Abstract
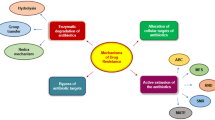
The treatment of bacterial infections is becoming increasingly ineffective due to rapid mutation which leads to antibiotic resistant and resistant bacteria become more prevalent. As a result the existing antibiotics are gradually obsolete and again new drugs are needed to be designed for the same threat. However, the prediction of evolutionary processes/antibiotic resistance is uncertain. Still, the understanding of mode of evolution of resistance in bacteria is a determining step in the preclinical development of new antibiotics, because drug developers assess the risk of resistance arising against a drug during preclinical development. Multidrug efflux pump systems play an important role for making multidrug resistance to a range of clinically important antibiotics in gram-negative bacteria like Pseudomonas aeruginosa, which lower the intracellular drug concentration by exporting incoming antibiotics across the membranes. We tried to show that the wild type susceptible bacteria P. aeruginosa modified its genetic makeup at mutational hotspots under stress. This strain may either become multidrug resistant or remain susceptible depending on position of amino acid changes in regulatory proteins of efflux pump. Multidrug resistant strain made significant changes at the amino acid positions, 103rd (G → A) and 126th (E → V) through the mutation on the nucleotide position of 308th (G → C); both 377th (A → T) and 378th (G → T), respectively in mexR, a repressor of mexAB-oprM efflux pump. This mutant protein showed low affinity with their operator. But the alteration at 103th position (G → A) in mexR may provide almost similar structural and functional stability as wild type. It was found that mutation was seemed to be well regulated within the limit and position specific under stress which might be back to its original form by supplying counter stress unless addition or deletion takes place.


Similar content being viewed by others
References
Toprak E, Veres A, Michel BJ, Chait R, Hart DL, Kishony R (2011) Evolutionary paths to antibiotic resistance under dynamically sustained drug selection. Nat Genet 44:101–105. https://doi.org/10.1038/ng.1034
Yen P, Papin JA (2017) History of antibiotic adaptation influences microbial evolutionary dynamics during subsequent treatment. PLoS Biol 15:e2001586. https://doi.org/10.1371/journal.pbio.2001586
York A (2017) Historical influences on antibiotic resistance. Nat Rev Microbiol. https://doi.org/10.1038/nrmicro.2017.111
Kelly E, Lateef A, Keith P (2001) MexR repressor of the mexAB-oprM multidrug efflux operon of Pseudomonas aeruginosa: identification of MexR binding sites in the mexA-mexR intergenic region. J Bacteriol 183:807–812. https://doi.org/10.1128/JB.183.3.807-812.2001
Tjong H, Zhou HX (2007) DISPLAR: an accurate method for predicting DNA-binding sites on protein surfaces. Nucl Acids Res 35:1465–1477
Yan Y, Zhang D, Zhou P, Li B, Huang SY (2017) HDOCK: a web server for protein–protein and protein–DNA/RNA docking based on a hybrid strategy. Nucl Acids Res 1:1. https://doi.org/10.1093/nar/gkx407
Daniel L, Keith Natali N CJ (2002) Crystal structure of the MexR repressor of the mexRAB-oprM multidrug efflux operon of Pseudomonas aeruginosa. J Biochem Chem 277:29253–29259. https://doi.org/10.1074/jbc.M111381200
Blázquez J, Gómez-Gómez JM, Oliver A, Juan C, Kapur V, Martín S (2006) PBP3 inhibition elicits adaptive responses in Pseudomonas aeruginosa. Mol Microbiol 62:84–99. https://doi.org/10.1111/j.1365-2958.2006.05366.x
Mahoney TF, Silhavy TJ (2013) The Cpx stress response confers resistance to some, but not all, bactericidal antibiotics. J Bacteriol 195:1869–1874. https://doi.org/10.1128/JB.02197-12
Tian Z, Yi X, Cho A, O’Gara F, Wang Y (2016) CpxR activates MexAB-OprM efflux pump expression and enhances antibiotic resistance in both laboratory and clinical nalB-type isolates of Pseudomonas aeruginosa. PLoS Pathogen 12:e1005932. https://doi.org/10.1371/journal.ppat.1005932
Hermann T (2005) Drugs targeting the ribosome. Curr Opin Struct Biol 15:355–366. https://doi.org/10.1016/j.sbi.2005.05.001
Raivio TL, Silhavy TJ (2001) Periplasmic stress and ECF sigma factors. Annu Rev Microbiol 55:591–624. https://doi.org/10.1146/annurev.micro.55.1.591
Fraud S, Poole K (2011) Oxidative stress induction of the MexXY multidrug efflux genes and promotion of aminoglycoside resistance development in Pseudomonas aeruginosa. Antimicrob Agents Chemother 55:1068–1074. https://doi.org/10.1128/AAC.01495-10
Hay T, Fraud S, Lau CH, Gilmour C, Poole K (2013) Antibiotic Inducibility of the mexXY multidrug efflux operon of Pseudomonas aeruginosa: involvement of the MexZ anti-repressor ArmZ. PLoS ONE 8:e56858. https://doi.org/10.1371/journal.pone.0056858
Kalia VC (2014) Microbes, antimicrobials and resistance: the battle goes on. Indian J Microbiol 54:1–2. https://doi.org/10.1007/s12088-013-0443-7
Kalia VC (2013) Quorum sensing inhibitors: an overview. Biotechnol Adv 31:224–245. https://doi.org/10.1016/j.biotechadv.2012.10.004
Kalia VC, Patel SKS, Kang YC, Lee JK (2019) Quorum sensing inhibitors as antipathogens: biotechnological applications. Biotechnol Adv 37:68–90. https://doi.org/10.1016/j.biotechadv.2018.11.006
Acknowledgements
Raju Biswas is thankful to CSIR for Junior Research Fellowship (File No: 09/025(0216)/2015-EMR-I) and Principal, Symsundar College, Shymsundar, Burdwan for conducting the research. Authors are thankful to UGC-Center of Advanced Study and DST-FIST, Department of Botany, The University of Burdwan for pursuing research activities.
Author information
Authors and Affiliations
Contributions
Author’s Contributions
RB 1 and 2 adopted the idea. RB 1 and ASP conducted some in silico work and interpreted the data. RB 1, 2 and ASP wrote and all authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Biswas, R., Panja, A.S. & Bandopadhyay, R. Molecular Mechanism of Antibiotic Resistance: The Untouched Area of Future Hope. Indian J Microbiol 59, 254–259 (2019). https://doi.org/10.1007/s12088-019-00781-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-019-00781-6