Abstract
Background
Recent studies have reported that cuproptosis, a novel cell death pathway, strongly correlates with mitochondrial metabolism. In addition, the studies reported that cuproptosis plays a role in the development of several cancers and is regulated by protein lipoylation. During cuproptosis, copper binds to the lipoylated proteins and mediates cancer progression. However, the role of cuproptosis in acute myeloid leukemia (AML) patients is yet to be explored.
Methods
This study curated seven cuproptosis-related-genes (CRGs): FDX1, DLAT, PDHB, PDHA1, DLD, LIAS, and LIPT1 to determine cuproptosis modification patterns and the CRGs signature in AML. The CIBERSORT and ssGSEA algorithms were utilized to evaluate the infiltration levels of different immune cell subtypes. A cuproptosis score system based on differentially expressed genes (DEGs) was constructed using the least absolute shrinkage and selection operator (LASSO) regression analysis. The developed cuproptosis score system was validated using two immunotherapy datasets, IMvigor210 and GSE78220.
Results
Three distinct cuproptosis regulation patterns were identified using the Beat AML cohort. The results demonstrated that the three cuproptosis regulation patterns were correlated with various biological pathways and clinical outcomes. Tumor microenvironment (TME) characterization revealed that the identified cuproptosis regulation patterns were consistent with three immune profiles: immune-desert, immune-inflamed, and immune-excluded. The AML patients were grouped into low- and high-score groups based on the cuproptosis score system abstracted from 486 cuproptosis-related DEGs. Patients with lower cuproptosis scores were characterized by longer survival time and attenuated immune infiltration. It was found that lower cuproptosis scores were strongly correlated with lower somatic mutation frequency. Moreover, patients with lower cuproptosis scores presented more favorable immune responses and dual clinical benefits among external validation cohorts.
Conclusions
Cuproptosis phenotypes are significantly correlated with immune microenvironment complexity and variety. Cuprotopsis regulates the response of cancer cells to the immune system. Quantitatively assessing cuproptosis phenotypes in AML improves the understanding and knowledge regarding immune microenvironment characteristics and promotes the development of therapeutic interventions.






Similar content being viewed by others
Data availability
The data could be download at Beat AML dataset (http://www.vizome.org/aml/); AMLCG (GSE106291:https://portal.gdc.cancer.gov/); TCGA-LAML dataset (https://www.cancer.gov/aboutnci/organization/ccg/research/-structuralgenomics/tcga). Immunotherapy datasets were downloaded from IMvigor210 (http://research-pub.gene.com/IMvigor210CoreBiologies/packageVersions/) and GSE78220 (https://www.ncbi.nlm.nih.gov/geo/).
Abbreviations
- AMLCG:
-
AML Cooperative Group
- AML:
-
Acute myeloid leukemia
- CRGs:
-
Cuproptosis-related genes
- DEGs:
-
Differentially expressed genes
- GEO:
-
Gene expression omnibus
- GO:
-
Gene ontology
- GSEA:
-
Gene set enrichment analysis
- GSVA:
-
Gene set variation analysis
- ICB:
-
Immune checkpoint blockade
- ICIs:
-
Immune checkpoint inhibitors
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- LASSO:
-
The least absolute shrinkage and selection operator
- ROC:
-
Receiver operating characteristic
- TME:
-
Tumor microenvironment
- TCGA-LAML:
-
The cancer genome atlas-acute myeloid leukemia
- TCA:
-
The tricarboxylic acid
References
Thomas D, Majeti R. Biology and relevance of human acute myeloid leukemia stem cells. Blood. 2017;129:1577–85. https://doi.org/10.1182/blood-2016-10-696054.
American Cancer Society, <https://cancerstatisticscenter.cancer.org/#!/data-analysis/NewCaseEstimates/compare/DeathEstimates> (2023).
Forte D, Garcia-Fernandez M, Sanchez-Aguilera A, Stavropoulou V, Fielding C, Martin-Perez D, et al. Bone marrow mesenchymal stem cells support acute myeloid leukemia bioenergetics and enhance antioxidant defense and escape from chemotherapy. Cell Metab. 2020;32(829–843):e829. https://doi.org/10.1016/j.cmet.2020.09.001.
Baryawno N, Przybylski D, Kowalczyk MS, Kfoury Y, Severe N, Gustafsson K, et al. A Cellular taxonomy of the bone marrow stroma in homeostasis and leukemia. Cell. 2019;177(1915–1932):e1916. https://doi.org/10.1016/j.cell.2019.04.040.
