Abstract
Background
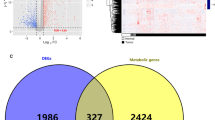
The importance of metabolism-related alterations in the development of gastric cancer (GC) is increasingly recognized. The present study aimed to identify metabolism-related genes to facilitate prognosis of GC patients.
Methods
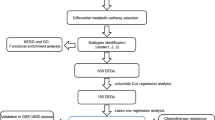
Gene expression datasets and clinical information of GC patients were downloaded from TCGA and GEO databases. We scored the enrichment of human metabolism-related pathways (n = 86) in GC samples by GSV, constructed prognostic risk models using LASSO algorithm and multivariate Cox regression analysis, combined with clinical information to construct a nomogram, and finally cis score algorithm to analyze the abundance of immune-related cells in different subtypes. We used Weka software to screen for prognosis-related marker genes and finally validated the expression of the selected genes in clinical cancer patient tissues.
Results
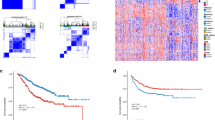
We identified that two GC metabolism-related signatures were strongly associated with OS and the levels of immune cell infiltration. Moreover, a survival prediction model for GC was established based on six GC metabolism-related genes. Time-dependent ROC analysis showed good stability of the risk prediction scoring model. The model was successfully validated in an independent ACRG cohort, and the expression trends of key genes were also verified in the GC tissues of patients. DLX1, LTBP2, FGFR1 and MMP2 were highly expressed in the cluster with poorer prognosis while SLC13A2 and SLCO1B3 were highly expressed in the cluster with better prognosis.
Conclusions
We identified a risk predictive score model based on six metabolism-related genes related to survival, which may serve as prognostic indicators and potential therapeutic targets for GC.







Similar content being viewed by others
Data availability statement
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding author.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49. https://doi.org/10.3322/caac.21660.
Zong L, Abe M, Seto Y, Ji J. The challenge of screening for early gastric cancer in China. Lancet. 2016;388(10060):2606. https://doi.org/10.1016/S0140-6736(16)32226-7.
Chen YC, Fang WL, Wang RF, Liu CA, Yang MH, Lo SS, et al. Clinicopathological variation of Lauren classification in gastric cancer. Pathol Oncol Res. 2016;22(1):197–202. https://doi.org/10.1007/s12253-015-9996-6.
Hu B, El HN, Sittler S, Lammert N, Barnes R, Meloni-Ehrig A. Gastric cancer: classification, histology and application of molecular pathology. J Gastrointest Oncol. 2012;3(3):251–61. https://doi.org/10.3978/j.issn.2078-6891.2012.021.
Chia NY, Tan P. Molecular classification of gastric cancer. Ann Oncol. 2016;27(5):763–9. https://doi.org/10.1093/annonc/mdw040.
Park JH, Pyun WY, Park HW. Cancer metabolism: phenotype, signaling and therapeutic targets. Cells Basel. 2020. https://doi.org/10.3390/cells9102308.
Jurisic V, Radenkovic S, Konjevic G. The actual role of LDH as tumor marker, biochemical and clinical aspects. Adv Exp Med Biol. 2015;867:115–24. https://doi.org/10.1007/978-94-017-7215-0_8.
Yang GX, Li X, Snyder M. Investigating metabolite-protein interactions: an overview of available techniques. Methods. 2012;57(4):459–66. https://doi.org/10.1016/j.ymeth.2012.06.013.
Bhattacharyya A, Chattopadhyay R, Mitra S, Crowe SE. Oxidative stress: an essential factor in the pathogenesis of gastrointestinal mucosal diseases. Physiol Rev. 2014;94(2):329–54. https://doi.org/10.1152/physrev.00040.2012.
Xu LB, Zhang HH, Shi MM, Huang ZX, Zhang WT, Chen XD, et al. Metabolic syndrome-related sarcopenia is associated with worse prognosis in patients with gastric cancer: a prospective study. Eur J Surg Oncol. 2020;46(12):2262–9. https://doi.org/10.1016/j.ejso.2020.07.032.
