Abstract
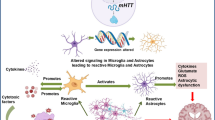
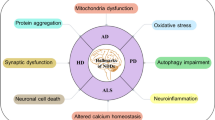
Neurodegenerative disorders are often a culmination of the accumulation of abnormally folded proteins and defective organelles. Autophagy is a process of removing these defective proteins, organelles, and harmful substances from the body, and it works to maintain homeostasis. If autophagic removal of defective proteins has interfered, it affects neuronal health. Some of the autophagic genes are specifically found to be associated with neurodegenerative phenotypes. Non-functional, mutated, or gene copies having silent mutations, often termed synonymous variants, might explain this. However, these synonymous variant which codes for exactly similar proteins have different translation rates, stability, and gene expression profiling. Hence, it would be interesting to study the pattern of synonymous variant usage. In the study, synonymous variant usage in various transcripts of autophagic genes ATG5, ATG7, ATG8A, ATG16, and ATG17/FIP200 reported to cause neurodegeneration (if dysregulated) is studied. These genes were analyzed for their synonymous variant usage; nucleotide composition; any possible nucleotide skew in a gene; physical properties of autophagic protein including GRAVY and AROMA; hydropathicity; instability index; and frequency of acidic, basic, neutral amino acids; and gene expression level. The study will help understand various evolutionary forces acting on these genes and the possible augmentation of a gene if showing unusual behavior.






Similar content being viewed by others
Data Availability
All the associated data is available within the manuscript.
References
Mizushima N, Yoshimori T, Ohsumi Y (2011) The role of Atg proteins in autophagosome formation. Annu Rev Cell Dev Biol 27:107–132
Jiang Q, Zhao L, Dai J, Wu Q (2012) Analysis of autophagy genes in microalgae: chlorella as a potential model to study mechanism of autophagy. PLoS ONE 7(7):e41826
Guo F, Liu X, Cai H, Le W (2018) Autophagy in neurodegenerative diseases: pathogenesis and therapy. Brain Pathol 28(1):3–13
Park H, Kang JH, Lee S (2020) Autophagy in neurodegenerative diseases: a hunter for aggregates. Int J Mol Sci 21(9):E3369
Yuk JM, Yoshimori T, Jo EK (2012) Autophagy and bacterial infectious diseases. Exp Mol Med 44(2):99–108
Ahmad L, Mostowy S, Sancho-Shimizu V (2018) Autophagy-virus interplay: from cell biology to human disease. Front Cell Dev Biol 6:155
Yun CW, Lee SH (2018) The roles of autophagy in cancer. Int J Mol Sci 19(11):E3466
Mulcahy Levy JM, Thorburn A (2020) Autophagy in cancer: moving from understanding mechanism to improving therapy responses in patients. Cell Death Differ 27(3):843–857
Bravo-San Pedro JM, Kroemer G, Galluzzi L (2017) Autophagy and mitophagy in cardiovascular disease. Circ Res 120(11):1812–1824
Schiattarella GG, Hill JA (2016) Therapeutic targeting of autophagy in cardiovascular disease. J Mol Cell Cardiol 95:86–93
Zhang Y, Sowers JR, Ren J (2018) Targeting autophagy in obesity: from pathophysiology to management. Nat Rev Endocrinol 14(6):356–376
Namkoong S, Cho CS, Semple I, Lee JH (2018) Autophagy dysregulation and obesity-associated pathologies. Mol Cells 41(1):3–10
Choi AMK, Ryter SW, Levine B (2013) Autophagy in human health and disease. N Engl J Med 368(7):651–662
