Abstract
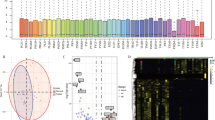
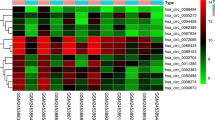
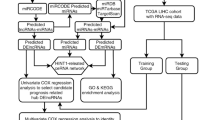
Thyroid cancer (TC) is the most common endocrine cancer, accounting for 1.7% of all cancer cases. It has been reported that the existing approach to diagnosing TC is problematic. Therefore, it is essential to develop molecular biomarkers to improve the accuracy of the diagnosis. This study aimed to screen hub lncRNAs in the ceRNA network (ceRNET) connected to TC formation and progression based on the overall survival rate. In this study, first, RNA-seq data from the GDC database were collected. A package called edgeR in R programming language was then used to obtain differentially expressed lncRNAs (DElncRNAs), miRNAs (DEmiRNAs), and mRNAs (DEmRNAs) in TC patients' samples compared to normal samples. Second, DEmRNAs were analyzed for their functional enrichment. Third, to identify RNAs associated with overall survival, the overall survival of these RNAs was analyzed using the Kaplan-Meier plotter database to create a survival associated with the ceRNA network (survival-related ceRNET). Next, the GeneMANIA plugin was used to construct a PPI network to better understand survival-related DEmRNA interactions. The survival ceRNET was then visualized with the Cytoscape software, and hub genes, including hub lncRNAs and hub mRNAs, were identified using the CytoHubba plugin. We found 45 DElncRNAs, 28 DEmiRNAs, and 723 DEmRNAs among thyroid tumor tissue and non-tumor tissue samples. According to KEGG, GO and DO analyses, 723 DEmRNAs were mainly enriched in cancer-related pathways. Importantly, the results found that ten DElncRNAs, four DEmiRNAs, and 68 DEmRNAs are associated with overall survival. In this account, the PPI network was constructed for 68 survival-related DEmRNAs, and ADAMTS9, DTX4, and CLDN10 were identified as hub genes. The ceRNET was created by combining six lncRNAs, 109 miRNAs, and 22 mRNAs related to survival using Cytoscape. in this network, ten hub RNAs were identified by the CytoHubba plugin, including mRNAs (CTXND1, XKRX, IGFBP2, ENTPD1, GALNT7, ADAMTS9) and lncRNAs (AC090673.1, AL162511.1, LINC02454, AL365259.1). This study suggests that three lncRNAs, including AL162511.1, AC090673.1, and AL365259.1, could be reliable diagnostic biomarkers for TC. The findings of this study provide a basis for future studies on the therapeutic potential of these lncRNAs.
Graphical abstract










Similar content being viewed by others
References
Al-Hashimi A, Venugopalan V, Sereesongsaeng N, Tedelind S, Pinzaru AM, Hein Z et al (2020) Significance of nuclear cathepsin V in normal thyroid epithelial and carcinoma cells. Biochim Biophys Acta Mol Cell Res 1867(12):118846
Bian W, Jiang XX, Wang Z, Zhu YR, Zhang H, Li X et al (2021) Comprehensive analysis of the ceRNA network in coronary artery disease. Sci Rep 11(1):1–11
Brito JP, Morris JC, Montori VM (2013)Thyroid cancer: zealous imaging has increased detection and treatment of low risk tumours. Bmj 347
Cai Y, Wan J (2018) Competing endogenous RNA regulations in neurodegenerative disorders: current challenges and emerging insights. Front Mol Neurosci 11:370
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY (2014) cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8(Supp 4):S11
Dong M, Xie Y, Xu Y (2019) miR-7-5p regulates the proliferation and migration of colorectal cancer cells by negatively regulating the expression of Krüppel-like factor 4. Oncol Lett 17(3):3241–3246
