Abstract
Purpose
The de novo lipogenesis has been a longstanding observation in hepatocellular carcinoma (HCC). However, the prognostic value and carcinogenic roles of the enzyme Acetyl-CoA carboxylase alpha (ACACA) in HCC remains unknown.
Methods
The proteins with remarkable prognostic significance were screened out from The Cancer Proteome Atlas Portal (TCPA) database. Furthermore, the expression characteristics and prognostic value of ACACA were evaluated in multiple databases and the local HCC cohort. The loss-of-function assays were performed to uncover the potential roles of ACACA in steering malignant behaviors of HCC cells. The underlying mechanisms were conjectured by bioinformatics and validated in HCC cell lines.
Results
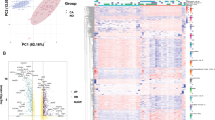
ACACA was identified as a crucial factor of HCC prognosis. Bioinformatics analyses showed that HCC patients with higher expression of ACACA protein or mRNA levels had poor prognosis. Knockdown of ACACA remarkably crippled the proliferation, colony formation, migration, invasion, epithelial−mesenchymal transition (EMT) process of HCC cells and induced the cell cycle arrest. Mechanistically, ACACA might facilitate the malignant phenotypes of HCC through aberrant activation of Wnt/β-catenin signaling pathway. In addition, ACACA expression was associated with the dilute infiltration of immune cells including plasmacytoid DC (pDC) and cytotoxic cells by utilization of relevant database analysis.
Conclusion
ACACA could be a potential biomarker and molecular target for HCC.






Similar content being viewed by others
Availability of data and materials
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary Material.
References
Mao X, Cheung KS, Peng C, Mak LY, Cheng HM, Fung J, et al. Steatosis, HBV-related hepatocellular carcinoma, cirrhosis & HBsAg seroclearance: a systematic review and meta-analysis. Hepatology. 2022. https://doi.org/10.1002/hep.32792.
Zhang J, Wang L, Song Q, Xiao M, Gao J, Cao X, et al. Organoids in recapitulating tumorigenesis driven by risk factors: Current trends and future perspectives. Int J Biol Sci. 2022;18(7):2729–43.
Glišić TM, Perišić MD, Dimitrijevic S, Jurišić V. Doppler assessment of splanchnic arterial flow in patients with liver cirrhosis: correlation with ammonia plasma levels and MELD score. J Clin Ultrasound. 2014;42(5):264–9.
Nikulina D, Terentyev A, Galimzyanov K, Jurisic V. Fifty years of discovery of alpha-fetoprotein as the first tumor marker. Srp Arh Celok Lek. 2015;143(1–2):100–4.
Jurisic V, Radenkovic S, Konjevic G. The Actual role of LDH as tumor marker, biochemical and clinical aspects. Adv Exp Med Biol. 2015;867:115–24.
Kotoh K, Kato M, Kohjima M, Tanaka M, Miyazaki M, Nakamura K, et al. Lactate dehydrogenase production in hepatocytes is increased at an early stage of acute liver failure. Exp Ther Med. 2011;2(2):195–9.
Yau T, Tai D, Chan SL, Huang YH, Choo SP, Hsu C, et al. Systemic treatment of advanced unresectable hepatocellular carcinoma after first-line therapy: expert recommendations from Hong Kong, Singapore, and Taiwan. Liver Cancer. 2022;11(5):426–39.
Sonbol MB, Riaz IB, Naqvi SAA, Almquist DR, Mina S, Almasri J, et al. Systemic therapy and sequencing options in advanced hepatocellular carcinoma: a systematic review and network meta-analysis. JAMA Oncol. 2020;6(12): e204930.
Wang C, Xu C, Sun M, Luo D, Liao DF, Cao D. Acetyl-CoA carboxylase-alpha inhibitor TOFA induces human cancer cell apoptosis. Biochem Biophys Res Commun. 2009;385(3):302–6.
Zhang H, Liu S, Cai Z, Dong W, Ye J, Cai Z, et al. Down-regulation of ACACA suppresses the malignant progression of prostate cancer through inhibiting mitochondrial potential. J Cancer. 2021;12(1):232–43.