Stahl M, Goldberg AD. Immune checkpoint inhibitors in acute myeloid leukemia: novel combinations and therapeutic targets. Curr Oncol Rep. 2019;21:37. https://doi.org/10.1007/s11912-019-0781-7.
Kim BE, Nevitt T, Thiele DJ. Mechanisms for copper acquisition, distribution and regulation. Nat Chem Biol. 2008;4:176–85. https://doi.org/10.1038/nchembio.72.
Ge EJ, Bush AI, Casini A, Cobine PA, Cross JR, DeNicola GM, et al. Connecting copper and cancer: from transition metal signalling to metalloplasia. Nat Rev Cancer. 2022;22:102–13. https://doi.org/10.1038/s41568-021-00417-2.
Hunsaker EW, Franz KJ. Emerging opportunities to manipulate metal trafficking for therapeutic benefit. Inorg Chem. 2019;58:13528–45. https://doi.org/10.1021/acs.inorgchem.9b01029.
Bellia F, Grasso GI, Ahmed IMM, Oliveri V, Vecchio G. Carnoquinolines target copper dyshomeostasis, aberrant protein-protein interactions, and oxidative stress. Chemistry. 2020;26:16690–705. https://doi.org/10.1002/chem.202001591.
Singh RP, Jeyaraju DV, Voisin V, Hurren R, Xu C, Hawley JR, et al. Disrupting mitochondrial copper distribution inhibits leukemic stem cell self-renewal. Cell Stem Cell. 2020;26(926–937):e910. https://doi.org/10.1016/j.stem.2020.04.010.
Tsvetkov P, Coy S, Petrova B, Dreishpoon M, Verma A, Abdusamad M, et al. Copper induces cell death by targeting lipoylated TCA cycle proteins. Science. 2022;375:1254–61. https://doi.org/10.1126/science.abf0529.
Chen DS, Mellman I. Elements of cancer immunity and the cancer-immune set point. Nature. 2017;541:321–30. https://doi.org/10.1038/nature21349.
Mariathasan S, Turley SJ, Nickles D, Castiglioni A, Yuen K, Wang Y, et al. TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature. 2018;554:544–8. https://doi.org/10.1038/nature25501.
Hugo W, Zaretsky JM, Sun L, Song C, Moreno BH, Hu-Lieskovan S, et al. Genomic and transcriptomic features of response to Anti-PD-1 therapy in metastatic melanoma. Cell. 2016;165:35–44. https://doi.org/10.1016/j.cell.2016.02.065.
Layton CJ, McMahon PL, Greenleaf WJ. Large-Scale, quantitative protein assays on a high-throughput DNA sequencing chip. Mol Cell. 2019;73(1075–1082):e1074. https://doi.org/10.1016/j.molcel.2019.02.019.
Ni M, Solmonson A, Pan C, Yang C, Li D, Notzon A, et al. Functional assessment of lipoyltransferase-1 deficiency in cells, mice, and humans. Cell Rep. 2019;27(1376–1386):e1376. https://doi.org/10.1016/j.celrep.2019.04.005.
Tsvetkov P, Detappe A, Cai K, Keys HR, Brune Z, Ying W, et al. Mitochondrial metabolism promotes adaptation to proteotoxic stress. Nat Chem Biol. 2019;15:681–9. https://doi.org/10.1038/s41589-019-0291-9.
Sharma S, Bhattarai S, Ara H, Sun G, St Clair DK, Bhuiyan MS, et al. SOD2 deficiency in cardiomyocytes defines defective mitochondrial bioenergetics as a cause of lethal dilated cardiomyopathy. Redox Biol. 2020;37:101740–101740. https://doi.org/10.1016/j.redox.2020.101740.
Saudino G, Ciofi-Baffoni S, Banci L. Protein-interaction affinity gradient drives [4Fe-4S] cluster insertion in human lipoyl synthase. J Am Chem Soc. 2022;144:5713–7. https://doi.org/10.1021/jacs.1c13626.
Rowland EA, Snowden CK, Cristea IM. Protein lipoylation: an evolutionarily conserved metabolic regulator of health and disease. Curr Opin Chem Biol. 2018;42:76–85. https://doi.org/10.1016/j.cbpa.2017.11.003.
Galon J, Bruni D. Approaches to treat immune hot, altered and cold tumours with combination immunotherapies. Nat Rev Drug Discov. 2019;18:197–218. https://doi.org/10.1038/s41573-018-0007-y.
