Lancet. 2016;387(10021):894–906. https://doi.org/10.1016/S0140-6736(15)00046-X.
Kroeger N, Zimmermann U, Burchardt M. One decade of improving palliative care of metastatic renal cell carcinoma by antiangiogenic therapies: time to move toward RCC cure. Int J Cancer. 2015;136(7):1483–4. https://doi.org/10.1002/ijc.29189.
Tan X, Liu Y, Hou J, Cao G. Targeted therapies for renal cell carcinoma in Chinese patients: focus on everolimus. Onco Targets Ther. 2015;8:313–21. https://doi.org/10.2147/OTT.S64660.
Huang D, Ding Y, Luo WM, Bender S, Qian CN, Kort E, et al. Inhibition of MAPK kinase signaling pathways suppressed renal cell carcinoma growth and angiogenesis in vivo. Cancer Res. 2008;68(1):81–8. https://doi.org/10.1158/0008-5472.CAN-07-5311.
Dong B, Zhang J, Chen Y, Chen H, Chen Q, Guo S, et al. Data analysis of renal cancer database of Shanghai Renji Hospital. Chin J Urol. 2008;04:222–5.
Kabaria R, Klaassen Z, Terris MK. Renal cell carcinoma: links and risks. Int J Nephrol Renovasc Dis. 2016;9:45–52. https://doi.org/10.2147/IJNRD.S75916.
Mellemgaard A, Niwa S, Mehl ES, Engholm G, McLaughlin JK, Olsen JH. Risk factors for renal cell carcinoma in Denmark: role of medication and medical history. Int J Epidemiol. 1994;23(5):923–30. https://doi.org/10.1093/ije/23.5.923.
Muscat JE, Hoffmann D, Wynder EL. The epidemiology of renal cell carcinoma. A second look. Cancer. 1995;75(10):2552–7. https://doi.org/10.1002/1097-0142(19950515)75:10%3c2552:aid-cncr2820751023%3e3.0.co;2-1.
Parker A, Lohse C, Cheville J, Leibovich B, Igel T, Blute M. Evaluation of the association of current cigarette smoking and outcome for patients with clear cell renal cell carcinoma. Int J Urol. 2008;15(4):304–8. https://doi.org/10.1111/j.1442-2042.2008.01994.x.
Lindblad P. Epidemiology of renal cell carcinoma. Scand J Surg. 2004;93(2):88–96. https://doi.org/10.1177/145749690409300202.
Pischon T, Lahmann PH, Boeing H, Tjonneland A, Halkjaer J, Overvad K, et al. Body size and risk of renal cell carcinoma in the European Prospective Investigation into Cancer and Nutrition (EPIC). Int J Cancer. 2006;118(3):728–38. https://doi.org/10.1002/ijc.21398.
Jung JE, Lee HG, Cho IH, Chung DH, Yoon SH, Yang YM, et al. STAT3 is a potential modulator of HIF-1-mediated VEGF expression in human renal carcinoma cells. Faseb J. 2005;19(10):1296–8. https://doi.org/10.1096/fj.04-3099fje.
Li M, Wang Y, Song Y, Bu R, Yin BO, Fei X, et al. MicroRNAs in renal cell carcinoma: a systematic review of clinical implications (Review). Oncol Rep. 2015;33(4):1571–8. https://doi.org/10.3892/or.2015.3799.
Wilhelm M, Veltman JA, Olshen AB, Jain AN, Moore DH, Presti JJ, et al. Array-based comparative genomic hybridization for the differential diagnosis of renal cell cancer. Cancer Res. 2002;62(4):957–60.
Avery AK, Beckstead J, Renshaw AA, Corless CL. Use of antibodies to RCC and CD10 in the differential diagnosis of renal neoplasms. Am J Surg Pathol. 2000;24(2):203–10. https://doi.org/10.1097/00000478-200002000-00006.
Hemal AK, Kumar A, Kumar R, Wadhwa P, Seth A, Gupta NP. Laparoscopic versus open radical nephrectomy for large renal tumors: a long-term prospective comparison. J Urol. 2007;177(3):862–6. https://doi.org/10.1016/j.juro.2006.10.053.