Chen L, Duan Y, Wei H, Ning H, Bi C, Zhao Y, et al. Acetyl-CoA carboxylase (ACC) as a therapeutic target for metabolic syndrome and recent developments in ACC1/2 inhibitors. Expert Opin Investig Drugs. 2019;28(10):917–30.
Abu-Elheiga L, Almarza-Ortega DB, Baldini A, Wakil SJ. Human acetyl-CoA carboxylase 2 Molecular cloning, characterization, chromosomal mapping, and evidence for two isoforms. J Biol Chem. 1997;272(16):10669–77.
Ponce-Castaneda MV, Lopez-Casillas F, Kim KH. Acetyl-coenzyme A carboxylase messenger ribonucleic acid metabolism in liver, adipose tissues, and mammary glands during pregnancy and lactation. J Dairy Sci. 1991;74(11):4013–21.
Witters LA, Widmer J, King AN, Fassihi K, Kuhajda F. Identification of human acetyl-CoA carboxylase isozymes in tissue and in breast cancer cells. Int J Biochem. 1994;26(4):589–94.
Ma J, Yan R, Zu X, Cheng JM, Rao K, Liao DF, et al. Aldo-keto reductase family 1 B10 affects fatty acid synthesis by regulating the stability of acetyl-CoA carboxylase-alpha in breast cancer cells. J Biol Chem. 2008;283(6):3418–23.
Milgraum LZ, Witters LA, Pasternack GR, Kuhajda FP. Enzymes of the fatty acid synthesis pathway are highly expressed in in situ breast carcinoma. Clin Cancer Res. 1997;3(11):2115–20.
Lally JSV, Ghoshal S, DePeralta DK, Moaven O, Wei L, Masia R, et al. Inhibition of Acetyl-CoA Carboxylase by Phosphorylation or the Inhibitor ND-654 Suppresses Lipogenesis and Hepatocellular Carcinoma. Cell Metab. 2019;29(1):174-82 e5.
Chen MM, Li J, Wang Y, Akbani R, Lu Y, Mills GB, et al. TCPA v30: an integrative platform to explore the pan-cancer analysis of functional proteomic data. Mol Cell Proteom. 2019;18(8):S15-s25.
Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D, et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004;6(1):1–6.
Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS, et al. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 2017;77(21):e108–10.
Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi B, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19(8):649–58.
Vasaikar SV, Straub P, Wang J, Zhang B. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucl Acids Res. 2018;46(D1):D956–63.
Malta TM, Sokolov A, Gentles AJ, Burzykowski T, Poisson L, Weinstein JN, et al. Machine learning identifies stemness features associated with oncogenic dedifferentiation. Cell. 2018;173(2):338-54 e15.
Sun D, Wang J, Han Y, Dong X, Ge J, Zheng R, et al. TISCH: a comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucl Acids Res. 2021;49(D1):D1420–30.
Jurisic V, Srdic-Rajic T, Konjevic G, Bogdanovic G, Colic M. TNF-α induced apoptosis is accompanied with rapid CD30 and slower CD45 shedding from K-562 cells. J Membr Biol. 2011;239(3):115–22.
Bian S, Ni W, Zhu M, Zhang X, Qiang Y, Zhang J, et al. Flap endonuclease 1 facilitated hepatocellular carcinoma progression by enhancing USP7/MDM2-mediated P53 inactivation. Int J Biol Sci. 2022;18(3):1022–38.
Zhu M, Wu M, Bian S, Song Q, Xiao M, Huang H, et al. DNA primase subunit 1 deteriorated progression of hepatocellular carcinoma by activating AKT/mTOR signaling and UBE2C-mediated P53 ubiquitination. Cell Biosci. 2021;11(1):42.
Bian S, Ni W, Zhu M, Song Q, Zhang J, Ni R, et al. Identification and validation of the N6-Methyladenosine RNA methylation regulator YTHDF1 as a novel prognostic marker and potential target for hepatocellular carcinoma. Front Mol Biosci. 2020;7: 604766.
Ni W, Bian S, Zhu M, Song Q, Zhang J, Xiao M, et al. Identification and validation of ubiquitin-specific proteases as a novel prognostic signature for hepatocellular carcinoma. Front Oncol. 2021;11: 629327.