Garber K. Energy deregulation: licensing tumors to grow. Science. 2006;312(5777):1158–9. https://doi.org/10.1126/science.312.5777.1158.
Bourlieu C, Paboeuf G, Chever S, Pezennec S, Cavalier JF, Guyomarc’H F, et al. Adsorption of gastric lipase onto multicomponent model lipid monolayers with phase separation. Colloids Surf B Biointerfaces. 2016;143:97–106. https://doi.org/10.1016/j.colsurfb.2016.03.032.
Yuan LW, Yamashita H, Seto Y. Glucose metabolism in gastric cancer: The cutting-edge. World J Gastroenterol. 2016;22(6):2046–59. https://doi.org/10.3748/wjg.v22.i6.2046.
Ren Q, Zhu P, Zhang H, Ye T, Liu D, Gong Z, et al. Identification and validation of stromal-tumor microenvironment-based subtypes tightly associated with PD-1/PD-L1 immunotherapy and outcomes in patients with gastric cancer. Cancer Cell Int. 2020;20:92. https://doi.org/10.1186/s12935-020-01173-3.
Tahara T, Arisawa T. DNA methylation as a molecular biomarker in gastric cancer. Epigenomics Uk. 2015;7(3):475–86. https://doi.org/10.2217/epi.15.4.
Obradovic J, Todosijevic J, Jurisic V. Application of the conventional and novel methods in testing EGFR variants for NSCLC patients in the last 10 years through different regions: a systematic review. Mol Biol Rep. 2021;48(4):3593–604. https://doi.org/10.1007/s11033-021-06379-w.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12(5):453–7. https://doi.org/10.1038/nmeth.3337.
Xu YH, Li ZL, Qiu SF. IFN-gamma induces gastric cancer cell proliferation and metastasis through upregulation of integrin beta3-mediated NF-kappaB signaling. Transl Oncol. 2018;11(1):182–92. https://doi.org/10.1016/j.tranon.2017.11.008.
Chen D, Liu Z, Liu W, Fu M, Jiang W, Xu S, et al. Predicting postoperative peritoneal metastasis in gastric cancer with serosal invasion using a collagen nomogram. Nat Commun. 2021;12(1):179. https://doi.org/10.1038/s41467-020-20429-0.
Ganapathy-Kanniappan S, Geschwind JF. Tumor glycolysis as a target for cancer therapy: progress and prospects. Mol Cancer. 2013;12:152. https://doi.org/10.1186/1476-4598-12-152.
Ghanemi A, Melouane A, Yoshioka M, St-Amand J. Secreted protein acidic and rich in cysteine and bioenergetics: extracellular matrix, adipocytes remodeling and skeletal muscle metabolism. Int J Biochem Cell Biol. 2019;117: 105627. https://doi.org/10.1016/j.biocel.2019.105627.
Pickup MW, Mouw JK, Weaver VM. The extracellular matrix modulates the hallmarks of cancer. Embo Rep. 2014;15(12):1243–53. https://doi.org/10.15252/embr.201439246.
Moreira AM, Pereira J, Melo S, Fernandes MS, Carneiro P, Seruca R, et al. The extracellular matrix: an accomplice in gastric cancer development and progression. Cells Basel. 2020. https://doi.org/10.3390/cells9020394.
Cooper J, Giancotti FG. Integrin signaling in cancer: mechanotransduction, stemness, epithelial plasticity, and therapeutic resistance. Cancer Cell. 2019;35(3):347–67. https://doi.org/10.1016/j.ccell.2019.01.007.
Kawashima A, Tsugawa S, Boku A, Kobayashi M, Minamoto T, Nakanishi I, et al. Expression of alphav integrin family in gastric carcinomas: increased alphavbeta6 is associated with lymph node metastasis. Pathol Res Pract. 2003;199(2):57–64. https://doi.org/10.1078/0344-0338-00355.
Lian PL, Liu Z, Yang GY, Zhao R, Zhang ZY, Chen YG, et al. Integrin alphavbeta6 and matrix metalloproteinase 9 correlate with survival in gastric cancer. World J Gastroenterol. 2016;22(14):3852–9. https://doi.org/10.3748/wjg.v22.i14.3852.
