Abstract
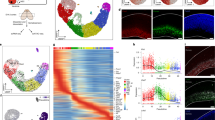
Multivalent binding of CTCF to variable DNA sequences is thought to underlie its ability to mediate diverse cellular functions. CTCF typically binds a 20 base-pair consensus DNA sequence, but the full diversity of CTCF binding sites (CBS) within the genome has not been interrogated. We assessed CTCF occupancy in cultured cortical neurons and observed surprisingly that ~ 22% of CBS lack the consensus CTCF motif. We report here that sequence diversity at most of these atypical CBS involves degeneracy at specific nucleotide positions within the consensus CTCF motif, which likely affect the binding of CTCF zinc fingers 6 and 7. This mode of atypical CTCF binding defines most CBS at gene promoters, as well as CBS that are dynamically altered during neural differentiation and following neuronal stimulation, revealing how atypical CTCF binding could influence gene activity. Dynamic CBS are distributed both within and outside loop anchors and TAD boundaries, suggesting both looping-dependent and independent roles for CTCF. Finally, we describe a second mode of atypical CTCF binding to DNA sequences that are completely unrelated to the consensus CTCF motif, which are enriched within the bodies of tissue-specific genes. These tissue-specific atypical CBS are also enriched in H3K27ac, which marks cis-regulatory elements within chromatin, including enhancers. Overall, these results indicate how atypical CBS could dynamically regulate gene activity patterns during differentiation, development, and in response to environmental cues.




Similar content being viewed by others
Data Availability
Original data produced for this manuscript, including raw and processed CTCF and SMC1 ChIP-seq data, are openly available from the NCBI GEO database at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE235972, reference number GSE235972.
References
Kim TH, Abdullaev ZK, Smith AD et al (2007) Analysis of the vertebrate insulator protein CTCF-binding sites in the human genome. Cell 128:1231–1245. https://doi.org/10.1016/j.cell.2006.12.048
Chen H, Tian Y, Shu W et al (2012) Comprehensive identification and annotation of cell type-specific and ubiquitous CTCF-binding sites in the human genome. PLoS ONE 7:e41374. https://doi.org/10.1371/journal.pone.0041374
Rubio ED, Reiss DJ, Welcsh PL et al (2008) CTCF physically links cohesin to chromatin. PNAS 105:8309–8314. https://doi.org/10.1073/pnas.0801273105
Pugacheva EM, Kubo N, Loukinov D et al (2020) CTCF mediates chromatin looping via N-terminal domain-dependent cohesin retention. PNAS 117:2020–2031. https://doi.org/10.1073/pnas.1911708117
Davidson IF, Bauer B, Goetz D et al (2019) DNA loop extrusion by human cohesin. Science 366:1338–1345. https://doi.org/10.1126/science.aaz3418
Kim Y, Shi Z, Zhang H et al (2019) Human cohesin compacts DNA by loop extrusion. Science 366:1345–1349. https://doi.org/10.1126/science.aaz4475
Hansen AS (2020) CTCF as a boundary factor for cohesin-mediated loop extrusion: evidence for a multi-step mechanism. Nucleus 11:132–148. https://doi.org/10.1080/19491034.2020.1782024
Nora EP, Goloborodko A, Valton AL et al (2017) Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169:930-944.e22. https://doi.org/10.1016/j.cell.2017.05.004
Aljahani A, Hua P, Karpinska MA et al (2022) Analysis of sub-kilobase chromatin topology reveals nano-scale regulatory interactions with variable dependence on cohesin and CTCF. Nat Commun 13:1–13. https://doi.org/10.1038/s41467-022-29696-5
Braccioli L, De Wit E (2019) CTCF: A Swiss-army knife for genome organization and transcription regulation. Essays Biochem 63:157–165. https://doi.org/10.1042/EBC20180069
Kim S, Yu NK, Kaang BK (2015) CTCF as a multifunctional protein in genome regulation and gene expression. Exp Mol Med 47:1–5. https://doi.org/10.1038/emm.2015.33
Kubo N, Ishii H, Xiong X et al (2021) Promoter-proximal CTCF binding promotes distal enhancer-dependent gene activation. Nat Struct Mol Biol 28:152–161. https://doi.org/10.1038/s41594-020-00539-5
Ohlsson R, Renkawitz R, Lobanenkov V (2001) CTCF is a uniquely versatile transcription regulator linked to epigenetics and disease. Trends Genet 17:520–527. https://doi.org/10.1016/S0168-9525(01)02366-6
Filippova GN, Qi C-F, Ulmer JE et al (2002) Tumor-associated zinc finger mutations in the CTCF transcription factor selectively alter its DNA-binding specificity. Cancer Res 62:48–52
