Abstract
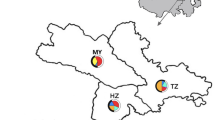
Single-nucleotide polymorphisms (SNPs), microsatellites and copy number variation (CNV) were studied on the Y chromosome to understand the paternal origin and phylogenetic relationships for resource protection, rational development and utilization of the domestic Bactrian camel in China. Our sample set consisted of 94 Chinese domestic Bactrian camels from four regions (Inner Mongolia, Gansu, Qinghai and Xinjiang), we screened 29 Y-chromosome-specific loci for SNPs, analysed 40 bovine-derived microsatellite loci and measured CNVs of HSFY and SRY through Sanger sequencing, automated fluorescence-based microsatellite analysis and quantitative real-time PCR, respectively. A multicopy gene, SRY, was first found, and sequence variation was only detected in SRY in a screen of 29 loci in 13 DNA pools of individual camels. In addition, a TG repeat in the USP9Y gene was identified as the first polymorphic microsatellite in the camel Y chromosome, whereas microsatellite based on bovine sequences were not detected. The frequency of each allele varied among different populations. For the Nanjiang, Hexi and Alashan populations, a 243-bp allele was found. For the Sunite population, 241-bp, 243-bp and 247-bp alleles were detected, and the frequencies of these alleles were \(22.2\%\), \(44.5\%\) and \(33.3\%\), respectively; 241-bp and 243-bp alleles were found in other populations. Finally, CNVs in two Y-chromosomal genes were detected; CNV for HSFY and SRY ranged from 1 to 3 and from 1 to 9, respectively.






Similar content being viewed by others
References
Chang T. C., Yang Y., Retzel E. F. and Liu W. S. 2013 Male-specific region of the bovine Y chromosome is gene rich with a high transcriptomic activity in testis development. Proc. Natl. Acad. Sci. 110, 12373--12378.
Cheng J., Ren Z. J., Wang L., Wang H. L., Wang Y. X., Azhati Z. L. P. K. E. et al. 2009 Molecular phylogeny of domestic Bactrian camel revealed by partial cytochrome b gene. J. Northwest A & F University-Natural Science Edition 37, 17–21 (in Chinese).
Cliffe K. M., Day A. E., Bagga M., Siggens K., Quilter C. R., Lowden S. et al. 2010 Analysis of the non-recombining Y chromosome defines polymorphisms in domestic pig breeds: ancestral bases identified by comparative sequencing. Anim. Genet. 41, 619--629.
Douadi M. I., Gatti S., Levrero F., Duhamel G., Bermejo M., Vallet D. et al. 2007 Sex-biased dispersal in western lowland gorillas (Gorilla gorilla gorilla). Mol. Ecol. 16, 2247--2259.
Giachini C., Nuti F., Turner D. J., Laface I., Xue Y., Daguin F. et al. 2009 TSPY1 copy number variation influences spermatogenesis and shows differences among Y lineages. J. Clin. Endocrinol Metab. 94, 4016–4022.
Hamilton C. K., Favetta L. A., Di Meo G. P., Floriot S., Perucatti A., Peippo J. et al. 2009 Copy number variation of testis-specific protein, Y-encoded (TSPY) in 14 different breeds of cattle (Bos taurus). Sex. Dev. 3, 205–213.
Hammond R. L., Handley L. J. L., Winney B. J., Bruford M. W. and Perrin N. 2006 Genetic evidence for female-biased dispersal and gene flow in a polygynous primate. Proc. R. Soc. London, Ser. B 273, 479–484.
Han H. Y., Zhang Q., Gao K. X., Yue X. P., Zhang T., Dang R. H. et al. 2015 Y-single nucleotide polymorphisms diversity in Chinese indigenous horse. Asian-Aust. J. Anim. Sci. 28, 1066.
Helen S., Tomoko K. K., Minx P. J., Cordum H. S., Ladeana H., Brown L. G. et al. 2003 The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 423, 825–837.
Ji R., Cui P., Ding F., Geng J., Gao H., Zhang H. et al. 2009 Monophyletic origin of domestic Bactrian camel (Camelus bactrianus) and its evolutionary relationship with the extant wild camel (Camelus bactrianus ferus). Anim. Genet. 40, 377--382.
Ji R. M. T., Wang Z., Ding G., Chen G., Sun Y., Sun Z. et al. 2013 Corrigendum: genome sequences of wild and domestic Bactrian camels. Nat. Commun. 4, 2089.
Kreutzmann N., Brem G. and Wallner B. 2014 The domestic horse harbours Y-chromosomal microsatellite polymorphism only on two widely distributed male lineages. Anim. Genet. 45, 460.
Lei C. Z., Chen H., Zhang H. C., Cai X., Liu R. Y., Luo L. Y. et al. 2006 Origin and phylogeographical structure of Chinese cattle. Anim. Genet. 37, 579--582.
Li R., Zhang X. M., Campana M. G., Huang J. P., Chang Z. H., Qi X. B. et al. 2013 Paternal origins of Chinese cattle. Anim. Genet. 44, 446--449.
Liu W. S., Mariani P., Beattie C. W., Alexander L. J. and De León F. A. P. 2002 A radiation hybrid map for the bovine Y Chromosome. Mam. Genome 13, 320–326.
Liu W. S., Beattie C. and Ponce de Leon F. 2004 Bovine Y chromosome microsatellite polymorphisms. Cytogenet. Genome Res. 102, 53–58.
