Abstract
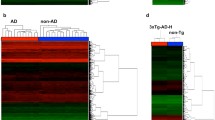
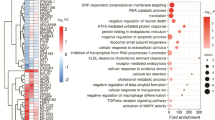
Sporadic early-onset Alzheimer’s disease (EOAD) and autosomal dominant Alzheimer’s disease (ADAD) provide the opportunity to investigate the physiopathological mechanisms in the absence of aging, present in late-onset forms. Frontotemporal dementia (FTD) causes early-onset dementia associated to tau or TDP43 protein deposits. A 15% of FTD cases are caused by mutations in C9orf72, GRN, or MAPT genes. Lymphoblastoid cell lines (LCLs) have been proposed as an alternative to brain tissue for studying earlier phases of neurodegenerative diseases. The aim of this study is to investigate the expression profile in EOAD, ADAD, and sporadic and genetic FTD (sFTD and gFTD, respectively), using brain tissue and LCLs. Sixty subjects of the following groups were included: EOAD, ADAD, sFTD, gFTD, and controls. Gene expression was analyzed with Clariom D microarray (Affymetrix). Brain tissue pairwise comparisons revealed six common differentially expressed genes (DEG) for all the patients’ groups compared with controls: RGS20, WIF1, HSPB1, EMP3, S100A11 and GFAP. Common up-regulated biological pathways were identified both in brain and LCLs (including inflammation and glial cell differentiation), while down-regulated pathways were detected mainly in brain tissue (including synaptic signaling, metabolism and mitochondrial dysfunction). CD163, ADAMTS9 and LIN7A gene expression disruption was validated by qPCR in brain tissue and NrCAM in LCLs in their respective group comparisons. In conclusion, our study highlights neuroinflammation, metabolism and synaptic signaling disturbances as common altered pathways in different AD and FTD forms. The use of LCLs might be appropriate for studying early immune system and inflammation, and some neural features in neurodegenerative dementias.








Similar content being viewed by others
Data Availability
The datasets generated during the current study are available in the NCBI’s GEO database under the accession number GSE195872.
References
Association A (2020) 2020 Alzheimer’s disease facts and figures. Alzheimers Dement 16:391–460. https://doi.org/10.1002/alz.12068
Falgàs N, Ruiz-Peris M, Pérez-Millan A et al (2020) Contribution of CSF biomarkers to early-onset Alzheimer’s disease and frontotemporal dementia neuroimaging signatures. Hum Brain Mapp 41:2004–2013. https://doi.org/10.1002/hbm.24925
Serrano-pozo A, Das S, Hyman BT (2021) APOE and Alzheimer’s disease: advances in genetics, pathophysiology, and therapeutic approaches. Lancet Neurol 20:68–80
Rascovsky K, Hodges JR, Knopman D et al (2011) Sensitivity of revised diagnostic criteria for the behavioural variant of frontotemporal dementia. Brain 134:2456–2477. https://doi.org/10.1093/brain/awr179
Gorno-Tempini ML, Hillis AE, Weintraub S et al (2011) Classification of primary progressive aphasia and its variants. Neurology 76:1006–1014. https://doi.org/10.1212/WNL.0b013e31821103e6
Lee EB, Porta S, Michael Baer G et al (2017) Expansion of the classification of FTLD-TDP: distinct pathology associated with rapidly progressive frontotemporal degeneration. Acta Neuropathol 134:65–78. https://doi.org/10.1007/s00401-017-1679-9
Froelich S, Houlden H, Pickering-Brown S et al (1998) Association of missense and 5’-splice-site mutations in tau with the inherited dementia FTDP-17. Nature 393:702–705
Baker M, Mackenzie IR, Pickering-Brown SM et al (2006) Mutations in progranulin cause tau-negative frontotemporal dementia linked to chromosome 17. Nature 442:916–919. https://doi.org/10.1038/nature05016
Dejesus-hernandez M, Mackenzie IR, Boeve BF et al (2012) Expanded GGGGCC hexanucleotide repeat in non-coding region of C9ORF72 causes chromosome 9p-linked frontotemporal dementia and amyotrophic lateral sclerosis. Neuron 72:245–256. https://doi.org/10.1016/j.neuron.2011.09.011.Expanded
Stopa EG, Tanis KQ, Miller MC et al (2018) Comparative transcriptomics of choroid plexus in Alzheimer’s disease, frontotemporal dementia and Huntington’s disease: implications for CSF homeostasis. Fluids Barriers CNS 15:18. https://doi.org/10.1186/s12987-018-0102-9
Mathys H, Davila-Velderrain J, Peng Z et al (2019) Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570:332–337. https://doi.org/10.1038/s41586-019-1195-2.Single-cell