Hemal AK, Kumar A. A prospective comparison of laparoscopic and robotic radical nephrectomy for T1–2N0M0 renal cell carcinoma. World J Urol. 2009;27(1):89–94. https://doi.org/10.1007/s00345-008-0321-9.
Ou YC, Kuan YH, Li JR, Raung SL, Wang CC, Hung YY, et al. Induction of apoptosis by luteolin involving akt inactivation in human 786-o renal cell carcinoma cells. Evid Based Complement Alternat Med. 2013. https://doi.org/10.1155/2013/109105.
Yang B. Transport characteristics of urea transporter-B. Subcell Biochem. 2014;73:127–35. https://doi.org/10.1007/978-94-017-9343-8_8.
Hediger MA, Smith CP, You G, Lee WS, Kanai Y, Shayakul C. Structure, regulation and physiological roles of urea transporters. Kidney Int. 1996;49(6):1615–23. https://doi.org/10.1038/ki.1996.235.
Sands JM, Timmer RT, Gunn RB. Urea transporters in kidney and erythrocytes. Am J Physiol. 1997;273(3 Pt 2):F321–39. https://doi.org/10.1152/ajprenal.1997.273.3.F321.
Klein JD, Blount MA, Sands JM. Molecular mechanisms of urea transport in health and disease. Pflugers Arch. 2012;464(6):561–72. https://doi.org/10.1007/s00424-012-1157-0.
Shayakul C, Hediger MA. The SLC14 gene family of urea transporters. Pflugers Arch. 2004;447(5):603–9. https://doi.org/10.1007/s00424-003-1124-x.
Berger UV, Tsukaguchi H, Hediger MA. Distribution of mRNA for the facilitated urea transporter UT3 in the rat nervous system. Anat Embryol (Berl). 1998;197(5):405–14. https://doi.org/10.1007/s004290050152.
Olives B, Neau P, Bailly P, Hediger MA, Rousselet G, Cartron JP, et al. Cloning and functional expression of a urea transporter from human bone marrow cells. J Biol Chem. 1994;269(50):31649–52.
Trinh-Trang-Tan MM, Lasbennes F, Gane P, Roudier N, Ripoche P, Cartron JP, et al. UT-B1 proteins in rat: tissue distribution and regulation by antidiuretic hormone in kidney. Am J Physiol Renal Physiol. 2002;283(5):F912–22. https://doi.org/10.1152/ajprenal.00359.2001.
Tsukaguchi H, Shayakul C, Berger UV, Tokui T, Brown D, Hediger MA. Cloning and characterization of the urea transporter UT3: localization in rat kidney and testis. J Clin Invest. 1997;99(7):1506–15. https://doi.org/10.1172/JCI119313.
Li C, Xue H, Lei Y, Zhu J, Yang B, Gai X. Clinical significance of the reduction of UT-B expression in urothelial carcinoma of the bladder. Pathol Res Pract. 2014;210(12):799–803. https://doi.org/10.1016/j.prp.2014.09.012.
Liu L, Sun Y, Zhao Y, Wang Q, Guo H, Guo R, et al. Urea transport B gene induces melanoma B16 cell death via activation of p53 and mitochondrial apoptosis. Cancer Sci. 2018;109(12):3762–73. https://doi.org/10.1111/cas.13825.
Lau WK, Blute ML, Weaver AL, Torres VE, Zincke H. Matched comparison of radical nephrectomy vs nephron-sparing surgery in patients with unilateral renal cell carcinoma and a normal contralateral kidney. Mayo Clin Proc. 2000;75(12):1236–42. https://doi.org/10.4065/75.12.1236.
Ghoneim IA, Fergany AF. Minimally invasive surgery for renal cell carcinoma. Expert Rev Anticancer Ther. 2009;9(7):989–97. https://doi.org/10.1586/era.09.59.
