Abstract
Background
Neonatal tumors represent an extremely rare and heterogeneous disease with an unknown etiology. Due to its early onset, it has been proposed that genetic factors could play a critical role; however, germline genetic analysis is not usually performed in neonatal cancer patients
Patients and methods
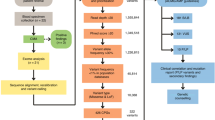
To improve the identification of cancer genetic predisposition syndromes, we retrospectively review clinical characteristics in 45 patients with confirmed tumor diagnosis before 28 days of age, and we carried out germline genetic analysis in 20 patients using next-generation sequencing and directed sequencing.
Results
The genetic studies did not find any germline mutation except patients diagnosed with bilateral retinoblastoma who harbored RB1 germline mutations.
Conclusions
Our results suggest that genetic factors have almost no higher impact in most neonatal tumors. However, since the heterogeneity of the tumors and the small sample size analyzed, we recommend complementary and centralized germline studies to discard the early onset as an additional criterion to take into account to improve the identification of cancer genetic predisposition syndromes in neonates.


Similar content being viewed by others
References
Isaacs H Jr. Congenital and neonatal malignant tumors. A 28-year experience at Children’s Hospital of Los Angeles. Am J Pediatr Hematol/oncol. 1987;9(2):121–9.
Moore SW. Neonatal tumours. Pediatr Surg Int. 2013;29(12):1217–29. https://doi.org/10.1007/s00383-013-3424-3.
Moore SW, Satge D, Sasco AJ, Zimmermann A, Plaschkes J. The epidemiology of neonatal tumours. Report of an international working group. Pediatr Surg Int. 2003;19(7):509–19. https://doi.org/10.1007/s00383-003-1048-8.
Gurney JG, Ross JA, Wall DA, Bleyer WA, Severson RK, Robison LL. Infant cancer in the US: histology-specific incidence and trends, 1973 to 1992. J Pediatr Hematol Oncol. 1997;19(5):428–32.
Parkes SE, Muir KR, Southern L, Cameron AH, Darbyshire PJ, Stevens MC. Neonatal tumours: a thirty-year population-based study. Med Pediatr Oncol. 1994;22(5):309–17.
Vasilatou-Kosmidis H. Cancer in neonates and infants. Med Pediatr Oncol. 2003;41(1):7–9. https://doi.org/10.1002/mpo.10153.
Lichtenstein P, Holm NV, Verkasalo PK, Iliadou A, Kaprio J, Koskenvuo M, et al. Environmental and heritable factors in the causation of cancer–analyses of cohorts of twins from Sweden, Denmark, and Finland. N Engl J Med. 2000;343(2):78–85. https://doi.org/10.1056/NEJM200007133430201.
Orbach D, Sarnacki S, Brisse HJ, Gauthier-Villars M, Jarreau PH, Tsatsaris V, et al. Neonatal cancer. Lancet Oncol. 2013;14(13):e609–20. https://doi.org/10.1016/S1470-2045(13)70236-5.
Grobner SN, Worst BC, Weischenfeldt J, Buchhalter I, Kleinheinz K, Rudneva VA, et al. The landscape of genomic alterations across childhood cancers. Nature. 2018;555(7696):321–7. https://doi.org/10.1038/nature25480.
Zhang J, Walsh MF, Wu G, Edmonson MN, Gruber TA, Easton J, et al. Germline mutations in predisposition genes in pediatric cancer. N Engl J Med. 2015;373(24):2336–46. https://doi.org/10.1056/NEJMoa1508054.
Chang W, Brohl AS, Patidar R, Sindiri S, Shern JF, Wei JS, et al. MultiDimensional ClinOmics for precision therapy of children and adolescent young adults with relapsed and refractory cancer: a report from the center for cancer research. Clin Cancer Res. 2016;22(15):3810–20. https://doi.org/10.1158/1078-0432.CCR-15-2717.
Parsons DW, Roy A, Yang Y, Wang T, Scollon S, Bergstrom K, et al. Diagnostic yield of clinical tumor and germline whole-exome sequencing for children with solid tumors. JAMA Oncol. 2016. https://doi.org/10.1001/jamaoncol.2015.5699.