Antonell A, Lladó A, Altirriba J et al (2013) A preliminary study of the whole-genome expression profile of sporadic and monogenic early-onset Alzheimer’s disease. Neurobiol Aging 34:1772–1778. https://doi.org/10.1016/j.neurobiolaging.2012.12.026
Martínez M, Inestrosa NC (2021) The transcriptional landscape of Alzheimer’s disease and its association with Wnt signaling pathway. Neurosci Biobehav Rev 128:454–466. https://doi.org/10.1016/j.neubiorev.2021.06.029
Patel H, Dobson RJB, Newhouse SJ (2019) A meta-analysis of Alzheimer’s disease brain transcriptomic data. J Alzheimers Dis 68:1635–1656. https://doi.org/10.3233/JAD-181085
Chen-Plotkin AS, Geser F, Plotkin JB et al (2008) Variations in the progranulin gene affect global gene expression in frontotemporal lobar degeneration. Hum Mol Genet 17:1349–1362. https://doi.org/10.1093/hmg/ddn023
Andrés-Benito P, Gelpi E, Povedano M et al (2019) Combined transcriptomics and proteomics in frontal cortex area 8 in frontotemporal lobar degeneration linked to C9ORF72 expansion. J Alzheimers Dis 68:1287–1307. https://doi.org/10.3233/JAD-181123
Andres-Benito P, Gelpi E, Povedano M et al (2018) Gene expression profile in frontal cortex in sporadic frontotemporal lobar degeneration-TDP. J Neuropathol Exp Neurol 77:608–627. https://doi.org/10.1093/jnen/nly037
Dezfulian M (2018) A new Alzheimer’s disease cell model using B cells to induce beta amyloid plaque formation and increase TNF alpha expression. Int Immunopharmacol 59:106–112. https://doi.org/10.1016/j.intimp.2018.04.012
Coskun P, Helguera P, Nemati Z et al (2017) Metabolic and growth rate alterations in lymphoblastic cell lines discriminate between Down syndrome and Alzheimer’s disease. J Alzheimers Dis 55:737–748. https://doi.org/10.3233/JAD-160278
Leuner K, Schulz K, Schütt T et al (2012) Peripheral mitochondrial dysfunction in Alzheimer’s disease: focus on lymphocytes. Mol Neurobiol 46:194–204. https://doi.org/10.1007/s12035-012-8300-y
Hadar A, Milanesi E, Squassina A et al (2016) RGS2 expression predicts amyloid-β sensitivity, MCI and Alzheimer’s disease: Genome-wide transcriptomic profiling and bioinformatics data mining. Transl Psychiatry 6:e909–e911. https://doi.org/10.1038/tp.2016.179
Gentleman R, Carey V, Huber W et al (2005) Bioinformatics and computational biology solutions using R and Bioconductor. Springer, New York
Irizarry RA, Hobbs B, Collin F et al (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264. https://doi.org/10.1007/978-1-4614-1347-9_15
Kauffmann A, Gentleman R, Huber W (2009) arrayQualityMetrics - a bioconductor package for quality assessment of microarray data. Bioinformatics 25:415–416. https://doi.org/10.1093/bioinformatics/btn647
Smyth GK (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3. https://doi.org/10.2202/1544-6115.1027
Subramaniana A, Tamayoa P, Moothaa VK, Mukherjeed S, Eberta BL, Gillettea MA, Paulovichg A, Pomeroyh SL, Goluba TR, Landera ES, JPM (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Jassal B, Matthews L, Viteri G et al (2020) The reactome pathway knowledgebase. Nucleic Acids Res 48:D498–D503. https://doi.org/10.1093/nar/gkz1031
Huntley RP, Binns D, Dimmer E et al (2009) QuickGO: a user tutorial for the web-based Gene Ontology browser. Database 2009:1–19. https://doi.org/10.1093/database/bap010
Wu T, Hu E, Xu S et al (2021) clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innov 2:100141. https://doi.org/10.1016/j.xinn.2021.100141
Gu Z, Hübschmann D (2021) simplifyEnrichment: an R/Bioconductor package for Clustering and Visualizing Functional Enrichment Results. bioRxiv 2020.10.27.312116
Begcevic I, Brinc D, Brown M et al (2018) Brain-related proteins as potential CSF biomarkers of Alzheimer’s disease: a targeted mass spectrometry approach. J Proteome 182:12–20. https://doi.org/10.1016/j.jprot.2018.04.027
Pey P, Pearce RKB, Kalaitzakis ME et al (2014) Phenotypic profile of alternative activation marker CD163 is different in Alzheimer’s and Parkinson’s disease. Acta Neuropathol Commun 2:1–14. https://doi.org/10.1186/2051-5960-2-21
Recabarren D, Alarcón M (2017) Gene networks in neurodegenerative disorders. Life Sci 183:83–97. https://doi.org/10.1016/j.lfs.2017.06.009
