Abstract
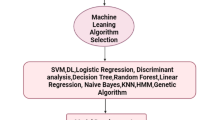
Tumors have drawn increasing attention recently because of their heterogeneous interior structures. Particularly, single-cell RNA (scRNA) mechanics have made important contributions to the field of tumor research. To investigate the cell types and identify similar types of gene markers present inside a tumor, machine learning classifier, optimization, and neural network models were applied to scRNA sequencing data. Indeed, even though single-cell analysis is a more powerful tool, several issues have been identified, such as transcriptional noise that alters gene expression and degrades mRNA. Recently, optimization models for single-cell analysis have been developed to address these kinds of issues, and encouraging results have been reported. scRNA sequencing is popular because it produces biological information in the form of patterns that are displayed within the transcriptome profile. The neural network approach plays an important role in understanding and identifying these distinct patterns. A single layer perceptron was introduced to better analyze the data pattern within gene expression profiles. Finally, recently developed optimization models with machine learning classifiers are compared with the proposed single layer perceptron. The single layer perceptron performs better compared with other models such as extra tree classifier with genetic algorithm, k-nearest neighbors with bat optimization, decision tree with gray wolf optimization, random forest with firefly optimization, and Gaussian naïve Bayes with artificial bee colony optimization. This study also focused on classifying these unique cell types and gene markers using scRNA sequence datasets. The proposed single layer perceptron was assessed using two datasets: normal mucosa and colorectal tumors. Our findings showed that the proposed single layer perceptron performed exceptionally well with accuracy, precision, recall, and F1 value.




















Similar content being viewed by others
Data availability
The scRNAseq tumor datasets have been collected from the gene expression omnibus database Article 17 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE81861] (GSE81861) which contains information about normal mucosa and colorectal tumors.
References
Aiwale A and Ansari S 2019 Brain tumor detection using knn. Int. J. Sci. Engineer. Res. 10 187–193
Anand MV, KiranBala B, Srividhya S, et al. 2022 Gaussian naive Bayes algorithm: A reliable technique involved in the assortment of the segregation in cancer. Mobile Inform. Syst. https://doi.org/10.1155/2022/2436946
Asada K, Takasawa K, Machino H, et al. 2021 Single-cell analysis using machine learning techniques and its application to medical research. Biomedicines 9 1513
Chellamuthu G, Kandasamy P and Kanagaraj S 2017 Biomarker selection from gene expression data for tumour categorization using bat algorithm. Methods 10 401–408
Chithambaram T and Perumal K 2017 Brain tumor segmentation using genetic algorithm and ann techniques. 2017 IEEE International Conference on Power, Control, Signals and Instrumentation Engineering (ICPCSI) 970 –982
Elbashir MK, Ezz M, Mohammed M, et al. 2019 Lightweight con- volutional neural network for breast cancer classification using RNA-seq gene expression data. IEEE Access 7 185338–185348
Jiao S, Zou Q, Guo H, et al. 2021 IITCA-RF: a random forest predictor for tumor T cell antigens. J. Transl. Med. 19 449
Kuo WJ, Chang RF, Chen DR, et al. 2001 Data mining with decision trees for diagnosis of breast tumor in medical ultrasonic images. Breast Cancer Res. Treat. 66 51–57
Li H, Courtois ET, Sengupta D, et al. 2017 Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat. Genet. 49 708–718
Lin P, Troup M and Ho JW 2017 CIDR: Ultrafast and accurate clustering through imputation for single-cell RNA-seq data. Genome Biol. 18 2–11
Liu HP, Wang D and Lai HM 2022 Can we infer tumor presence of single cell transcriptomes and their tumor of origin from bulk transcriptomes by machine learning. Comput. Struct. Biotechnol. J. 20 2672–2679
Mathew TE 2022 An optimized extremely randomized tree model for breast cancer classification. J. Theor. Appl. Informat. Technol. 100 5234–5246
Mothe R, Senapati B, Das R, et al. 2023 Identifying colorectal tumor for single cell RNA sequence using rectified linear unit with stochastic gradient descent. Procedia Comput. Sci. 218 189–198
Patel SA and Shah UV 2014 Tumor location and size identification in brain tissues using fuzzy c-clustering and artificial bee colony algorithm. Int. J. Engineer. Dev. Res. 2 2321–9939
Senapati B and Das R 2022 Single-cell RNA sequence data analysing using fuzzy de based clustering technique. Adv. Distributed Comput. Machine Learn. Proc. ICADCML 2022 427 479–487
SreeDevi KD, Karthikeyan P, Moorthy U, et al. 2022 Tumor detection on microarray data using grey wolf optimization with gain information. Math. Problems Eng. https://doi.org/10.1155/2022/4092404
Wang B, Zhu J, Pierson E, et al. 2017 Visualization and analysis of single cell RNA-seq data by kernel-based similarity learning. Nat. Methods 14 414–416
Acknowledgements
The research project was funded by DST and SERB (Grant number EEQ/2020/000104).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All of the authors agree that there are no conflicting interests.
Additional information
Corresponding editor: Mohit Kumar Jolly
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Senapati, B., Das, R. Tumor cell type and gene marker identification by single layer perceptron neural network on single-cell RNA sequence data. J Biosci 49, 47 (2024). https://doi.org/10.1007/s12038-023-00368-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12038-023-00368-w