Zeng D, Ye Z, Wu J, Zhou R, Fan X, Wang G, et al. Macrophage correlates with immunophenotype and predicts anti-PD-L1 response of urothelial cancer. Theranostics. 2020;10:7002–14. https://doi.org/10.7150/thno.46176.
Sallman DA, McLemore AF, Aldrich AL, Komrokji RS, McGraw KL, Dhawan A, et al. TP53 mutations in myelodysplastic syndromes and secondary AML confer an immunosuppressive phenotype. Blood. 2020;136:2812–23. https://doi.org/10.1182/blood.2020006158.
Ciurea SO, Chilkulwar A, Saliba RM, Chen J, Rondon G, Patel KP, et al. Prognostic factors influencing survival after allogeneic transplantation for AML/MDS patients with TP53 mutations. Blood. 2018;131:2989–92. https://doi.org/10.1182/blood-2018-02-832360.
Grob T, Al Hinai ASA, Sanders MA, Kavelaars FG, Rijken M, Gradowska PL, et al. Molecular characterization of mutant TP53 acute myeloid leukemia and high-risk myelodysplastic syndrome. Blood. 2022;139:2347–54. https://doi.org/10.1182/blood.2021014472.
Long J, Wang A, Bai Y, Lin J, Yang X, Wang D, et al. Development and validation of a TP53-associated immune prognostic model for hepatocellular carcinoma. EBioMedicine. 2019;42:363–74. https://doi.org/10.1016/j.ebiom.2019.03.022.
Prassek VV, Rothenberg-Thurley M, Sauerland MC, Herold T, Janke H, Ksienzyk B, et al. Genetics of acute myeloid leukemia in the elderly: mutation spectrum and clinical impact in intensively treated patients aged 75 years or older. Haematologica. 2018;103:1853–61. https://doi.org/10.3324/haematol.2018.191536.
Bamopoulos SA, Batcha AMN, Jurinovic V, Rothenberg-Thurley M, Janke H, Ksienzyk B, et al. Clinical presentation and differential splicing of SRSF2, U2AF1 and SF3B1 mutations in patients with acute myeloid leukemia. Leukemia. 2020;34:2621–34. https://doi.org/10.1038/s41375-020-0839-4.
Lindsley RC, Mar BG, Mazzola E, Grauman PV, Shareef S, Allen SL, et al. Acute myeloid leukemia ontogeny is defined by distinct somatic mutations. Blood. 2015;125:1367–76. https://doi.org/10.1182/blood-2014-11-610543.
Grimm J, Jentzsch M, Bill M, Backhaus D, Brauer D, Kupper J, et al. Clinical implications of SRSF2 mutations in AML patients undergoing allogeneic stem cell transplantation. Am J Hematol. 2021;96:1287–94. https://doi.org/10.1002/ajh.26298.
Tyner JW, Tognon CE, Bottomly D, Wilmot B, Kurtz SE, Savage SL, et al. Functional genomic landscape of acute myeloid leukaemia. Nature. 2018;562:526–31. https://doi.org/10.1038/s41586-018-0623-z.
Cancer Genome Atlas Research Network, Ley TJ, Miller C, Ding L, Raphael BJ, Mungall AJ, et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368:2059–74. https://doi.org/10.1056/NEJMoa1301689.
Herold T, Jurinovic V, Batcha AMN, Bamopoulos SA, Rothenberg-Thurley M, Ksienzyk B, et al. A 29-gene and cytogenetic score for the prediction of resistance to induction treatment in acute myeloid leukemia. Haematologica. 2018;103:456–65. https://doi.org/10.3324/haematol.2017.178442.
Wilkerson MD, Hayes DN. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics. 2010;26:1572–3. https://doi.org/10.1093/bioinformatics/btq170.
Yoshihara K, Shahmoradgoli M, Martinez E, Vegesna R, Kim H, Torres-Garcia W, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612. https://doi.org/10.1038/ncomms3612.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12:453–7. https://doi.org/10.1038/nmeth.3337.
Barbie DA, Tamayo P, Boehm JS, Kim SY, Moody SE, Dunn IF, et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature. 2009;462:108–12. https://doi.org/10.1038/nature08460.
Charoentong P, Finotello F, Angelova M, Mayer C, Efremova M, Rieder D, et al. Pan-cancer Immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep. 2017;18:248–62. https://doi.org/10.1016/j.celrep.2016.12.019.