Levine B, Mizushima N, Virgin HW (2011) Autophagy in immunity and inflammation. Nature 469(7330):323–335
Murrow L, Debnath J (2013) Autophagy as a stress-response and quality-control mechanism: implications for cell injury and human disease. Annu Rev Pathol 24(8):105–137
Cherra SJ, Chu CT (2008) Autophagy in neuroprotection and neurodegeneration: a question of balance. Future Neurol 3(3):309–323
Ravikumar B, Vacher C, Berger Z, Davies JE, Luo S, Oroz LG et al (2004) Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat Genet 36(6):585–595
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R et al (2006) Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441(7095):885–889
Komatsu M, Waguri S, Chiba T, Murata S, Iwata JI, Tanida I et al (2006) Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441(7095):880–4
Liang CC, Wang C, Peng X, Gan B, Guan JL (2010) Neural-specific deletion of FIP200 leads to cerebellar degeneration caused by increased neuronal death and axon degeneration. J Biol Chem 285(5):3499–3509
Heckmann BL, Teubner BJW, Boada-Romero E, Tummers B, Guy C, Fitzgerald P et al (2020) Noncanonical function of an autophagy protein prevents spontaneous Alzheimer’s disease. Sci Adv 6(33):eabb9036
Lipinski MM, Zheng B, Lu T, Yan Z, Py BF, Ng A et al (2010) Genome-wide analysis reveals mechanisms modulating autophagy in normal brain aging and in Alzheimer’s disease. Proc Natl Acad Sci U S A 107(32):14164–14169
Simonsen A, Cumming RC, Brech A, Isakson P, Schubert DR, Finley KD (2008) Promoting basal levels of autophagy in the nervous system enhances longevity and oxidant resistance in adult Drosophila. Autophagy 4(2):176–184
Quax TEF, Claassens NJ, Söll D, van der Oost J (2015) Codon bias as a means to fine-tune gene expression. Mol Cell 59(2):149–161
Tuller T, Carmi A, Vestsigian K, Navon S, Dorfan Y, Zaborske J et al (2010) An evolutionarily conserved mechanism for controlling the efficiency of protein translation. Cell 141(2):344–354
Purvis IJ, Bettany AJ, Santiago TC, Coggins JR, Duncan K, Eason R et al (1987) The efficiency of folding of some proteins is increased by controlled rates of translation in vivo. A hypothesis J Mol Biol 193(2):413–417
Pechmann S, Frydman J (2013) Evolutionary conservation of codon optimality reveals hidden signatures of cotranslational folding. Nat Struct Mol Biol 20(2):237–243
Gustafsson C, Govindarajan S, Minshull J (2004) Codon bias and heterologous protein expression. Trends Biotechnol 22(7):346–353
Gingold H, Tehler D, Christoffersen NR, Nielsen MM, Asmar F, Kooistra SM et al (2014) A dual program for translation regulation in cellular proliferation and differentiation. Cell 158(6):1281–1292
Neafsey DE, Galagan JE (2007) Positive selection for unpreferred codon usage in eukaryotic genomes. BMC Evol Biol 18(7):119
Arella D, Dilucca M, Giansanti A (2021) Codon usage bias and environmental adaptation in microbial organisms. Mol Genet Genomics 296(3):751–762
Galtier N, Roux C, Rousselle M, Romiguier J, Figuet E, Glémin S et al (2018) Codon usage bias in animals: disentangling the effects of natural selection, effective population size, and GC-biased gene conversion. Mol Biol Evol 35(5):1092–1103
Plotkin JB, Robins H, Levine AJ (2004) Tissue-specific codon usage and the expression of human genes. Proc Natl Acad Sci U S A 101(34):12588–12591
