Abstract
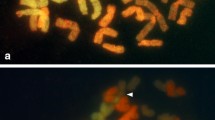
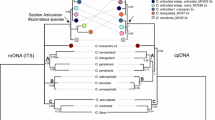
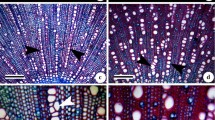
Aponogeton microphyllus, previously placed under the synonymy of A. undulatus, is recognized here as a distinct species based on morphology, chromosome number, and molecular phylogenetics (nuclear ribosomal internal transcribed (ITS) spacer region). Observations on the type and live specimens revealed morphological differences between the two species. Aponogeton microphyllus flowered regularly and set seeds. Aponogeton undulatus flowered rarely, did not set seeds, but showed formation of young plantlets on the inflorescence axis. Similarly, different chromosome numbers were recorded in Aponogeton microphyllus and the two forms of A. undulatus, viz., AF1 and AF2, which occur in distinct populations. Aponogeton microphyllus exhibited polysomaty with root-tip cells showing 2n=40, 42, and 44 chromosomes. The two forms of A. undulatus, AF1 and AF2, showed 2n=84 and 86 chromosomes, respectively. Based on the ITS data, both species occupied two separate clades. Plastid trnK intron region indicated a close relationship between both species. Our study suggests the need for comprehensive phylogenetic analyses of A. undulatus across its distribution range based on more advanced techniques such as high-throughput sequencing data to understand the A. undulatus species complex and to detect natural hybrids of this species.







Similar content being viewed by others
References
Ackerfield J and Wen J 2003 Evolution of Hedera (the Ivy Genus, Araliaceae): Insights from chloroplast DNA data. Int. J. Plant. Sci. 164 593–602
Bonfield JK and Whitwham A 2010 Gap5—editing the billion-fragment sequence assembly. Bioinformatics 26 1699–1703
van Bruggen HWE 1985 Monograph of the genus Aponogeton (Aponogetonaceae). Bioblioth. Bot. 33 1–76
Chen LU, Grimm GW, Wang QF, et al. 2015 A phylogeny and biogeography analysis for the Cape-Pondweed family Aponogetonaceae (Alismatales). Mol. Phylogenet. Evol. 82 111–117
Darriba D, Taboada GL, Doallo R, et al. 2012 jModelTest 2: more models, new heuristics and parallel computing. Nat. Methods 9 772
Doyle JJ 1992 Gene trees and species trees: molecular systematics as one character taxonomy. Syst. Biol. 17 144–163
Doyle JJ and Doyle JL 1987 A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19 11–15
de Wit HCD 1958 Aponogeton stachyosporous sp. nov. Meded. Bot. Tuinen Belmonte Arbor. Wageningen. 2 96
Dey D, Yadav SR and Devi N 2021 Rediscovery of Aponogeton lakhonensis A. Camus (Aponogetonaceae): a long-lost aquatic plant of India. J. Threat. Taxa 13 19632–19635
Fukai S, Hasegawa A and Goi M 2002 Polysomaty in Cymbidium. HortScience 37 1088–1091
Gaikwad SP 2000 Studies on biology of some endemic aquatic monocots. Ph.D. Thesis, Department of Botany, Shivaji University, Kolhapur, India
Gaikwad SP, Sardesai MM and Yadav SR 1998 Naturliche hybriden (?) zwischen Aponogeton appendiculatus HWE van Bruggen und A. undulatus Roxburgh und ihr Nutzen fur die Aquaristik. Aqua Planta 2 54–67
Gaikwad SP, Yadav SR, Gore RD, et al. 2014 Karyomorphological studies of Aponogeton appendiculatus Bruggen and Aponogeton crispus Thunb. J. Natn. Sci. Foundation Sri Lanka 42 163–167
Gargiulo GM, Vilardo I, Gemelli F, et al. 2020 Aneusomaty and polysomaty in Cymodocea nodosa (Ucria) Ascherson from Mediterranean Sea (Sicily, Italy). Aquat. Bot. 162 103206
Gargiulo GM, Balkkouri BEl, Crisafulli A, et al. 2022 Polysomaty and chromosome number variation in a population of Ceratophyllum demersum L. from Aquila Lake (Aspromonte Mountains, Calabria, Italy). Aquat. Bot. 180 103530
Hickey M and King C 2001 The Cambridge illustrated glossary of botanical terms (Cambridge: Cambridge University Press)
Kim S and Donoghue MJ 2008 Incongruence between cpDNA and nrITS trees indicates extensive hybridization within Eupersicaria (Polygonaceae). Am. J. Bot. 95 1122–1135
Larsson A 2014 AliView: a fast and lightweight alignment viewer and editor for large data sets. Bioinformatics 30 3276–3278
Les DH and Tippery NP 2013 In time and with water…the systematics of alismatid monocotyledons. in Early events in monocot evolution. Eds. P. Wilkinand S. J Mayo, (Cambridge, United Kingdom: Cambridge University Press). pp 118–164