Zhang G, Ha SA, Kim HK, Yoo J, Kim S, Lee YS, et al. Combined analysis of AFP and HCCR-1 as an useful serological marker for small hepatocellular carcinoma: a prospective cohort study. Dis Markers. 2012;32(4):265–71.
Agopian VG, Harlander-Locke MP, Markovic D, Zarrinpar A, Kaldas FM, Cheng EY, et al. Evaluation of patients with hepatocellular carcinomas that do not produce alpha-fetoprotein. JAMA Surg. 2017;152(1):55–64.
Gallo M, Sapio L, Spina A, Naviglio D, Calogero A, Naviglio S. Lactic dehydrogenase and cancer: an overview. Front Biosci (Landmark Ed). 2015;20(8):1234–49.
Garcia R, Hernandez JM, Caballero MD, Gonzalez M, Galende J, del Canizo MC, et al. Serum lactate dehydrogenase level as a prognostic factor in Hodgkin’s disease. Br J Cancer. 1993;68(6):1227–31.
Scartozzi M, Faloppi L, Bianconi M, Giampieri R, Maccaroni E, Bittoni A, et al. The role of LDH serum levels in predicting global outcome in HCC patients undergoing TACE: implications for clinical management. PLoS ONE. 2012;7(3): e32653.
Faloppi L, Scartozzi M, Bianconi M, Svegliati Baroni G, Toniutto P, Giampieri R, et al. The role of LDH serum levels in predicting global outcome in HCC patients treated with sorafenib: implications for clinical management. BMC Cancer. 2014;14:110.
Chajes V, Cambot M, Moreau K, Lenoir GM, Joulin V. Acetyl-CoA carboxylase alpha is essential to breast cancer cell survival. Cancer Res. 2006;66(10):5287–94.
Mohapatra P, Chandrasekaran N. Wnt/beta-catenin targeting in liver carcinoma through nanotechnology-based drug repurposing: A review. Biomed Pharmacother. 2022;155: 113713.
Xu C, Xu Z, Zhang Y, Evert M, Calvisi DF, Chen X. beta-Catenin signaling in hepatocellular carcinoma. J Clin Invest. 2022. https://doi.org/10.1172/JCI154515.
Wang Z, Li B, Zhou L, Yu S, Su Z, Song J, et al. Prodigiosin inhibits Wnt/beta-catenin signaling and exerts anticancer activity in breast cancer cells. Proc Natl Acad Sci USA. 2016;113(46):13150–5.
Liu J, Sun G, Pan S, Qin M, Ouyang R, Li Z, et al. The Cancer Genome Atlas (TCGA) based m(6)A methylation-related genes predict prognosis in hepatocellular carcinoma. Bioengineered. 2020;11(1):759–68.
Yang Y, Cai J, Yang X, Wang K, Sun K, Yang Z, et al. Dysregulated m6A modification promotes lipogenesis and development of non-alcoholic fatty liver disease and hepatocellular carcinoma. Mol Ther. 2022;30(6):2342–53.
Ding J, Zheng Y, Wang G, Zheng J, Chai D. The performance and perspectives of dendritic cell vaccines modified by immune checkpoint inhibitors or stimulants. Biochim Biophys Acta Rev Cancer. 2022;1877(5): 188763.
Sahin Y. LncRNA H19 is a potential biomarker and correlated with immune infiltration in thyroid carcinoma. Clin Exp Med. 2022. https://doi.org/10.1007/s10238-022-00853-w.
Funding
This study was supported by grants from National Natural Science Foundation (82070622, 82272839, 81702419), the Key Research and Development Plan of Jiangsu Province (BE2020668), and Science project of Nantong City (JCZ21041).
Author information
Authors and Affiliations
Contributions
WZ and JZ designed and supervised this study. YS, ZN, and SX conducted the experiments. YS drafted the manuscript. JZ and WZ performed the revision. XW and SQ performed the bioinformatics and statistics. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The studies involving human participants were reviewed and approved by the Ethics Committee of Affiliated Hospital of Nantong University (2022-L092).
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shen, Y., Wang, X., Ni, Z. et al. Identification of acetyl-CoA carboxylase alpha as a prognostic and targeted candidate for hepatocellular carcinoma. Clin Transl Oncol 25, 2499–2513 (2023). https://doi.org/10.1007/s12094-023-03137-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-023-03137-1