Payne BL, Alvarez-Ponce D (2019) Codon usage differences among genes expressed in different tissues of Drosophila melanogaster. Genome Biol Evol 11(4):1054–1065
Allen SR, Stewart RK, Rogers M, Ruiz IJ, Cohen E, Laederach A et al (2022) Distinct responses to rare codons in select Drosophila tissues. Elife 6(11):e76893
Goodman DB, Church GM, Kosuri S (2013) Causes and effects of N-terminal codon bias in bacterial genes. Science 342(6157):475–479
Miller JB, Brase LR, Ridge PG (2019) ExtRamp: a novel algorithm for extracting the ramp sequence based on the tRNA adaptation index or relative codon adaptiveness. Nucleic Acids Res 47(3):1123–1131
Esposito E, Weidemann DE, Rogers JM, Morton CM, Baybay EK, Chen J et al (2022) Mitotic checkpoint gene expression is tuned by codon usage bias. EMBO J 41(15):e107896
Fu H, Liang Y, Zhong X, Pan Z, Huang L, Zhang H et al (2020) Codon optimization with deep learning to enhance protein expression. Sci Rep 10(1):17617
Lorenzo MM, Nogales A, Chiem K, Blasco R, Martínez-Sobrido L (2022) Vaccinia virus attenuation by codon deoptimization of the A24R gene for vaccine development. Microbiol Spectr 10(3):e0027222
Ullah S, Ross TM (2022) Next generation live-attenuated influenza vaccine platforms. Expert Rev Vaccines 21(8):1097–1110
Khandia R, Ali Khan A, Alexiou A, Povetkin SN, Verevkina MN (2022) Codon usage analysis of pro-apoptotic Bim gene isoforms. J Alzheimers Dis 86(4):1711–1725. https://doi.org/10.3233/JAD-215691
Ingusci S, Verlengia G, Soukupova M, Zucchini S, Simonato M (2019) Gene therapy tools for brain diseases. Front Pharmacol 10:724
Karlin S, Mrázek J, Campbell AM (1998) Codon usages in different gene classes of the Escherichia coli genome. Mol Microbiol 29(6):1341–1355
Camiolo S, Sablok G, Porceddu A (2017) The evolutionary basis of translational accuracy in plants. G3 Bethesda 7(7):2363–73
Sharp PM, Li WH (1987) The codon adaptation index–a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res 15(3):1281–1295
Munjal A, Khandia R, Shende KK, Das J (2020) Mycobacterium lepromatosis genome exhibits unusually high CpG dinucleotide content and selection is key force in shaping codon usage. Infect Genet Evol 84:104399
Yang X, Luo X, Cai X (2014) Analysis of codon usage pattern in Taenia saginata based on a transcriptome dataset. Parasit Vectors 2(7):527
Mirsafian H, Mat Ripen A, Singh A, Teo PH, Merican AF, Mohamad SB (2014) A comparative analysis of synonymous codon usage bias pattern in human albumin superfamily. ScientificWorldJournal 2014:639682
Wright F (1990) The, “effective number of codons” used in a gene. Gene 87(1):23–29
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157(1):105–132
Khandia R, Alqahtani T, Alqahtani AM (2021) Genes common in primary immunodeficiencies and cancer display overrepresentation of codon CTG and dominant role of selection pressure in shaping codon usage. Biomedicines 9(8):1001
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, et al (2005) Protein identification and analysis tools on the ExPASy Server. In: Walker JM, editor. The Proteomics Protocols Handbook [Internet]. Totowa, NJ: Humana Press; 2005 [cited 2022 Apr 13]. p. 571–607. (Springer Protocols Handbooks). Available from: https://doi.org/10.1385/1-59259-890-0:571
Hammer O, Harper DAT, Ryan PD. PAST: paleontological statistics software package for education and data analysis. :9.