He Y, Jiang Z, Chen C, Wang X. Classification of triple-negative breast cancers based on immunogenomic profiling. J Exp Clin Cancer Res. 2018;37:327. https://doi.org/10.1186/s13046-018-1002-1.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47. https://doi.org/10.1093/nar/gkv007.
Chen H, Boutros PC. VennDiagram: a package for the generation of highly-customizable venn and euler diagrams in R. BMC Bioinformatics. 2011;12:35. https://doi.org/10.1186/1471-2105-12-35.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50. https://doi.org/10.1073/pnas.0506580102.
Funding
This work was supported by the National Natural Science Foundation of China (No. 81700104) and Natural Science Foundation of Guangdong, China (No.2019A043135067).
Author information
Authors and Affiliations
Contributions
NX revised the manuscript and make final approval of the version. DL designed the study, interpreted results and wrote the manuscripts. SL analyzed data and prepared figures. JL, ZH, ZG, HC, and XL performed review and revision of the manuscripts. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
The study was reviewed by the Medical Ethics Committee of Southern Medical University Nanfang Hospital, and confirmed the research range as part of ‘can be exempted from medical ethical review on’ the article 3: for always archived data file records pathological specimens or diagnostic specimen collection or research, and these resources are public resources, or in the way they were unable to contact (directly or through identifier) records the information. Therefore, the application for exemption from medical ethics review of this study was approved.
Human and animal rights
This study dose not involving any human or animal participants directly, since it is a retrospective analysis from open access databases.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12094_2023_3118_MOESM2_ESM.png
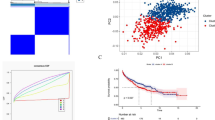
Supplementary file2 Identification of consistent clustering based on the copper cell death-related DEGs in Beat AML and AMLCG cohort. A The result of consistency clustering when k = 3. B, E The cumulative distribution function (CDF) when k = 2–10 (B Beat AML cohort; C AMLCG cohort). C, F The relative change of the area under the CDF curve when k = 2–10 (C Beat AML cohort; F AMLCG cohort). G, H The expression of 4 copper cell death-related DEGs. (G Beat AML cohort, H AMLCG cohort) (PNG 215 KB)
12094_2023_3118_MOESM3_ESM.png
Supplementary file3 The relative abundances of the immune cells or pathways in AMLCG cohort. A The different distribution of 22 TME infiltrating cells in three clusters. B Comparison of the AML immunity among three different clusters. C The different value of ESTIMATE score in three clusters. (****p < 0.0001; ***p < 0.001; **p < 0.01; *p < 0.05) (PNG 1059 KB)
12094_2023_3118_MOESM4_ESM.png
Supplementary file4 The relationship of biological signatures in Beat AML cohort. A Comparison of different biological signatures among three different clusters. (****p < 0.0001; ***p < 0.001; **p < 0.01; *p < 0.05). B The correlation of different biological signatures. (***p < 0.001; **p < 0.01; *p < 0.05) (PNG 351 KB)
12094_2023_3118_MOESM5_ESM.png
Supplementary file5 The development of copper cell death-related model. A Identification of the DEGs among three subclusters. B, C LASSO-cox regression of the genes extracted by univariate cox regression (PNG 283 KB)
12094_2023_3118_MOESM6_ESM.png
Supplementary file6 Kaplan–Meier estimates of BeatAML patients’ OS. A THG1L, B SYPL1, C MRPS36, D ACADS, E KLF9 (PNG 99 KB)
12094_2023_3118_MOESM7_ESM.png
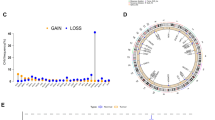
Supplementary file7 Analysis of factors affecting the prognosis of patients with AML. A, B Principal component analysis for the expression of 5 genes extracted from LASSO (A Beat AML cohort; B AMLCG cohort). C Rate of clinical response predicted by 29-gene signature defined by a previous study. D The landscape of Top 20 frequently mutated genes of the patients from TCGA-AML cohort (right bottom: bar plot of the alteration type). E, F Forest plot summary of the multivariable Cox analysis of the copper score (risk score) and clinicopathological characteristics (*p < 0.05; **p < 0.01; ***p < 0.001) (PNG 630 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Luo, D., Liu, S., Luo, J. et al. Characterization of cuproptosis identified immune microenvironment and prognosis in acute myeloid leukemia. Clin Transl Oncol 25, 2393–2407 (2023). https://doi.org/10.1007/s12094-023-03118-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-023-03118-4