Jonasch E, Wood CG, Matin SF, Tu SM, Pagliaro LC, Corn PG, et al. Phase II presurgical feasibility study of bevacizumab in untreated patients with metastatic renal cell carcinoma. J Clin Oncol. 2009;27(25):4076–81. https://doi.org/10.1200/JCO.2008.21.3660.
Coppin C, Porzsolt F, Awa A, Kumpf J, Coldman A, Wilt T. Immunotherapy for advanced renal cell cancer. Cochrane Database Syst Rev. 2005;1:D1425. https://doi.org/10.1002/14651858.CD001425.pub2.
Li Y, Hu Z, Ye Z. COX model analysis of three years clinical data of renal cell carcinoma database. J Modern Urol. 2008;04:294–6.
Yang B, Bankir L, Gillespie A, Epstein CJ, Verkman AS. Urea-selective concentrating defect in transgenic mice lacking urea transporter UT-B. J Biol Chem. 2002;277(12):10633–7. https://doi.org/10.1074/jbc.M200207200.
Sands JM, Blount MA. Genes and proteins of urea transporters. Subcell Biochem. 2014;73:45–63. https://doi.org/10.1007/978-94-017-9343-8_4.
Fenton RA, Yang B. Urea transporter knockout mice and their renal phenotypes. Subcell Biochem. 2014;73:137–52. https://doi.org/10.1007/978-94-017-9343-8_9.
Hou R. Study on the mechanism of abnormal expression of SLC14A1 gene in bladder cancer: Jilin University; 2017. 111p.
Ma Z, Li X, Mao Y, Wei C, Huang Z, Li G, et al. Interferon-dependent SLC14A1(+) cancer-associated fibroblasts promote cancer stemness via WNT5A in bladder cancer. Cancer Cell. 2022. https://doi.org/10.1016/j.ccell.2022.11.005.
Dong Z, Ran J, Zhou H, Chen J, Lei T, Wang W, et al. Urea transporter UT-B deletion induces DNA damage and apoptosis in mouse bladder urothelium. PLoS ONE. 2013;8(10):e76952–8. https://doi.org/10.1371/journal.pone.0076952.
Chan T, Wu W, Li W, Shiao M, Shiue Y, Li C. SLC14A1 prevents oncometabolite accumulation and recruits HDAC1 to transrepress oncometabolite genes in urothelial carcinoma. Theranostics. 2020;10(25):11775–93. https://doi.org/10.7150/thno.51655.
Xie B, Liu X, Longjiang T. Prognostic model of renal cell carcinoma and its clinical application. Int J Urol. 2011;06:782–6.
Cui T, Guo L, Guo S, Qin Q, Zhang K, Dong S. Factors affecting the prognosis of renal cell carcinoma. J Modern Urol. 2009;14(01):42–4.
Xu J, Zhang J, Geng T, Wang Y, Zhang A, Zuo L. Multivariate analysis of survival factors of patients with renal cell carcinoma after radical nephrectomy. Chin Oncol Clin. 2009;36(14):784–7.
Patard JJ, Leray E, Cindolo L, Ficarra V, Rodriguez A, De La Taille A, et al. Multi-institutional validation of a symptom-based classification for renal cell carcinoma. J Urol. 2004;172(3):858–62. https://doi.org/10.1097/01.ju.0000135837.64840.55.
Prendergast GC, Smith C, Thomas S, Mandik-Nayak L, Laury-Kleintop L, Metz R, et al. Indoleamine 2,3-dioxygenase pathways of pathogenic inflammation and immune escape in cancer. Cancer Immunol Immunother. 2014;63(7):721–35. https://doi.org/10.1007/s00262-014-1549-4.
Wang Z, Dabrosin C, Yin X, Fuster MM, Arreola A, Rathmell WK, et al. Broad targeting of angiogenesis for cancer prevention and therapy. Semin Cancer Biol. 2015;35:S224–43. https://doi.org/10.1016/j.semcancer.2015.01.001.
Seyfried TN, Huysentruyt LC. On the origin of cancer metastasis. Crit Rev Oncog. 2013;18(1–2):43–73. https://doi.org/10.1615/critrevoncog.v18.i1-2.40.