Renda M, Baglivo I, Burgess-Beusse B et al (2007) Critical DNA binding interactions of the insulator protein CTCF: a small number of zinc fingers mediate strong binding, and a single finger-DNA interaction controls binding at imprinted loci. J Biol Chem 282:33336–33345. https://doi.org/10.1074/jbc.m706213200
Nakahashi H, Kwon KRK, Resch W et al (2013) A genome-wide map of CTCF multivalency redefines the CTCF code. Cell Rep 3:1678–1689. https://doi.org/10.1016/j.celrep.2013.04.024
Yin M, Wang J, Wang M et al (2017) Molecular mechanism of directional CTCF recognition of a diverse range of genomic sites. Cell Res 27:1365–1377. https://doi.org/10.1038/cr.2017.131
Guo J, Li N, Han J et al (2018) DNA recognition patterns of the multi-zinc-finger protein CTCF: a mutagenesis study. Acta Pharm Sin B 8:900–908. https://doi.org/10.1016/j.apsb.2018.08.007
Essien K, Vigneau S, Apreleva S et al (2009) CTCF binding site classes exhibit distinct evolutionary, genomic, epigenomic and transcriptomic features. Genome Biol 10:1–15. https://doi.org/10.1186/gb-2009-10-11-r131
Plasschaert RN, Vigneau S, Tempera I et al (2014) CTCF binding site sequence differences are associated with unique regulatory and functional trends during embryonic stem cell differentiation. Nucleic Acids Res 42:774–789. https://doi.org/10.1093/nar/gkt910
Arzate-Mejía RG, Recillas-Targa F, Corces VG (2018) Developing in 3D: the role of CTCF in cell differentiation. Development 145:1–16. https://doi.org/10.1242/dev.137729/48856
Moore JM, Rabaia NA, Smith LE et al (2012) Loss of maternal CTCF Is associated with peri-implantation lethality of Ctcf null embryos. PLoS ONE 7:e34915. https://doi.org/10.1371/journal.pone.0034915
Kim S, Yu NK, Shim KW et al (2018) Remote memory and cortical synaptic plasticity require neuronal CCCTC-binding factor (CTCF). J Neurosci 38:5042–5052. https://doi.org/10.1523/jneurosci.2738-17.2018
Il CD, Kim M, Kim S et al (2021) Conditional knock out of transcription factor CTCF in excitatory neurons induces cognitive deficiency. Mol Brain 14:1–4. https://doi.org/10.1186/s13041-020-00716-z
Sams DS, Nardone S, Getselter D et al (2016) Neuronal CTCF is necessary for basal and experience-dependent gene regulation, memory formation, and genomic structure of BDNF and Arc. Cell Rep 17:2418–2430. https://doi.org/10.1016/j.celrep.2016.11.004
Watson LA, Wang X, Elbert A et al (2014) Dual effect of CTCF loss on neuroprogenitor differentiation and survival. J Neurosci 34:2860–2870. https://doi.org/10.1523/jneurosci.3769-13.2014
Landt SG, Marinov GK, Kundaje A et al (2012) ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res 22:1813–1831. https://doi.org/10.1101/gr.136184.111
Zhang Y, Liu T, Meyer CA et al (2008) Model-based analysis of ChIP-Seq (MACS). Genome Biol 9:1–9. https://doi.org/10.1186/gb-2008-9-9-r137
Li Q, Brown JB, Huang H, Bickel PJ (2011) Measuring reproducibility of high-throughput experiments. Ann Appl Stat 5:1752–1779. https://doi.org/10.1214/11-aoas466
Bonev B, Mendelson Cohen N, Szabo Q et al (2017) Multiscale 3D genome rewiring during mouse neural development. Cell 171:557-572.e24. https://doi.org/10.1016/j.cell.2017.09.043
Durand NC, Shamim MS, Machol I et al (2016) Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst 3:95–98. https://doi.org/10.1016/j.cels.2016.07.002
Ma W, Noble WS, Bailey TL (2014) Motif-based analysis of large nucleotide data sets using MEME-ChIP. Nat Protoc 9:1428–1450. https://doi.org/10.1038/nprot.2014.083
Quinlan AR, Hall IM (2010) BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26:841–842. https://doi.org/10.1093/bioinformatics/btq033
Fornes O, Castro-Mondragon JA, Khan A et al (2020) JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res 48:D87–D92. https://doi.org/10.1093/nar/gkz1001
Ambrosini G, Groux R, Bucher P (2018) PWMScan: a fast tool for scanning entire genomes with a position-specific weight matrix. Bioinformatics 34:2483–2484. https://doi.org/10.1093/bioinformatics/bty127
Fu S, Wang Q, Moore JE et al (2018) Differential analysis of chromatin accessibility and histone modifications for predicting mouse developmental enhancers. Nucleic Acids Res 46:11184–11201. https://doi.org/10.1093/NAR/GKY753
Yu G, Wang LG, He QY (2015) ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31:2382–2383. https://doi.org/10.1093/bioinformatics/btv145
Ernst J (2017) Kellis M (2017) Chromatin-state discovery and genome annotation with ChromHMM. Nat Protoc 1212(12):2478–2492. https://doi.org/10.1038/nprot.2017.124
Abascal F, Acosta R, Addleman NJ et al (2020) Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 583:699–710. https://doi.org/10.1038/s41586-020-2493-4
Wu T, Hu E, Xu S et al (2021) clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innov 2:100141. https://doi.org/10.1016/j.xinn.2021.100141