Mukherjee A., Dass G., Mohanarao J., Gohain M., Brahma B., Datta T. K. et al. 2013 Absolute copy number differences of Y chromosomal genes between crossbred (Bos taurus \(\times \) Bos indicus) and Indicine bulls. J. Anim. Sci. Biotechnol. 4, 1–7.
Musiani M., Leonard J. A., Cluff H., Gates C. C., Mariani S., Paquet P. C. et al. 2007 Differentiation of tundra/taiga and boreal coniferous forest wolves: genetics, coat colour and association with migratory caribou. Mol. Ecol. 16, 4149--4170.
Natanaelsson C., Oskarsson M. C., Angleby H., Lundeberg J., Kirkness E. and Savolainen P. 2006 Dog Y chromosomal DNA sequence: identification, sequencing and SNP discovery. BMC Genet. 7, 45.
Paria N., Raudsepp T., Wilkerson A. J. P., O’Brien P. C., Ferguson-Smith M. A., Love C. C. et al. 2011 A gene catalogue of the euchromatic male-specific region of the horse Y chromosome: comparison with human and other mammals. PLoS One 6, e21374.
Pearks Wilkerson A. J., Raudsepp T., Graves T., Albracht D., Warren W., Chowdhary B. P. et al. 2008 Gene discovery and comparative analysis of X-degenerate genes from the domestic cat Y chromosome. Genomics 92, 329--338.
Prokop J. W., Underwood A. C., Turner M. E., Miller N., Pietrzak D., Scott S. et al. 2013 Analysis of Sry duplications on the Rattus norvegicus Y-chromosome. BMC Genomics 14, 117–124.
Sambrook J. and Russell D. W. 2002 Translated by Huang, PT molecular cloning a laboratory manual. Science Press, Beijing, China.
Skaletsky H., Kuroda-Kawaguchi T., Minx P. J., Cordum H. S., Hillier L., Brown L. G. et al. 2003 The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 423, 825--837.
Skinner B. M., Lachani K., Sargent C. A., Yang F., Ellis P., Hunt T. et al. 2015 Expansion of the HSFY gene family in pig lineages. BMC Genomics 16, 1--11.
Tessari A., Salata E., Ferlin A., Bartoloni L., Slongo M. L. and Foresta C. 2004 Characterization of HSFY, a novel AZFb gene on the Y chromosome with a possible role in human spermatogenesis. Mol. Hum. Reprod. 10, 253--258.
Turner M. E., Martin C., Martins A. S., Dunmire J., Farkas J., Ely D. L. et al. 2007 Genomic and expression analysis of multiple SRY loci from a single Rattus norvegicus Y chromosome. BMC Genet. 8, 1--11.
Underhill P. A., Shen P., Lin A. A., Jin L., Passarino G., Yang W. H. et al. 2000 Y chromosome sequence variation and the history of human populations. Nat. Genet. 26, 358--361.
Vollmer N. L. and Rosel P. E. 2012 Developing genomic resources for the common bottlenose dolphin (Tursiops truncatus): isolation and characterization of 153 single nucleotide polymorphisms and 53 genotyping assays. Mol. Ecol. Resou. 12, 1124--1132.
Wallner B., Piumi F., Brem G., Müller M. and Achmann R. 2004 Isolation of Y chromosome-specific microsatellites in the horse and cross-species amplification in the genus Equus. J. Hered. 95, 158–164.
Wallner B., Brem G., Müller M. and Achmann R. 2003 Fixed nucleotide differences on the Y chromosome indicate clear divergence between Equus przewalskii and Equus caballus. Anim. Genet. 34, 453--456.
Wilkerson A. J. P., Raudsepp T., Graves T., Albracht D., Warren W., Chowdhary B. P. et al. 2008 Gene discovery and comparative analysis of X-degenerate genes from the domestic cat Y chromosome. Genomics 92, 329--338.
Yue X. P., Dechow C., Chang T. C., DeJarnette J. M., Marshall C. E., Lei C. Z. et al. 2014 Copy number variations of the extensively amplified Y-linked genes, HSFY and ZNF280BY, in cattle and their association with male reproductive traits in Holstein bulls. BMC Genomics 15, 1.
Zhang C. D., Ren Z. J. and Yang X. J. 2015a Genetic diversity of Sunite domestic Bactrian Camel revealed by partial cytochrome b gene and D-loop sequence of mitochondrial DNA. Journal of Northwest A & F University-Natural Science Edition 43, 32–42 (in Chinese).
Zhang L. Z., Jia S. G., Plath M., Huang Y. Z., Li C. J., Lei C. H. et al. 2015b Impact of parental Bos taurus and Bos indicus origins on copy number variation in traditional Chinese cattle breeds. Genome Biol. Evol. 7, 2352–2361.
Zhang M., Peng W. F., Yang G. L., Lv F. H., Liu M. J., Li W. R. et al. 2014 Y chromosome haplotype diversity of domestic sheep (Ovis aries) in northern Eurasia. Anim. Genet. 45, 903–907.
Zhang P. Y., Na S., Bao Y., Xi R. M. and Zhang Y. X. 1991 Advances in breeding of Sunite white camel. Animal Husbandry and Veterinary Medicine 2, 50–51 (in Chinese).
Acknowledgements
This study was supported by the National Natural Science Foundation of China, 31172178.
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding editor: Indrajit Nanda
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, H., Ren, Z., Zhao, J. et al. Y-chromosome polymorphisms of the domestic Bactrian camel in China. J Genet 97, 3–10 (2018). https://doi.org/10.1007/s12041-017-0852-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-017-0852-1