Piszter G, Kertész K, Bálint Z, Biró LP (2020) Stability and selective vapor sensing of structurally colored lepidopteran wings under humid conditions. Sensors (Basel) 20(11):E3258
Hernández-Barreto DF, Giraldo L, Moreno-Piraján JC (2020) Dataset on adsorption of phenol onto activated carbons: equilibrium, kinetics and mechanism of adsorption. Data Brief 32:106312
Moura G, Pinheiro M, Arrais J, Gomes AC, Carreto L, Freitas A et al (2007) Large scale comparative codon-pair context analysis unveils general rules that fine-tune evolution of mRNA primary structure. PLoS ONE 2(9):e847
Kim DJ, Kim J, Lee DH, Lee J, Woo HM (2022) DeepTESR: a deep learning framework to predict the degree of translational elongation short ramp for gene expression control. ACS Synth Biol 11(5):1719–1726
Miller JB, Meurs TE, Hodgman MW, Song B, Miller KN, Ebbert MTW et al (2022) The ramp atlas: facilitating tissue and cell-specific ramp sequence analyses through an intuitive web interface. NAR Genom Bioinform 4(2):Iqac039
Zahdeh F, Carmel L (2019) Nucleotide composition affects codon usage toward the 3’-end. PLoS ONE 14(12):e0225633
Zhang J, Wang M, Liu WQ, Zhou JH, Chen HT, Ma LN et al (2011) Analysis of codon usage and nucleotide composition bias in polioviruses. Virol J 8:146
Deka H, Chakraborty S (2014) Compositional constraint is the key force in shaping codon usage bias in hemagglutinin gene in H1N1 subtype of influenza A virus. Int J Genomics 2014:349139
Di Giallonardo F, Schlub TE, Shi M, Holmes EC (2017) Dinucleotide composition in animal RNA viruses is shaped more by virus family than by host species. J Virol 91(8):e02381-e2416
Alqahtani T, Khandia R, Puranik N, Alqahtani AM, Almikhlafi MA, Algahtany MA (2021) Leucine encoding codon TTG shows an inverse relationship with GC content in genes involved in neurodegeneration with iron accumulation. J Integr Neurosci 20(4):905–918
Barbhuiya PA, Uddin A, Chakraborty S (2019) Compositional properties and codon usage of TP73 gene family. Gene 30(683):159–168
Elhaik E (2022) Principal component analyses (PCA)-based findings in population genetic studies are highly biased and must be reevaluated. Sci Rep 12(1):14683
Lazaridis I, Nadel D, Rollefson G, Merrett DC, Rohland N, Mallick S et al (2016) Genomic insights into the origin of farming in the ancient Near East. Nature 536(7617):419–424
de Freire CCM, Palmisano G, Braconi CT, Cugola FR, Russo FB, Beltrão-Braga PC et al (2018) NS1 codon usage adaptation to humans in pandemic Zika virus. Mem Inst Oswaldo Cruz 113(5):e170385
Hassan S, Mahalingam V, Kumar V (2009) Synonymous codon usage analysis of thirty two mycobacteriophage genomes. Adv Bioinformatics 2009:316936
Mazumder TH, Chakraborty S (2015) Gaining insights into the codon usage patterns of TP53 gene across eight mammalian species. PLoS ONE 10(3):e0121709
Wang L, Xing H, Yuan Y, Wang X, Saeed M, Tao J et al (2018) Genome-wide analysis of codon usage bias in four sequenced cotton species. PLoS ONE 13(3):e0194372
Zhou Z, Dang Y, Zhou M, Li L, Yu CH, Fu J et al (2016) Codon usage is an important determinant of gene expression levels largely through its effects on transcription. Proc Natl Acad Sci U S A 113(41):E6117–E6125
Sahoo S, Das SS, Rakshit R (2019) Codon usage pattern and predicted gene expression in Arabidopsis thaliana. Gene 721S:100012
Yannai A, Katz S, Hershberg R (2018) The codon usage of lowly expressed genes is subject to natural selection. Genome Biol Evol 10(5):1237–1246
Deb B, Uddin A, Chakraborty S (2020) Codon usage pattern and its influencing factors in different genomes of hepadnaviruses. Arch Virol 165(3):557–570
Butt AM, Nasrullah I, Qamar R, Tong Y (2016) Evolution of codon usage in Zika virus genomes is host and vector specific. Emerg Microbes Infect 5(10):e107
