Abstract
Background
About 5–10% of incidences of breast cancers have been reported as a result of germline mutations of BRCA genes. However, the mutational spectrum of BRCA1 and BRCA2 genes among breast cancer Saudi women patients is inadequate at present. Therefore, the present study aimed to report the specific germinal mutation of BRCA1 and BRCA2 in the entire coding regions, to investigate the prevalence rate of BRCA1 & BRCA2 mutations among Saudi women and the effect of these mutations, both benign and malignant tumors.
Methodology
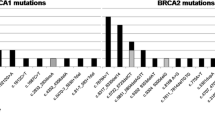
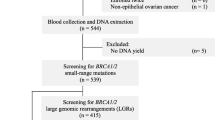
A total of 270 tissue samples of benign and malignant breast tumors were collected from Saudi women patients, Riyadh, Saudi Arabia. Examination of BRCA1 and BRCA2 germline mutations was performed using heteroduplex DNA analysis (HDA) or single-stranded conformation analysis (SSCA). 177 breast cancer women with malignant tumors and 93 with benign tumors were enrolled in the study. A total of 62 out of 177 breast cancer patients carried a BRCA1 or BRCA2 mutation (54 BRCA1 and 8 BRCA2). The analysis was done using the Sanger sequence assay.
Results
Point and frameshift mutations through the entire coding area of the two genes indicated that all the mutations were germline alterations and of early-onset breast cancers. The mean ages of diagnosed breast cancer women for BRCA1 and BRCA2 mutation carriers were 36.3 (± 3.5) and 37.9 (± 3.7) years, whereas that of benign control was 35(± 2.5) years.
Conclusion
Point and frameshift mutations across the entire coding region of BRCA1 and BRCA2 are responsible for many breast cancers cases.




Similar content being viewed by others
Change history
20 May 2021
A Correction to this paper has been published: https://doi.org/10.1007/s12094-021-02643-4
References
Ferlay J, et al. Global cancer observatory: cancer today. Lyon, France: International Agency for Research on Cancer. 2018.
Miki Y, et al. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science. 1994;266:66–71.
Abulkhair O, et al. Prevalence of BRCA1 and BRCA2 mutations among high-risk Saudi patients with breast cancer. J Global Oncol. 2018;3:1–9.
ALmutlaq BA, et al. Options in breast reconstruction and plastic surgery in regard to surgeon perceptions and patients acceptance in Saudi Arabia. J Mod Plast Surg. 2017;7:50–64.
ALmutlaq BA, et al. The influence of women's positioning in breast measurements for subsequent breast plastic surgery. Am J Clin Med Res. 2018;6:10–14.
Almutlaq BA, et al. Breast cancer in Saudi Arabia and its possible risk factors. J Cancer Policy. 2017;12:83–9.
Alrashidi AG, et al. Knowledge and perceptions of common breast cancer risk factors in Northern Saudi Arabia. Asian Pac J Cancer Prev. 2017;18:2755–62.
Othman A, et al. Knowledge, attitudes and practices of breast cancer screening among women in Jordan. Health Care Women Int. 2015;36:578–92.
Fang M, et al. Characterization of mutations in BRCA1/2 and the relationship with clinic-pathological features of breast cancer in a hereditarily high-risk sample of chinese population. Oncol Lett. 2018;15:3068–74.
Larsen MJ, et al. Hereditary breast cancer: clinical, pathological and molecular characteristics. Breast Cancer: Basic Clin Res. 2014;8:145–55.
Blackwood MA, Barbara LW. BRCA1 and BRCA2: from molecular genetics to clinical medicine. J Clin Oncol. 1998;16:1969–77
Torres D, et al. Prevalence and penetration of BRCA1 and BRCA2 germline mutationss in Colombian breast cancer patients. Sci Rep. 2018:7:1–9.
Ottini L, et al. BRCA1 and BRCA2 mutations status and tumor characteristics in male breast cancer: a population-based study in Italy. Cancer Res. 2003;63:342–7.
