Abstract
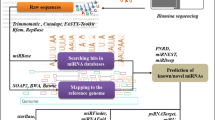
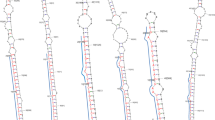
MicroRNAs (miRNAs) are generated in a cell endogenously which have a nucleotide length of 18 to 26 and are also known as short non-protein coding RNAs. The majority of them are evolutionarily conserved in nature, suggesting a logical foundation for the anticipation of new miRNAs in association with their clusters in numerous plants. Considering this study, physical, bioinformatics and efficient methods are integrated to predict the fresh miRNA clusters along with their targets in sugar cane. In sugar cane, there were a total of 21 new miRNA clusters identified, which were linked to 15 miRNA families. These families are as 165a, 166b, 528a, 827, 2118, 2120b, 5168, 5564c, 5565g, 5568c, 6220, 6225, 6226, 6232a and 7540a. Multiple characteristics of such miRNA clusters, including web logo, phylogenetic tree and secondary structures have been developed. The minimal free energy (MFE) of the secondary structures has been attained and reported as well. In addition, mature miRNAs have been sought in stem section of the structure. Consequently, 115 miRNA targets were also found. These targets include substantial GO term which have important targets in the reproduction, DNA packaging, multicellular organismal process, gene expression, translation, transcription factors, protein binding, transporter activity, secretion, cell division, binding, growth & development and aging. Hence, the achieved results of novel sugar cane miRNA clusters target several types of significant genes which help in managing the environment for sugar cane for better crop production.
Graphical Abstract







Similar content being viewed by others
References
Achakzai HK, Barozai MYK, Din M, Baloch IA, Achakzai AKK (2018) Identification and annotation of newly conserved microRNAs and their targets in wheat (Triticum aestivum L). PLoS ONE 13(7):e0200033. https://doi.org/10.1371/journal.pone.0200033
Achakzai HK, Barozai MYK, Achakzai AKK, Asghar M, Din M (2019a) Profiling of 21 novel microRNA clusters and their targets in an important grain: wheat (Triticum aestivum L). Pak J Bot 51(1):133–142. https://doi.org/10.30848/PJB2019-1(35)
Achakzai HK, Barozai MYK, Baloch IA, Achakzai AKK, Din M, Asghar M (2019b) The identification of eighteen precursor miRNA clusters and their targets in barley (Hordeum vulgare L). Pak J Bot 51(2):469–477
Ahmad M, Ali Q, Hafeez MM, Malik A (2021) Improvement for biotic and abiotic stress tolerance in crop plants. Biol Clin Sci Res 2021. https://doi.org/10.54112/bcsrj.v2021i1.50
Ambros V, Bartel B, Bartel DP (2003) A uniform system for microRNA annotation. RNA 9(3):277–279. http://www.rnajournal.org/cgi/doi/10.1261/rna.2183803
Awaad HA, Negm AM, Abu-hashim M (2021) Update, conclusions, and recommendations of Mitigating Environmental stresses for agricultural sustainability in Egypt. Mitigating Environmental Stresses for Agricultural Sustainability in Egypt. Springer, Cham, pp 561–590. https://doi.org/10.1007/978-3-030-64323-2_21
Balmer D, Mauch-Mani B (2013) Small yet mighty– microRNAs in plant-microbe interactions. MicroRNA 2(1):73–80
Baloch IA, Barozai MYK, Din M (2015a) Identification and characterization of 25 and their targeted proteins microRNAs in Apricot (Prunus armeniaca L). J Anim Plant Sci 25(5):1466–1476
Baloch IA, Barozai MYK, Din M, Achakzai AKK (2015b) Computational identification of 18 miRNAs and their targets in three species of Rose. Pak J Bot 47(4):1281–1285
Baloch IA, Barozai MYK, Din M (2018) Bioinformatics Prediction and Annotation of Cherry (Prunus avium L.) microRNAs and their targeted proteins. Turk J Bot 42. https://doi.org/10.3906/bot-1712-37
Baloch IA, Barozai MYK, Baloch AH, Din M (2019) Bioinformatic prediction and annotation of apple microRNAs and their targets. Pak J Bot 51:909–922
