Abstract
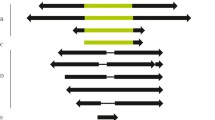
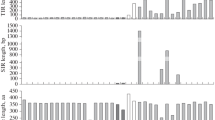
Mobilization of two P element subfamilies (canonical and O-type) from Drosophila sturtevanti and D. saltans was evaluated for copy number and transposition activity using the transposon display (TD) technique. Pairwise distances between strains regarding the insertion polymorphism profile were estimated. Amplification of the P element based on copy number estimates was highly variable among the strains (D. sturtevanti, canonical 20.11, O-type 9.00; D. saltans, canonical 16.4, O-type 12.60 insertions, on average). The larger values obtained by TD compared to our previous data by Southern blotting support the higher sensitivity of TD over Southern analysis for estimating transposable element copy numbers. The higher numbers of the canonical P element and the greater divergence in its distribution within the genome of D. sturtevanti (24.8%) compared to the O-type (16.7%), as well as the greater divergence in the distribution of the canonical P element, between the D. sturtevanti (24.8%) and the D. saltans (18.3%) strains, suggest that the canonical element occupies more sites within the D. sturtevanti genome, most probably due to recent transposition activity. These data corroborate the hypothesis that the O-type is the oldest subfamily of P elements in the saltans group and suggest that the canonical P element is or has been transpositionally active until more recently in D. sturtevanti.
Similar content being viewed by others
References
Almeida L. M., Langeani F. and Carareto C. M. A. 2003 Geographic polymorphism of P element populations of Drosophila sturtevanti. Genet. Mol. Biol. 26, 175–179.
Bartolomé C., Maside X. and Charlesworth B. 2002 On the abundance and distribution of transposable elements in the genome of Drosophila melanogaster. Mol. Biol. Evol. 19, 926–937.
Castro J. P. and Carareto C. M. A. 2004a P elements in the saltans group of Drosophila: a new evaluation of their distribution and number of genomic insertion sites. Mol. Phylogenet. Evol. 32, 383–387.
Castro J. P. and Carareto C. M. A. 2004b Canonical P elements are transcriptionally active in the saltans group of Drosophila. J. Mol. Evol. 59, 31–40.
Castro J. P. and Carareto C. M. A. 2004c Drosophila melanogaster P transposable elements: mechanisms of transposition and regulation. Genetica 121, 107–118.
Charlesworth B., Jarne P. and Assimacopoulos S. 1994 The distribution of transposable elements within and between chromosomes in a population of Drosophila melanogaster. III. Element abundances in heterochromatin. Genet. Res. 64, 183–197.
Clark J. B. and Kidwell M. G. 1997 A phylogenetic perspective on P transposable element evolution in Drosophila. Proc. Natl. Acad. Sci. USA 94, 11428–11433.
Clark J. B., Altheide T. K., Schlosser M. J. and Kidwell M. G. 1995 Molecular evolution of P transposable elements in the genus Drosophila I. The saltans and willistoni species groups. Mol. Biol. Evol. 12, 902–913.
Daniels S. B. and Strausbaugh L. D. 1986 The distribution of P element sequences in Drosophila: the willistoni and saltans species groups. J. Mol. Evol. 23, 138–148.
Daniels S. B., Peterson K. R., Strausbaugh L. D., Kidwell M. G. and Chovnick A. 1990 Evidence for horizontal transmission of the P element between Drosophila species. Genetics 124, 339–355.
Ellis T. H., Poyser S. J., Knox M. R., Vershinin A. V. and Ambrose M. J. 1998 Polymorphism of insertion sites of Tyl-copia class retrotransposons and its use for linkage and diversity analysis in pea. Mol. Gen. Genet. 260, 9–19.
Guimond N., Bideshi D. K., Pinkerton A. C., Atkinson P. W. and O’Brochta D. A. 2003 Patterns of Hermes transposition in Drosophila melanogaster. Mol. Genet. Genomics 268, 779–790.
Hagemann S., Miller W. J. and Pinsker W. 1992 Identification of a complete P element in the genome of Drosophila bifasciata. Nucleic Acids Res. 20, 409–413.
Hagemann S., Miller W. J. and Pinsker W. 1994 Two distinct P element subfamilies in the genome of Drosophila bifasciata. Mol. Gen. Genet. 244, 168–175.
Hagemann S., Haring E. and Pinsker W. 1996a Repeated horizontal transfer of P transposons between Scaptomyza pallida and Drosophila bifasciata. Genetica 98, 43–51.
Hagemann S., Haring E. and Pinsker W. 1996b A new P element subfamily from Drosophila tristis, D. ambigua and D. obscura. Genome 39, 978–985.
Haring E., Hagemann S. and Pinsker W. 1998 Transcription and splicing patterns of M-and O-type P elements in Drosophila bifasciata, D. helvetica, and Scaptomyza pallida. J. Mol. Evol. 46, 542–551.
Haring E., Hagemann S. and Pinsker W. 2000 Ancient and recent horizontal invasions of drosophilids by P elements. J. Mol. Evol. 51, 577–586.
Jowett T. 1986 Preparation of nucleic acids. In Drosophila: a practical approach (ed. D. B. Roberts), pp. 275–286. IRL Press, Oxford.
Kaplan N., Darden T. and Langley C. H. 1985 Evolution and extinction of transposable elements in Mendelian populations. Genetics 109, 459–480.
Loreto E. L., Valente V. L., Zaha A., Silva J. C. and Kidwell M. G. 2001 Drosophila mediopunctata P elements: a new example of horizontal transfer. J. Hered. 92, 375–381.
Maside X., Bartolomé C., Assimacopoulos S. and Charlesworth B. 2001 Rates of movement and distribution of transposable elements in Drosophila melanogaster: in situ hybridization vs Southern blotting data. Genet. Res. 78, 121–136.
Nouaud D., Quenesville H. and Anxolabéherè D. 2003 Recurrent exon shuffling between distant P-element families. Mol. Biol. Evol. 20, 190–199.
Oliveira de Carvalho M., Silva J. C. and Loreto E. L. 2004 Analyses of P-like transposable element sequences from the genome of Anopheles gambiae. Insect Mol. Biol. 13, 55–63.
Rizzon C., Marais G., Gouy M. and Biémont C. 2002 Recombination rate and the distribution of transposable elements in the Drosophila melanogaster genome. Genome Res. 12, 400–407.
Sarkar A., Sengupta R., Krzywinski J., Wang X., Roth C. and Collins F. H. 2003 P elements are found in the genomes of nematoceran insects of the genus Anopheles. Insect Biochem. Mol. Biol. 33, 381–387.
Silva J. C. and Kidwell M. G. 2000 Horizontal transfer and selection in the evolution of P elements. Mol. Biol. Evol. 17, 1542–1557.
Silva J. C. and Kidwell M. G. 2004 Evolution of P elements in natural populations of Drosophila willistoni and D. sturtevanti. Genetics 168, 1323–1335.
Swofford D. 2002 Phylogenetic analysis using parsimony ( * and other methods) version 4.0b10. Smithsonian Institution, Washington, DC, and Sinauer, Sunderland.
Van den Broeck D., Maes T., Sauer M., Zethof J., De Keukeleire P., D’Hauw M. et al. 1998 Transposon display identifies individual transposable elements in high copy number lines. Plant J. 13, 121–129.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
de Setta, N., Costa, A.P.P., Lopes, F.R. et al. Transposon display supports transpositional activity of P elements in species of the saltans group of Drosophila . J Genet 86, 37–43 (2007). https://doi.org/10.1007/s12041-007-0005-z
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-007-0005-z