Ramírez F, Dündar F, Diehl S et al (2014) deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res 42:W187–W191. https://doi.org/10.1093/NAR/GKU365
Kuhn RM, Haussler D, James Kent W (2013) The UCSC genome browser and associated tools. Brief Bioinform 14:144–161. https://doi.org/10.1093/bib/bbs038
Weth O, Renkawitz R (2011) CTCF function is modulated by neighboring DNA binding factors. Biochem Cell Biol 89:459–468. https://doi.org/10.1139/O11-033
Ohlsson R, Lobanenkov V, Klenova E (2010) Does CTCF mediate between nuclear organization and gene expression? BioEssays 32:37–50. https://doi.org/10.1002/bies.200900118
Ya Guo A, Xu Q, Canzio D et al (2015) CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162:900–910. https://doi.org/10.1016/j.cell.2015.07.038
Ren G, Jin W, Cui K et al (2017) CTCF-mediated enhancer-promoter interaction is a critical regulator of cell-to-cell variation of gene expression. Mol Cell 67:1049–1058. https://doi.org/10.1016/j.molcel.2017.08.026
Song Y, Ahn J, Suh Y et al (2013) Identification of novel tissue-specific genes by analysis of microarray databases: a human and mouse model. PLoS ONE 8:e64483. https://doi.org/10.1371/JOURNAL.PONE.0064483
Dubois-Chevalier J, Staels B, Lefebvre P, Eeckhoute J (2015) The ubiquitous transcription factor CTCF promotes lineage-specific epigenomic remodeling and establishment of transcriptional networks driving cell differentiation. Nucleus 6:15–18. https://doi.org/10.1080/19491034.2015.1004258
Balakrishnan SK, Witcher M, Berggren TW, Emerson BM (2012) Functional and molecular characterization of the role of CTCF in human embryonic stem cell biology. PLoS ONE 7:e42424. https://doi.org/10.1371/journal.pone.0042424
Phillips-Cremins JE, Sauria MEG, Sanyal A et al (2013) Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153:1281–1295. https://doi.org/10.1016/j.cell.2013.04.053
Nora EP, Caccianini L, Fudenberg G et al (2020) Molecular basis of CTCF binding polarity in genome folding. Nat Commun 11:1–13. https://doi.org/10.1038/s41467-020-19283-x
Beagan JA, Pastuzyn ED, Fernandez LR et al (2020) Three-dimensional genome restructuring across timescales of activity-induced neuronal gene expression. Nat Neurosci 23:707–717. https://doi.org/10.1038/s41593-020-0634-6
Davidson IF, Barth R, Zaczek M et al (2023) CTCF is a DNA-tension-dependent barrier to cohesin-mediated loop extrusion. Nature 616:822–827. https://doi.org/10.1038/s41586-023-05961-5
Corless S, Gilbert N (2016) Effects of DNA supercoiling on chromatin architecture. Biophys Rev 8:245–258. https://doi.org/10.1007/s12551-016-0210-1
Lee R, Kang MK, Kim YJ et al (2022) CTCF-mediated chromatin looping provides a topological framework for the formation of phase-separated transcriptional condensates. Nucleic Acids Res 50:207–226. https://doi.org/10.1093/NAR/GKAB1242
Chowdhary S, Kainth AS, Paracha S et al (2022) Inducible transcriptional condensates drive 3D genome reorganization in the heat shock response. Mol Cell 82:4386-4399.e7. https://doi.org/10.1016/j.molcell.2022.10.013
Hsieh THS, Cattoglio C, Slobodyanyuk E et al (2022) Enhancer–promoter interactions and transcription are largely maintained upon acute loss of CTCF, cohesin, WAPL or YY1. Nat Genet 54:1919–1932. https://doi.org/10.1038/s41588-022-01223-8
Acknowledgements
We would like to thank our colleges in the Madabhushi Lab, Lance Heady, Lahiri Konada, and Ilse Delint-Ramirez, for their helpful feedback on this manuscript.
Funding
This work was supported by NIMH under grant number MH120132 (RM) and CPRIT under grant number RP210041(MC).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation and data collection were performed by Morgan Crewe and Richard Rueda. Data analysis was performed by Morgan Crewe, Amir Segev, and Ram Madabhushi. The first draft of the manuscript was written by Morgan Crewe and Ram Madabhushi, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval
This study was performed in line with the principles outlined in the “Guide for the Care and Use of Laboratory Animals” and in accordance with all federal and state policies. Approval was granted by the UT Southwestern Institutional Care and Use Committee (IACUC).
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Crewe, M., Segev, A., Rueda, R. et al. Atypical Modes of CTCF Binding Facilitate Tissue-Specific and Neuronal Activity-Dependent Gene Expression States. Mol Neurobiol 61, 3240–3257 (2024). https://doi.org/10.1007/s12035-023-03762-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-023-03762-5