Baloch IA, Barozai MYK, Din M (2021) microRNAs: the mega regulators in eukaryotic genomes. Pure Appl Biol (PAB) 2(3):83–88. https://doi.org/10.19045/bspab.2013.23002
Barozai MYK (2013) Identification of microRNAs and their targets in Artemisia annua L. Pak J Bot 45(2):461–465
Barozai MYK, Wahid HA (2012) In-silico identification and characterization of cumulative abiotic stress responding genes in potato (Solanum tuberosum L). Pak J Bot 44:57–69
Barozai MYK, Irfan M, Yousaf R, Ali I, Qaisar U, Maqbool A, Zahoor M, Rashid B, Hussnain T, Riazuddin S (2008) Identification of micro-RNAs in cotton. Plant Physiol Biol 46(8–9):739–756. https://doi.org/10.1016/j.plaphy.2008.05.009
Barozai MYK, Baloch IA, Din M (2012) Identification of MicroRNAs and their targets in Helianthus. Mol Biol Rep 39(3):2523–2532
Barozai MYK, Shah SQ, Din M, Muhammad R (2014) Codon usage bias and RNA secondary structures analysis for virus resistant genes in Arabidopsis thaliana and Oryza sativa. Pure Appl Biol 3:81–91. https://doi.org/10.19045/bspab.2014.32005
Barozai MYK, Ye Z, Sangireddy SR, Zhou S (2018) Bioinformatics profiling and expressional studies of microRNAs in root, stem and leaf of the bioenergy plant switch grass (Panicum virgatum L.) under drought stress. Agri Gene 8:1–8. https://doi.org/10.1016/j.aggene.2018.02.001
Bartel DP (2009) MicroRNAs: Target Recognition and Regulatory functions. Cell 136(2):215–233. https://doi.org/10.1016/j.cell.2009.01.002
Chen X, GAO W, Zhang J, Zhang X, Lin Z (2013) Linkage mapping and expression analysis of miRNAs and their target genes during fiber development in cotton. BMC Genom 14(1):706
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) Web-Logo: a sequence logo generator. Genome Res 14(6):1188–1190. http://www.genome.org/cgi/doi/10.1101/gr.849004
Dai X, Zhao PX (2011) PsRNATarget: a plant small RNA target analysis server. Nucleic Acids Res 39(suppl–2). https://doi.org/10.1093/nar/gkr319. W155-W159
Din M, Barozai MYK, Baloch IA (2016) Profiling and annotation of microRNAs and their putative target genes in Chilli (Capsicum annuum L.) using ESTs. Gene Rep 5:62–69. https://doi.org/10.1016/j.genrep.2016.08.010
Din M, Barozai MYK, Aziz AN (2018) In Silico profiling and characterization of conserved microRNAs in Biofuel Plant Sorghum. Pak J Bot 50(6):2265–2275
Duarte-Almeida JM, Salatino A, Genovese MI, Lajolo FM (2011) Phenolic composition and antioxidant activity of culms and sugarcane (Saccharum officinarum L.) products. Food Chem 125(2):660–664. https://doi.org/10.1016/j.foodchem.2010.09.059
Eskandarynasab S, Roudbari Z, Bahreini Behzadi MR (2020) Clustering based on the ontology of microRNAs target genes affecting milk production. J Anim Environ 12(3):435–440. https://doi.org/10.22034/aej.2020.118071
Fontana DC, de Paula S, Torres AG, de Souza VHM, Pascholati SF, Schmidt D, Dourado Neto D (2021) Endophytic fungi: biological control and induced resistance to phytopathogens and abiotic stresses. Pathogens 10(5):570. https://doi.org/10.3390/pathogens10050570
Ghani A, Din M, Baloch IA, Barozai MYK (2013) Identification of microRNA in 12 plant species of Fabaceae. Pure Appl Biol 2(3):104–115
Islam W, Waheed A, Naveed H, Zeng F (2022) MicroRNAs Mediated Plant Responses to Salt Stress. Cells 11(18):2806. https://doi.org/10.3390/cells11182806
Iwakawa HO, Tomari Y (2021) Life of RISC: formation, action, and degradation of RNA-induced silencing complex. Mol Cell. https://doi.org/10.1016/j.molcel.2021.11.026
Jan R, Asaf S, Numan M, Kim KM (2021) Plant secondary metabolite biosynthesis and transcriptional regulation in response to biotic and abiotic stress conditions. Agronomy 11(5):968. https://doi.org/10.3390/agronomy11050968
Khvorova A, Reynolds A, Jayasena SD (2007) Functional siRNAs and miRNAs exhibit strand bias. Cell 131(2):41–49
Kirchner B (2022) Functional importance of intra-and extracellular microRNAs and their isoforms in blood and milk. Technische Universität München. https://mediatum.ub.tum.de/1617916. Doctoral dissertation