Esteller M (2011) Non-coding RNAs in human disease. Nat Rev Genet 12(12):861–74
Expansion of the Gene (2017) Ontology knowledgebase and resources. Nucleic Acids Res 45(D1):D331–D338
Feinberg AP, Ohlsson R, Henikoff S (2006) The epigenetic progenitor origin of human cancer. Nat rev genet 7(1):21–33
Goel MK, Khanna P, Kishore J (2010) Understanding survival analysis: kaplan-meier estimate. Int J Ayurveda Res 1(4):274
Gray RJ (2002) Modeling survival data: extending the cox model. J Am Stat Assoc 97(457):353–4. https://doi.org/10.1198/jasa.2002.s447
Grillone K, Riillo C, Scionti F, Rocca R, Tradigo G, Guzzi PH et al (2020) Non-coding RNAs in cancer: platforms and strategies for investigating the genomic dark matter. J Exp Clin Cancer Res 39(1):117
Guo K, Qian K, Shi Y, Sun T, Wang Z (2021) LncRNA-MIAT promotes thyroid cancer progression and function as ceRNA to target EZH2 by sponging miR-150-5p. Cell Death Dis 12(12):1–12
He Z, Deng W, Jiang B, Liu S, Tang M, Liu Y et al (2018) Hsa-let-7b inhibits cell proliferation by targeting PLK1 in HCC. Gene 673:46–55. https://doi.org/10.1016/j.gene.2018.06.047
Huang Q, Zhou Y, Li Y, Liao Z (2020) GGCT promotes colorectal cancer migration and invasion via epithelial-mesenchymal transition. Oncol Lett 20(2):1063–70
Huang HY, Lin YCD, Li J, Huang KY, Shrestha S, Hong HC et al (2020) miRTarBase 2020: updates to the experimentally validated microRNA-target interaction database. Nucleic Acids Res 48(D1):D148–D154
Hui L, Zheng F, Bo Y, Sen-Lin M, Ai-Jun L, Wei-Ping Z et al (2020) MicroRNA let-7b inhibits cell proliferation via upregulation of p21 in hepatocellular carcinoma. Cell Biosci 10(1):83
Jeggari A, Marks DS, Larsson E (2012) miRcode: a map of putative microRNA target sites in the long non-coding transcriptome. Bioinformatics 28(15):2062–2063
Jiang MC, Ni JJ, Cui WY, Wang BY, Zhuo W (2019) Emerging roles of lncRNA in cancer and therapeutic opportunities. Am J Cancer Res 9(7):1354
Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45(D1):D353-61
Kang H, Rho JG, Kim C, Tak H, Lee H, Ji E et al (2017) The miR-24-3p/p130Cas: a novel axis regulating the migration and invasion of cancer cells. Sci Rep 7:44847
Kawasaki K, Nojima S, Hijiki S, Tahara S, Ohshima K, Matsui T et al (2020) FAM111B enhances proliferation of KRAS-driven lung adenocarcinoma by degrading p16. Cancer Sci. 111(7):2635–46
Khella HWZ, Bakhet M, Allo G, Jewett MAS, Girgis AH, Latif A et al (2013) miR-192, miR-194 and miR-215: a convergent microRNA network suppressing tumor progression in renal cell carcinoma. Carcinogenesis 34(10):2231–2239. https://doi.org/10.1093/carcin/bgt184
Kilfoy BA, Zheng T, Holford TR, Han X, Ward MH, Sjodin A et al (2009) International patterns and trends in thyroid cancer incidence, 1973–2002. Cancer causes control 20(5):525–31
Law CW, Chen Y, Shi W, Smyth GK (2014) voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15(2):R29. https://doi.org/10.1186/gb-2014-15-2-r29
Li L, Kang L, Zhao W, Feng Y, Liu W, Wang T et al (2017) miR-30a-5p suppresses breast tumor growth and metastasis through inhibition of LDHA-mediated warburg effect. Cancer Lett 400:89–98. https://doi.org/10.1016/j.canlet.2017.04.034
Li R, Qu H, Wang S, Wei J, Zhang L, Ma R et al (2018) GDCRNATools: an R/Bioconductor package for integrative analysis of lncRNA, miRNA and mRNA data in GDC. Bioinformatics 34(14):2515–7
Liu X, Liu G, Lu Y, Shi Y (2020) Long non-coding RNA HOTAIR promotes cell viability, migration and invasion in thyroid cancer by sponging miR-17-5p. Neoplasma 67(2):229–37