Sarkar S, Horn G, Moulton K, Oza A, Byler S, Kokolus S, et al. Cancer development, progression, and therapy: an epigenetic overview. Int J Mol Sci. 2013;14(10):21087–113. https://doi.org/10.3390/ijms141021087.
Mishra P, Banerjee D, Ben-Baruch A. Chemokines at the crossroads of tumor-fibroblast interactions that promote malignancy. J Leukoc Biol. 2011;89(1):31–9. https://doi.org/10.1189/jlb.0310182.
Corn PG. The tumor microenvironment in prostate cancer: elucidating molecular pathways for therapy development. Cancer Manag Res. 2012;4:183–93. https://doi.org/10.2147/CMAR.S32839.
Alphonso A, Alahari SK. Stromal cells and integrins: conforming to the needs of the tumor microenvironment. Neoplasia. 2009;11(12):1264–71. https://doi.org/10.1593/neo.91302.
Olumi AF, Grossfeld GD, Hayward SW, Carroll PR, Tlsty TD, Cunha GR. Carcinoma-associated fibroblasts direct tumor progression of initiated human prostatic epithelium. Cancer Res. 1999;59(19):5002–11. https://doi.org/10.1186/bcr138.
Molavi O, Ma Z, Mahmud A, Alshamsan A, Samuel J, Lai R, et al. Polymeric micelles for the solubilization and delivery of STAT3 inhibitor cucurbitacins in solid tumors. Int J Pharm. 2008;347(1–2):118–27. https://doi.org/10.1016/j.ijpharm.2007.06.032.
Lee MH, Kundu JK, Keum YS, Cho YY, Surh YJ, Choi BY. Resveratrol inhibits IL-6-induced transcriptional activity of AR and STAT3 in human prostate cancer LNCaP-FGC Cells. Biomol Ther (Seoul). 2014;22(5):426–30. https://doi.org/10.4062/biomolther.2014.061.
Panni RZ, Sanford DE, Belt BA, Mitchem JB, Worley LA, Goetz BD, et al. Tumor-induced STAT3 activation in monocytic myeloid-derived suppressor cells enhances stemness and mesenchymal properties in human pancreatic cancer. Cancer Immunol Immunother. 2014;63(5):513–28. https://doi.org/10.1007/s00262-014-1527-x.
Wen W, Wu J, Liu L, Tian Y, Buettner R, Hsieh MY, et al. Synergistic anti-tumor effect of combined inhibition of EGFR and JAK/STAT3 pathways in human ovarian cancer. Mol Cancer. 2015;14:100. https://doi.org/10.1186/s12943-015-0366-5.
Li C, Ya G, Tang Z, Su K. MiR-153 targets PRDM2 gene and affects the invasion and migration of bladder cancer through JAK/STAT signaling pathway. Chin J Pathophysiol. 2018;34(01):58–63.
Acknowledgements
We thank the authors of the GSE161573 for their contribution, and we thank GEPIA database and Xiantao Academic database for providing the platform for data analysis.
Funding
This work was funded by “Clinical + X” project of Binzhou Medical University (BY2021LCX07).
Author information
Authors and Affiliations
Contributions
WZQ designed and conducted the whole research. LC uses R language to analyze BLCA data from GEO database and TCGA database. WZQ completed the basic experiments related to this study, WZQ and LC completed the data analysis and drafted the manuscript. WYL revised and finalized the manuscript. All authors approved the submitted version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval and consent to participate
This study was approved by the Ethics Committee of Binzhou Medical University, and all subjects signed the informed consent. Meanwhile, all methods were carried out in accordance with relevant guidelines and regulations or declaration of Helsinki.
Consent for publication
Not applicable.
Additional information
Publisher's Note Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wan, Z., Wang, Y., Li, C. et al. SLC14A1 is a new biomarker in renal cancer. Clin Transl Oncol 25, 2607–2623 (2023). https://doi.org/10.1007/s12094-023-03140-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-023-03140-6