Jongmans MC, Loeffen JL, Waanders E, Hoogerbrugge PM, Ligtenberg MJ, Kuiper RP, et al. Recognition of genetic predisposition in pediatric cancer patients: an easy-to-use selection tool. Eur J Med Genet. 2016;59(3):116–25. https://doi.org/10.1016/j.ejmg.2016.01.008.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9(4):357–9. https://doi.org/10.1038/nmeth.1923.
Broad Institute G. “Picard Toolkit.”. 2019. http://broadinstitute.github.io/picard/.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43(5):491–8. https://doi.org/10.1038/ng.806.
Cingolani P, Platts A, le Wang L, Coon M, Nguyen T, Wang L, et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly. 2012;6(2):80–92. https://doi.org/10.4161/fly.19695.
Pedersen BS, Layer RM, Quinlan AR. Vcfanno: fast, flexible annotation of genetic variants. Genome Biol. 2016;17(1):118. https://doi.org/10.1186/s13059-016-0973-5.
Peris-Bonet RPE, Ríos I, Sayas N, Valero S. Cáncer Infantil en España. Estadísticas 1980–2015. Registro Español de Tumores Infantiles (RETI-SEHOP). Valencia: Univesitat de València; 2016.
Lopez Almaraz R, Villafruela Alvarez C, Rodriguez Luis J, Domenech Martinez E. Neonatal neoplasms: a single-centre experience. An Pediatr. 2006;65(6):529–35.
Birch JM, Blair V. The epidemiology of infant cancers. Br J Cancer Suppl. 1992;18:S2–4.
Borch K, Jacobsen T, Olsen JH, Hirsch FR, Hertz H. Neonatal cancer in Denmark 1943-1985. Ugeskr Laeger. 1994;156(2):176–9.
Isaacs H Jr. Fetal and neonatal neuroblastoma: retrospective review of 271 cases. Fetal Pediatr Pathol. 2007;26(4):177–84. https://doi.org/10.1080/15513810701696890.
Minard-Colin V, Orbach D, Martelli H, Bodemer C, Oberlin O. Soft tissue tumors in neonates. Arch Pediatr. 2009;16(7):1039–48. https://doi.org/10.1016/j.arcped.2009.03.005.
Veal GJ, Boddy AV. Chemotherapy in newborns and preterm babies. Semin Fetal Neonatal Med. 2012;17(4):243–8. https://doi.org/10.1016/j.siny.2012.03.002.
Littman P, D’Angio GJ. Radiation therapy in the neonate. Am J Pediatr Hematol Oncol. 1981;3(3):279–85.
Caldwell KJ, De La Cuesta E, Morin C, Pappo A, Helmig S. A newborn with a large NTRK fusion positive infantile fibrosarcoma successfully treated with larotrectinib. Pediatr Blood Cancer. 2020. https://doi.org/10.1002/pbc.28330.
Dimaras H, Corson TW, Cobrinik D, White A, Zhao J, Munier FL, et al. Retinoblastoma. Nature reviews Disease primers. 2015;1:15021. https://doi.org/10.1038/nrdp.2015.21.
Porter CC. Germ line mutations associated with leukemias. Hematology American Society of Hematology Education Program. 2016;2016(1):302–8. https://doi.org/10.1182/asheducation-2016.1.302.
Aretz S, Koch A, Uhlhaas S, Friedl W, Propping P, von Schweinitz D, et al. Should children at risk for familial adenomatous polyposis be screened for hepatoblastoma and children with apparently sporadic hepatoblastoma be screened for APC germline mutations? Pediatr Blood Cancer. 2006;47(6):811–8. https://doi.org/10.1002/pbc.20698.
Acknowledgements
This study was supported by the CRIS Cancer Foundation (http://criscancer.org).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to disclose.
Ethical approval
Research was approved by the ethics committee of La Paz University Hospital.
Informed consent
Informed consent was obtained from all individual participants included on the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Escudero, A., Ruz-Caracuel, B., Bueno, D. et al. Genetic predisposition to fetal and neonatal cancer. Clin Transl Oncol 23, 1179–1184 (2021). https://doi.org/10.1007/s12094-020-02508-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-020-02508-2