Lee RC, Ambros V (2001) An extensive class of small RNAs in Caenorhabditis elegans. Science 294:862–864. https://doi.org/10.1126/science.1065329
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH, Kim VM (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23(20):4051–4060. https://doi.org/10.1038/sj.emboj.7600385
Pantaleo V, Szittya G, Moxon S, Miozzi L, Moulton V, Dalmay T, Burgyan J (2010) Identification of grapevine microRNAs and their targets using high throughput sequencing and degraded analysis. Plants J 62(6):960–976. https://doi.org/10.1111/j.1365-313X.2010.04208.x
Paul S, Bravo Vazquez LA, Marquez Nafarrate M, Gutierrez Resendiz AI, Srivastava A, Sharma A (2021) The regulatory activities of microRNAs in non-vascular plants: a mini review. Planta 254(3):1–23. https://doi.org/10.1007/s00425-021-03707-z
Rani V, Sengar RS (2022) Biogenesis and mechanisms of microRNA-mediated gene regulation. Biotechnol Bioeng 119(3):685–692. https://doi.org/10.1002/bit.28029
Rasheed A, Hassan MU, Aamer M, Batool M, Sheng FANG, Ziming WU, Huijie LI (2020) A critical review on the improvement of drought stress tolerance in rice (Oryza sativa L). Notulae Botanicae Horti Agrobotanici Cluj-Napoca 48(4). https://doi.org/10.15835/48412128
Schwarz DS, Zamore PD (2002) Why do miRNAs live in the miRNP? Genes Dev 16(9):1025–1031. https://doi.org/10.1101/gad.992502
Tian T, Liu Y, Yan H, You Q, Yi X, Du Z, Xu W, Su Z (2017) agriGO v2. 0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res 45(W1):W122–W129. https://doi.org/10.1093/nar/gkx382
Vaucheret H, Mallory A, Lepere G (2012) European patent no. EP 2436768. European Patent Office, Munich, Germany
Wang J, Yang X, Xu H, Chi X, Zhang M, Hou X (2012) Identification and characterization of microRNAs and their target genes in Brassica oleracea. Gene 505(2):300–308. https://doi.org/10.1016/j.gene.2012.06.002
Xue A, Li Z, Cai M, Zhang Q, Zhang X, Ming R, Zhang J (2017) Identification and characterization of microRNAs from Saccharum officinarum L by Deep Sequencing. Trop Plant Biol 10(2):134–150. https://doi.org/10.1007/s12042-017-9190-y
Yu J, Wang F, Yang GH, Wang FL, Ma YN, Du ZW, Zhang JW (2006) Human microRNA clusters: genomic organization and expression profile in Leukemia cell lines. Biochem Biophys Res Commun 349(1):59–68. https://doi.org/10.1016/j.bbrc.2006.07.207
Zhang B, Pan X, Wang Q, Cobb GP, Anderson TA (2006) Computational identification of microRNAs and their targets. Comput Biol Chem 30(6):395–407. https://doi.org/10.1016/j.compbiolchem.2006.08.006
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31(13):3406–3415. https://doi.org/10.1093/nar/gkg595
Acknowledgements
The authors are thankful to the faculty members of Center for Applied Molecular Biology, University of Punjab, Lahore (CAMB) for encouragement and providing necessary facilities. The authors also acknowledge the Department of Botany, University of Balochistan for providing the necessary resources.
Funding
No funding information is available from any author.
Author information
Authors and Affiliations
Contributions
Conceived and designed the data: Abdul Baqi, Samiullah & Muhammad Zafar Saleem. Arranged the tools: Abdul Baqi & Muhammad Zafar Saleem. Analyzed the data: Abdul Baqi & Muhammad Zafar Saleem. Contributed tools: Muhammad Zafar Saleem, Muhammad Ayub & Shazia Saeed. Wrote the paper: Abdul Baqi, Samiullah & Muhammad Zafar Saleem.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Communicated by Rui Xia.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Baqi, A., Samiullah, Saleem, M.Z. et al. Identification and Annotation of the 21 Novel Sugar Cane (Saccharum officinarum) MicroRNA Clusters and Their Significant Biological, Molecular and Cellular Targets. Tropical Plant Biol. 17, 65–81 (2024). https://doi.org/10.1007/s12042-023-09352-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12042-023-09352-y