Jiang X, Wu M, Xu X, Zhang L, Huang Y, Xu Z, et al. COL12A1, a novel potential prognostic factor and therapeutic target in gastric cancer. Mol Med Rep. 2019;20(4):3103–12. https://doi.org/10.3892/mmr.2019.10548.
Zhang QN, Zhu HL, Xia MT, Liao J, Huang XT, Xiao JW, et al. A panel of collagen genes are associated with prognosis of patients with gastric cancer and regulated by microRNA-29c-3p: an integrated bioinformatics analysis and experimental validation. Cancer Manag Res. 2019;11:4757–72. https://doi.org/10.2147/CMAR.S198331.
Zhao Y, Zhou T, Li A, Yao H, He F, Wang L, et al. A potential role of collagens expression in distinguishing between premalignant and malignant lesions in stomach. Anat Rec (Hoboken). 2009;292(5):692–700. https://doi.org/10.1002/ar.20874.
Li T, Huang H, Shi G, Zhao L, Li T, Zhang Z, et al. TGF-beta1-SOX9 axis-inducible COL10A1 promotes invasion and metastasis in gastric cancer via epithelial-to-mesenchymal transition. Cell Death Dis. 2018;9(9):849. https://doi.org/10.1038/s41419-018-0877-2.
Ding B, Lou W, Xu L, Li R, Fan W. Analysis the prognostic values of solute carrier (SLC) family 39 genes in gastric cancer. Am J Transl Res. 2019;11(1):486–98.
Sato H, Ishihara S, Kawashima K, Moriyama N, Suetsugu H, Kazumori H, et al. Expression of peroxisome proliferator-activated receptor (PPAR)gamma in gastric cancer and inhibitory effects of PPARgamma agonists. Br J Cancer. 2000;83(10):1394–400. https://doi.org/10.1054/bjoc.2000.1457.
Wagner N, Wagner KD. PPAR beta/delta and the hallmarks of cancer. Cells Basel. 2020. https://doi.org/10.3390/cells9051133.
Kim KW, Kim N, Choi Y, Kim WS, Yoon H, Shin CM, et al. Different effects of p53 protein overexpression on the survival of gastric cancer patients according to Lauren histologic classification: a retrospective study. Gastric Cancer. 2021;24(4):844–57. https://doi.org/10.1007/s10120-021-01163-y.
Han Y, Zhang Y, Jia T, Sun Y. Molecular mechanism underlying the tumor-promoting functions of carcinoma-associated fibroblasts. Tumour Biol. 2015;36(3):1385–94. https://doi.org/10.1007/s13277-015-3230-8.
Noto CN, Hoft SG, DiPaolo RJ. Mast cells as important regulators in autoimmunity and cancer development. Front Cell Dev Biol. 2021;9: 752350. https://doi.org/10.3389/fcell.2021.752350.
Du Y, Wei Y. Therapeutic potential of natural killer cells in gastric cancer. Front Immunol. 2018;9:3095. https://doi.org/10.3389/fimmu.2018.03095.
Wei M, Shen D, Mulmi SS, Liu J, Zhang J, Yin Y. The progress of T cell immunity related to prognosis in gastric cancer. Biomed Res Int. 2018;2018:3201940. https://doi.org/10.1155/2018/3201940.
Goel S, Bhatia V, Kundu S, Biswas T, Carskadon S, Gupta N, et al. Transcriptional network involving ERG and AR orchestrates Distal-less homeobox-1 mediated prostate cancer progression. Nat Commun. 2021;12(1):5325. https://doi.org/10.1038/s41467-021-25623-2.
Chan DW, Hui WW, Wang JJ, Yung MM, Hui LM, Qin Y, et al. DLX1 acts as a crucial target of FOXM1 to promote ovarian cancer aggressiveness by enhancing TGF-beta/SMAD4 signaling. Oncogene. 2017;36(10):1404–16. https://doi.org/10.1038/onc.2016.307.