Forcelloni S, Giansanti A (2020) Evolutionary forces and codon bias in different flavors of intrinsic disorder in the human proteome. J Mol Evol 88(2):164–178
Majeed A, Kaur H, Bhardwaj P (2020) Selection constraints determine preference for A/U-ending codons in Taxus contorta. Genome 63(4):215–224
Yap CC, Winckler B (2012) Harnessing the power of the endosome to regulate neural development. Neuron 74(3):440–451
Guo T, Nan Z, Miao C, Jin X, Yang W, Wang Z et al (2019) The autophagy-related gene Atg101 in Drosophila regulates both neuron and midgut homeostasis. J Biol Chem 294(14):5666–5676
Lehtonen Š, Sonninen TM, Wojciechowski S, Goldsteins G, Koistinaho J (2019) Dysfunction of cellular proteostasis in Parkinson’s disease. Front Neurosci 13:457
Djajadikerta A, Keshri S, Pavel M, Prestil R, Ryan L, Rubinsztein DC (2020) Autophagy induction as a therapeutic strategy for neurodegenerative diseases. J Mol Biol 432(8):2799–2821
Grishkevich V, Yanai I (2014) Gene length and expression level shape genomic novelties. Genome Res 24(9):1497–1503
Norkiene M, Gedvilaite A (2012) Influence of codon bias on heterologous production of human papillomavirus type 16 major structural protein L1 in yeast. ScientificWorldJournal 2012:979218
Chamary JV, Hurst LD (2005) Evidence for selection on synonymous mutations affecting stability of mRNA secondary structure in mammals. Genome Biol 6(9):R75
Shen X, Song S, Li C, Zhang J (2022) Synonymous mutations in representative yeast genes are mostly strongly non-neutral. Nature 606(7915):725–731
Plotkin JB, Kudla G (2011) Synonymous but not the same: the causes and consequences of codon bias. Nat Rev Genet 12(1):32–42
Davis JJ, Olsen GJ (2010) Modal codon usage: assessing the typical codon usage of a genome. Mol Biol Evol 27(4):800–810
Beutler E, Gelbart T, Han JH, Koziol JA, Beutler B (1989) Evolution of the genome and the genetic code: selection at the dinucleotide level by methylation and polyribonucleotide cleavage. Proc Natl Acad Sci U S A 86(1):192–196
Peifer M, Karro JE, von Grünberg HH (2008) Is there an acceleration of the CpG transition rate during the mammalian radiation? Bioinformatics 24(19):2157–2164
Blake RD, Hess ST, Nicholson-Tuell J (1992) The influence of nearest neighbors on the rate and pattern of spontaneous point mutations. J Mol Evol 34(3):189–200
Hodgman MW, Miller JB, Meurs TE, Kauwe JSK (2020) CUBAP: an interactive web portal for analyzing codon usage biases across populations. Nucleic Acids Res 48(19):11030–11039
Miller JB, McKinnon LM, Whiting MF, Kauwe JSK, Ridge PG (2020) Codon pairs are phylogenetically conserved: a comprehensive analysis of codon pairing conservation across the tree of life. PLoS ONE 15(5):e0232260
Irwin B, Heck JD, Hatfield GW (1995) Codon pair utilization biases influence translational elongation step times. J Biol Chem 270(39):22801–22806
Huang Y, Lin T, Lu L, Cai F, Lin J, Jiang YE et al (2021) Codon pair optimization (CPO): a software tool for synthetic gene design based on codon pair bias to improve the expression of recombinant proteins in Pichia pastoris. Microb Cell Fact 20(1):209
Kunec D, Osterrieder N, Trimpert J (2022) Synthetically recoded virus sCPD9 - a tool to accelerate SARS-CoV-2 research under biosafety level 2 conditions. Comput Struct Biotechnol J 20:4376–4380
Groenke N, Trimpert J, Merz S, Conradie AM, Wyler E, Zhang H et al (2020) Mechanism of virus attenuation by codon pair deoptimization. Cell Rep 31(4):107586
Miller JB, Hippen AA, Belyeu JR, Whiting MF, Ridge PG (2017) Missing something? Codon aversion as a new character system in phylogenetics. Cladistics 33(5):545–556