Lu NH, Wei CY, Qi FZ, Gu JY (2020) Hsa-let-7b suppresses cell proliferation by targeting UHRF1 in melanoma. Cancer Invest 38(1):52–60
Nagaiah G, Hossain A, Mooney CJ, Parmentier J, Remick SC (2011) Anaplastic thyroid cancer: a review of epidemiology, pathogenesis, and treatment. J Oncol
Paci P, Colombo T, Farina L (2014) Computational analysis identifies a sponge interaction network between long non-coding RNAs and messenger RNAs in human breast cancer. BMC Syst Biol 8(1):83
Park CS, Kim SH, Jung SL, Kang BJ, Kim JY, Choi JJ et al (2010) Observer variability in the sonographic evaluation of thyroid nodules. J Clin Ultrasound 38(6):287–93
Peng X, Zhang K, Ma L, Xu J, Chang W (2020) The role of long non-coding RNAs in thyroid cancer. Front Oncol 10:941
Quinn JJ, Chang HY (2016) Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet 17(1):47–62
Ramírez-Moya J, Wert-Lamas L, Acuña-Ruíz A, Fletcher A, Wert-Carvajal C, McCabe CJ et al (2022) Identification of an interactome network between lncRNAs and miRNAs in thyroid cancer reveals SPTY2D1-AS1 as a new tumor suppressor. Sci Rep 12(1):1–13
Robinson MD, Oshlack A (2010) A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol 11(3):R25. https://doi.org/10.1186/gb-2010-11-3-r25
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP (2011) A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146(3):353–8
Schriml LM, Mitraka E, Munro J, Tauber B, Schor M, Nickle L et al (2019) Human Disease Ontology 2018 update: classification, content and workflow expansion. Nucleic Acids Res. 47(D1):D955–D962
See A, Iyer NG, Tan NC, Teo C, Ng J, Soo KC et al (2017) Distant metastasis as the sole initial manifestation of well-differentiated thyroid carcinoma. Europ Arch Oto-Rhino-Laryngol 274(7):2877–82
Seib CD, Sosa JA (2019) Evolving understanding of the epidemiology of thyroid cancer. Endocrinology and Metabolism Clinics. 48(1):23–35
Seo D, Kim D, Chae Y, Kim W.( 2020) The ceRNA network of lncRNA and miRNA in lung cancer. Genomics Informa 18(4)
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13(11):2498–504
Su K, Wang N, Shao Q, Liu H, Zhao B, Ma S (2021) The role of a ceRNA regulatory network based on lncRNA MALAT1 site in cancer progression. Biomed Pharmacother 137:111389
Sun H, Liu K, Huang J, Sun Q, Shao C, Luo J et al (2019) FAM111B, a direct target of p53, promotes the malignant process of lung adenocarcinoma. Onco Targets Ther 12:2829
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 71(3):209–49
Tam AA, Ozdemir D, Aydın C, Bestepe N, Ulusoy S, Sungu N et al (2018) Association between preoperative thyrotrophin and clinicopathological and aggressive features of papillary thyroid cancer. Endocrine 59(3):565–72
Tan J, Liu L, Zuo Z, Song B, Cai T, Ding D et al (2020) Overexpression of novel long intergenic non-coding RNA LINC02454 is associated with a poor prognosis in papillary thyroid cancer. Oncol Rep 44(4):1489–1501
Taniguchi K, Matsumura K, Ii H, Kageyama S, Ashihara E, Chano T et al (2018) Depletion of gamma-glutamylcyclotransferase in cancer cells induces autophagy followed by cellular senescence. Am J Cancer Res 8(4):650–61
4 USDOEJGIHT 4 BE 4 PP 4 RP 4 WS 4 ST 4 DN 4 CJF 4 OA 4 LS 4 EC 4 UE 4 FM, 9 RGSCSY 9 FA 9 HM 9 YT 9 TA 9 IT 9 KC 9 WH 9 TY 9 TT, 10 G and CU 8030: WJ 10 HR 10 SW 10 AF 10 BP 10 BT 10 PE 10 RC 10 WP, Department of Genome Analysis I of MBRA 12 PM 12 NG 12 TS 12 RA 12, 11 GTCSCSDR 11 DSL 11 RM 11 WK 11 LHM 11 DJ, 15 BGIGCYH 13 YJ 13 WJ 13 HG 14 GJ (2001) Initial sequencing and analysis of the human genome. nature.;409 (6822):860–921.