Wang J, Liang WJ, Min GT, Wang HP, Chen W, Yao N. LTBP2 promotes the migration and invasion of gastric cancer cells and predicts poor outcome of patients with gastric cancer. Int J Oncol. 2018;52(6):1886–98. https://doi.org/10.3892/ijo.2018.4356.
Chen Q, Zhu M, Xie J, Dong Z, Khushafah F, Yun D, et al. Design and synthesis of novel nordihydroguaiaretic acid (NDGA) analogues as potential FGFR1 kinase inhibitors with anti-gastric activity and chemosensitizing effect. Front Pharmacol. 2020;11: 518068. https://doi.org/10.3389/fphar.2020.518068.
Liu D, Kang H, Gao M, Jin L, Zhang F, Chen D, et al. Exosome-transmitted circ_MMP2 promotes hepatocellular carcinoma metastasis by upregulating MMP2. Mol Oncol. 2020;14(6):1365–80. https://doi.org/10.1002/1878-0261.12637.
Gobin E, Bagwell K, Wagner J, Mysona D, Sandirasegarane S, Smith N, et al. A pan-cancer perspective of matrix metalloproteases (MMP) gene expression profile and their diagnostic/prognostic potential. BMC Cancer. 2019;19(1):581. https://doi.org/10.1186/s12885-019-5768-0.
Han L, Sheng B, Zeng Q, Yao W, Jiang Q. Correlation between MMP2 expression in lung cancer tissues and clinical parameters: a retrospective clinical analysis. Bmc Pulm Med. 2020;20(1):283. https://doi.org/10.1186/s12890-020-01317-1.
Wang J, Cai H, Xia Y, Wang S, Xing L, Chen C, et al. Bufalin inhibits gastric cancer invasion and metastasis by down-regulating Wnt/ASCL2 expression. Oncotarget. 2018;9(34):23320–33. https://doi.org/10.18632/oncotarget.24157.
Sun L, Zhang Y, Lou J. ARHGAP9 siRNA inhibits gastric cancer cell proliferation and EMT via inactivating Akt, p38 signaling and inhibiting MMP2 and MMP9. Int J Clin Exp Pathol. 2017;10(12):11979–85.
Bergeron MJ, Clemencon B, Hediger MA, Markovich D. SLC13 family of Na(+)-coupled di- and tri-carboxylate/sulfate transporters. Mol Aspects Med. 2013;34(2–3):299–312. https://doi.org/10.1016/j.mam.2012.12.001.
Tang T, Wang G, Liu S, Zhang Z, Liu C, Li F, et al. Highly expressed SLCO1B3 inhibits the occurrence and development of breast cancer and can be used as a clinical indicator of prognosis. Sci Rep. 2021;11(1):631. https://doi.org/10.1038/s41598-020-80152-0.
Acknowledgements
Not applicable.
Funding
This study was supported by National Natural Science Foundation of China (No.81973782, No.81704031), Science and Technology Planning Project of Jiangsu Province, China (No. BK20211392), Postgraduate Research & Practice Innovation Program of Jiangsu Province (SJCX22_0749, SJCX21_0692, SJCX21-0740), The Nanjing Medical Science and Technique Development Foundation (ZKX19022), Jiangsu Provincial High level health talent “six one project” (LGY2019005).
Author information
Authors and Affiliations
Contributions
XXZ, XC, YQL, JW, MLC, RJZ, XTX, QMS and TYX: jointly designed the study and drafted the manuscript. JYL: was responsible for the collection of clinical specimens. JW, QMS, and TYX: supervised the study. All the authors contributed to the data collection, analysis and interpretation, manuscript writing and revision. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no potential conflicts of interest.
Ethics approval
This retrospective observational study involving human participants was in accordance with the ethical standards of the institutional research committee. The Ethics Committee of the Affiliated Hospital of Nanjing University of Traditional Chinese Medicine (Jiangsu Provincial Hospital of Traditional Chinese Medicine) approved this study (No. 20221NL18702).
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, X., Chen, X., Liu, J. et al. A novel metabolism-related prognostic gene development and validation in gastric cancer. Clin Transl Oncol 25, 447–459 (2023). https://doi.org/10.1007/s12094-022-02958-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-022-02958-w