Brest P, Lapaquette P, Souidi M, Lebrigand K, Cesaro A, Vouret-Craviari V et al (2011) A synonymous variant in IRGM alters a binding site for miR-196 and causes deregulation of IRGM-dependent xenophagy in Crohn’s disease. Nat Genet 43(3):242–245
Behura SK, Severson DW (2012) Comparative analysis of codon usage bias and codon context patterns between dipteran and hymenopteran sequenced genomes. PLoS ONE 7(8):e43111
Ahmed W, Gupta S, Mukherjee I, Babu V, Singh R (2022) Comparative studies of codon usage profile of Anisakis simplex (Nematoda) and Carassius gibelio (Prussian carp). J Environ Biol 7(43):123–132
Wang P, Mao Y, Su Y, Wang J (2020) Comparative analysis of the codon usage patterns in two closely related Marsupenaeus species based on comparative transcriptomics
Fei YJ, Stoming TA, Kutlar A, Huisman TH, Stamatoyannopoulos G (1989) One form of inclusion body beta-thalassemia is due to a GAA––TAA mutation at codon 121 of the beta chain. Blood 73(4):1075–1077
Sørensen MA, Pedersen S (1991) Absolute in vivo translation rates of individual codons in Escherichia coli. The two glutamic acid codons GAA and GAG are translated with a threefold difference in rate. J Mol Biol 222(2):265–80
Malakar AK, Halder B, Paul P, Chakraborty S (2016) Cytochrome P450 genes in coronary artery diseases: codon usage analysis reveals genomic GC adaptation. Gene 590(1):35–43
Nath Choudhury M, Uddin A, Chakraborty S (2017) Codon usage bias and its influencing factors for Y-linked genes in human. Comput Biol Chem 69:77–86
Gupta SK, Ghosh TC (2001) Gene expressivity is the main factor in dictating the codon usage variation among the genes in Pseudomonas aeruginosa. Gene 273(1):63–70
Hou ZC, Yang N (2003) Factors affecting codon usage in Yersinia pestis. Sheng Wu Hua Xue Yu Sheng Wu Wu Li Xue Bao (Shanghai) 35(6):580–586
Moriyama EN, Powell JR (1998) Gene length and codon usage bias in Drosophila melanogaster, Saccharomyces cerevisiae and Escherichia coli. Nucleic Acids Res 26(13):3188–3193
Eyre-Walker A (1996) Synonymous codon bias is related to gene length in Escherichia coli: selection for translational accuracy? Mol Biol Evol 13(6):864–872
Puigbò P, Bravo IG, Garcia-Vallve S (2008) CAIcal: a combined set of tools to assess codon usage adaptation. Biol Direct 16(3):38
Trotta E (2013) Selection on codon bias in yeast: a transcriptional hypothesis. Nucleic Acids Res 41(20):9382–9395
Zhou T, Weems M, Wilke CO (2009) Translationally optimal codons associate with structurally sensitive sites in proteins. Mol Biol Evol 26(7):1571–1580
Uddin A, Mazumder TH, Barbhuiya PA, Chakraborty S (2020) Similarities and dissimilarities of codon usage in mitochondrial ATP genes among fishes, aves, and mammals. IUBMB Life 72(5):899–914
Acknowledgements
We acknowledge the respective universities and institutes for providing support for the study.
Funding
This work was funded by the researchers supporting project number (RSP-2023R339) King Saud University, Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
The manuscript was written through the contributions of all authors. All authors have approved the final version of the manuscript. RK, conception and supervision; RK, MP, AAK, and AA, writing—reviewing and editing; RK and AAK, interpretation of data and data curation; RK and MP, writing—reviewing and editing. IVR contributed significantly during revisions. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript and take responsibility of the work.
Corresponding authors
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khandia, R., Pandey, M.K., Rzhepakovsky, I.V. et al. Synonymous Codon Variant Analysis for Autophagic Genes Dysregulated in Neurodegeneration. Mol Neurobiol 60, 2252–2267 (2023). https://doi.org/10.1007/s12035-022-03081-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-022-03081-1