Wang C, Yan G, Zhang Y, Jia X, Bu P (2015) Long non-coding RNA MEG3 suppresses migration and invasion of thyroid carcinoma by targeting of Rac1. Neoplasma 62(4):541–9
Wang L, Cho KB, Li Y, Tao G, Xie Z, Guo B (2019) Long non-coding RNA (lncRNA)-mediated competing endogenous RNA networks provide novel potential biomarkers and therapeutic targets for colorectal cancer. Int J Mol Sci. 20(22):5758
Warde-Farley D, Donaldson SL, Comes O, Zuberi K, Badrawi R, Chao P et al (2010) The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 38(2):W214–W220
Xiao H (2019) MiR-7-5p suppresses tumor metastasis of non-small cell lung cancer by targeting NOVA2. Cell Mol Biol Lett 24(1):60
Xu J, Xu J, Liu X, Jiang J (2022) The role of lncRNA-mediated ceRNA regulatory networks in pancreatic cancer. Cell Death Discov 8(1):1–11
Yan-Chun L, Hong-Mei Y, Zhi-Hong C, Qing H, Yan-Hong Z, Ji-Fang W (2017) MicroRNA-192-5p promote the proliferation and metastasis of hepatocellular carcinoma cell by targeting SEMA3A. Appl Immunohistochem Mol Morphol. 25(4):251–60
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 16(5):284–287
Zargari N, Mokhtari M (2019) Evaluation of diagnostic utility of immunohistochemistry markers of TROP-2 and HBME-1 in the diagnosis of thyroid carcinoma. Eur Thyroid J 8(1):1–6
Zhang Y (2019) Classification and diagnosis of thyroid carcinoma using reinforcement residual network with visual attention mechanisms in ultrasound images. J Med Syst 43(11):1–9
Zhang W, Chen L, Xiang H, Hu C, Shi W, Dong P et al (2016) Knockdown of GGCT inhibits cell proliferation and induces late apoptosis in human gastric cancer. BMC Biochem 17(1):19
Zhu J, Zeng Y, Li W, Qin H, Lei Z, Shen D et al (2017) CD73/NT5E is a target of miR-30a-5p and plays an important role in the pathogenesis of non-small cell lung cancer. Mol Cancer. 16(1):34
Zhu W, Wang Y, Zhang D, Yu X, Leng X (2018) MiR-7-5p functions as a tumor suppressor by targeting SOX18 in pancreatic ductal adenocarcinoma. Biochem Biophys Res Commun 497(4):963–70
Acknowledgments
We wish to acknowledge The Cancer Genome Atlas (TCGA) project, miRcode, Cytoscape, GO, KEGG, and GeneMANIA databases, and their contributors for presenting these valuable public data sets.
Author information
Authors and Affiliations
Contributions
PM and SM wrote the manuscript comprehensively in all parts, MH, FGZM, and MJK accompanied in many other sections of the paper, and AAS edited the manuscript scientifically and technically. All the authors read the manuscript comprehensively and confirmed the final edited version. Importantly, there is no conflict of interest.
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Morovat, P., Morovat, S., Hosseinpour, M. et al. Survival-based bioinformatics analysis to identify hub long non-coding RNAs along with lncRNA-miRNA-mRNA network for potential diagnosis/prognosis of thyroid cancer. J. Cell Commun. Signal. 17, 639–655 (2023). https://doi.org/10.1007/s12079-022-00697-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12079-022-00697-9