Ediriweera DS, et al. Mapping the risk of snakebite in Sri Lanka-a national survey with geospatial analysis. PLoS Negl Trop Dis. 2016;10:1–14.
Kumar D, et al. Quantification of DNA extracted from formalin fixed paraffin-embedded tissue comparison of three techniques: effect on PCR efficiency. J Clin Diagn Res. 2016;10:BC01–3.
Alcaide M, et al. A novel multiplex droplet digital pcr assay to identify and quantify KRAS mutations in clinical specimens. J Mol Diagn. 2019;21:214–27.
Nataraj AJ, et al. Single‐strand conformation polymorphism and heteroduplex analysis for gel‐based mutation detection. Electrophoresis. 1999;20:1177–85.
Suvarna KS, Christopher L, John DB, editors. Bancroft's theory and practice of histological techniques E-Book. Elsevier Health Sciences; 2018. p. 213–20.
Tian H, Lawrence CB, James PL. Rapid detection of deletion, insertion, and substitution mutations via heteroduplex analysis using capillary-and microchip-based electrophoresis. Genome Res. 2000;10:1403–13.
Lancaster JM, et al. BRCA2 mutations in primary breast and ovarian cancers. Nat Genet. 1996;13:238–40.
Apessos A, et al. Comprehensive BRCA mutation analysis in the Greek population. Experience from a single clinical diagnostic center. Cancer Genet. 2018;220:1–12.
Liu Y, et al. BRCA1/BRCA2 mutations in Japanese women with ductal carcinoma in situ. Mol Genet Genomic Med. 2019;7:e493.
Bayraktar S, et al. Predictive factors for BRCA1/BRCA2 mutations in women with ductal carcinoma in situ. Cancer. 2012;118:1515–22.
Abusanad A. BRCA testing dichotomy in Saudi Arabia. J Global Oncol. 2019;5:1–2.
Comen E. et al. Relative contributions of BRCA1 and BRCA2 mutations to “triple-negative” breast cancer in Ashkenazi Women. Breast Cancer Res Treat. 2011;129:185–90.
Na B, et al. Therapeutic targeting of BRCA1 and TP53 mutant breast cancer through mutant p53 reactivation. NPJ Breast Cancer. 2019;5:1–10.
Lara K, et al. BRCA1 and BRCA2 mutations in breast cancer patients from Venezuela. Biol Res. 2012;45:117–30.
Suliman et al. Role of human PTEN and TP53 sequence mutations in the etiology of breast cancer in Saudi Patients. Pak J Biol Sci. 2020;9:321–30.
Acknowledgements
This research project was supported by a grant from the Deanship of Scientific Research, Princess Nourah Bint Abdul Rahman University (233-ص-38).
Author information
Authors and Affiliations
Contributions
ASA: participated in the coordination of the study and acquisition and revised the manuscript. RSS: participated in the coordination of the practical work of the study and acquisition, collected the data and revised the manuscript. WSA: carried out the design of the study, collected the data, wrote, revised, and edited the manuscript text and was corresponding author for this study. All authors made a significant contribution to the work reported, read, revised and agreed to the final manuscript before submission.
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no competing interests.
Ethical approval
The ethical standards of the institutional research committee provided a framework for performing procedures on human participants in agreement with the 1964 Declaration of Helsinki and subsequent ethical frameworks. The Deanship of Scientific Research, Princess Nourah Bint Abdul Rahman University approved the study.
Informed consent
All human participants in the study provided informed consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original article was revised to correct the grant number in Acknowledgements.
Rights and permissions
About this article
Cite this article
Algebaly, A.S., Suliman, R.S. & Al-Qahtani, W.S. Comprehensive study for BRCA1 and BRCA2 entire coding regions in breast cancer. Clin Transl Oncol 23, 74–81 (2021). https://doi.org/10.1007/s12094-020-02385-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-020-02385-9