Les DH and Philbrick CT 1993 Studies of hybridization and chromosome number variation in aquatic angiosperms: evolutionary implications. Aquat. Bot. 44 181–228
Les DH, Crawford DJ, Kimball RT, et al. 2003 Biogeography of discontinuously distributed hydrophytes: A molecular appraisal of intercontinental disjunctions. Int. J. Plant Sci. 164 917–932
Les DH, Moody ML and Jacobs SWL 2005 Phylogeny and systematics of Aponogeton (Aponogetonaceae): the Australian species. Syst. Bot. 30 503–519
POWO 2023 Plants of the World Online (http://www.plantsoftheworldonline.org)
Morales-Briones DF, Liston A and Tank DC 2018 Phylogenomic analyses reveal a deep history of hybridization and polyploidy in the Neotropical genus Lachemilla (Rosaceae). New Phytol. 218 1668–1684
Miller MA, Pfeiffer W and Schwartz T 2010 Creating the CIPRES Science Gateway for inference of large phylogenetic trees. 2010 Gateway Computing Environments Workshop (GCE) pp 1–8
Rieseberg LH and Soltis DE 1991 Phylogenetic consequences of cytoplasmic gene flow in plants. Evol. Trends Plants 5 65–83
Rice A, Glick L, Abadi S, et al. 2014 The Chromosome Counts Database (CCDB) – a community resource of plant chromosome numbers. New Phytol. 206 19–26
Ronquist F, Teslenko M, van der Mark P, et al. 2012 MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61 539–542
Roxburgh W 1832 Flora Indica; or, descriptions of Indian Plants 2 (Thacker & Co., Serampore, Calcutta and Allen & Co., Parbury, London) p 211
Swofford DL 2003 PAUP* Phylogenetic analysis using parsimony (*and other methods). Version 4 (Sinauer Associates, Sunderland, Massachusetts)
Sen S 1973 Polysomaty and its significance in Liliales. Cytologia 38 737–751
Stamatakis A 2014 RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30 1312–1313
Soltis DE and Kuzoff RK 1995 Discordance between nuclear and chloroplast phylogenies in the Heuchera Group (Saxifragaceae). Evolution 49 727–742
Surveswaran S, Gowda V and Sun M 2018 Using an integrated approach to identify cryptic species, divergence patterns and hybrid species in Asian ladies’ tresses orchids (Spiranthes, Orchidaceae). Mol. Phylogenet. Evol. 124 106–121
Than MMM, Pal A and Jha S 2011 Chromosome number and modal karyotype in a polysomatic endangered orchid, Bulbophyllum auricomum Lindl., the Royal Flower of Myanmar Plant Syst. Evol. 294 167–175
White TJ, Bruns T, Lee S and Taylor J 1990 Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics; in PCR protocols: A guide to methods and applications (Eds.) MA Innis, DH Gelfand, JJ Sninsky, et al. (Academic Press, San Diego) pp 315–322
Yadav SR 2017 A new species of Aponogeton (Aponogetonaceae) from India with critical notes on embryo morphology. Phytotaxa 328 83–89
Yadav SR and Gaikwad SP 2003 A revision of the Indian Aponogetonaceae. Bull. Bot. Surv. India. 45 39–76
Yadav SR, Patil VS, Gholave AR, et al. 2015 Aponogeton nateshii (Aponogetonaceae): a new species from India. Rheedea 25 9–13
Yadav SR 1995 Die hybrid zwishen Aponogeton decaryi Jumelle and Aponogeton satarensis Raghavan, Kulkami and Yadav. Aqua Planta 20 71–80
Acknowledgements
The authors are grateful to the Head, Department of Botany, Shivaji University, Kolhapur, for providing necessary research facilities. Thanks are due to Dr. Susanne S. Renner, Honorary Professor of Biology, Washington University in St. Louis, Missouri, for going through previous versions of the manuscript and improving its content. SS thanks Pooja Kumari Shukla for help in sequence data analysis. SRY thanks Mr. Pushpendra Katiyar, NBRI, Lucknow, for his help during the field tour. SRY is grateful to University Grants Commission (UGC), New Delhi, for the award of the BSR faculty fellowship. We thank the Director, Royal Botanic Gardens, Kew, for permitting us to publish the selected images.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors have no conflict of interest to declare.
Additional information
Corresponding editor: R Geeta
Rights and permissions
About this article
Cite this article
Chougule, R.N., Surveswaran, S., George, A. et al. Roxburgh was right: Aponogeton microphyllus and Aponogeton undulatus are distinct species. J Biosci 48, 53 (2023). https://doi.org/10.1007/s12038-023-00366-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12038-023-00366-y