Lin CH, Lin E, Lane HY (2017) Genetic biomarkers on age-related cognitive decline. Front Psychiatry 8:1–9. https://doi.org/10.3389/fpsyt.2017.00247
Hondius DC, Van Nierop P, Li KW et al (2016) Profiling the human hippocampal proteome at all pathologic stages of Alzheimer’s disease. Alzheimers Dement 12:654–668. https://doi.org/10.1016/j.jalz.2015.11.002
Crist AM, Hinkle KM, Wang X et al (2021) Transcriptomic analysis to identify genes associated with selective hippocampal vulnerability in Alzheimer’s disease. Nat Commun 12:1–17. https://doi.org/10.1038/s41467-021-22399-3
Fan C, Chen K, Zhou J et al (2021) Systematic analysis to identify transcriptome-wide dysregulation of Alzheimer’s disease in genes and isoforms. Hum Genet 140:609–623. https://doi.org/10.1007/s00439-020-02230-7
Colangelo V, Schurr J, Ball MJ et al (2002) Gene expression profiling of 12633 genes in Alzheimer hippocampal CA1: Transcription and neurotrophic factor down-regulation and up-regulation of apoptotic and pro-inflammatory signaling. J Neurosci Res 70:462–473. https://doi.org/10.1002/jnr.10351
Patel H, Hodges AK, Curtis C et al (2019) Transcriptomic analysis of probable asymptomatic and symptomatic alzheimer brains. Brain Behav Immun 80:644–656. https://doi.org/10.1016/j.bbi.2019.05.009
Johnson TS, Xiang S, Dong T et al (2021) Combinatorial analyses reveal cellular composition changes have different impacts on transcriptomic changes of cell type specific genes in Alzheimer’s disease. Sci Rep 11:1–19. https://doi.org/10.1038/s41598-020-79740-x
Calabrò M, Rinaldi C, Santoro G, Crisafulli C (2021) The biological pathways of Alzheimer disease: a review. AIMS Neurosci 8:86–132. https://doi.org/10.3934/Neuroscience.2021005
Noori A, Mezlini AM, Hyman BT et al (2021) Systematic review and meta-analysis of human transcriptomics reveals neuroinflammation, deficient energy metabolism, and proteostasis failure across neurodegeneration. Neurobiol Dis Feb 149:105225. https://doi.org/10.1016/j.nbd.2020.105225
Piras IS, Krate J, Delvaux E et al (2019) Transcriptome changes in the Alzheimer’s disease middle temporal gyrus: importance of RNA metabolism and mitochondria-associated membrane genes. J Alzheimers Dis 70:691–713. https://doi.org/10.3233/JAD-181113
Weidling IW, Swerdlow RH (2020) Mitochondria in Alzheimer’s disease and their potential role in Alzheimer’s proteostasis. Exp Neurol 330:113321. https://doi.org/10.1016/j.expneurol.2020.113321
Yan MH, Wang X, Zhu X (2013) Mitochondrial defects and oxidative stress in Alzheimer disease and Parkinson disease. Free Radic Biol Med 62:90–101. https://doi.org/10.1016/j.freeradbiomed.2012.11.014
Grubman A, Chew G, Ouyang JF et al (2019) A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat Neurosci 22:2087–2097. https://doi.org/10.1038/s41593-019-0539-4
Canchi S, Raao B, Masliah D et al (2019) Integrating gene and protein expression reveals perturbed functional networks in Alzheimer’s disease. Cell Rep 28:1103–1116. https://doi.org/10.1016/j.celrep.2019.06.073
Jevtic S, Sengar AS, Salter MW, McLaurin JA (2017) The role of the immune system in Alzheimer disease: Etiology and treatment. Ageing Res Rev 40:84–94. https://doi.org/10.1016/j.arr.2017.08.005
Castillo E, Leon J, Mazzei G et al (2017) Comparative profiling of cortical gene expression in Alzheimer’s disease patients and mouse models demonstrates a link between amyloidosis and neuroinflammation. Sci Rep 7:1–16. https://doi.org/10.1038/s41598-017-17999-3
Liang WS, Dunckley T, Beach TG et al (2010) Neuronal gene expression in non-demented individuals with intermediate Alzheimer’s disease neuropathology. Neurobiol Aging 31:549–566. https://doi.org/10.1016/j.neurobiolaging.2008.05.013
Sebollela A, Freitas-Correa L, Oliveira FF et al (2012) Amyloid-β oligomers induce differential gene expression in adult human brain slices. J Biol Chem 287:7436–7445. https://doi.org/10.1074/jbc.M111.298471
Maes OC, Xu S, Yu B et al (2007) Transcriptional profiling of Alzheimer blood mononuclear cells by microarray. Neurobiol Aging 28:1795–1809. https://doi.org/10.1016/j.neurobiolaging.2006.08.004
Leandro GS, Evangelista AF, Lobo RR et al (2018) Changes in expression profiles revealed by transcriptomic analysis in peripheral blood mononuclear cells of Alzheimer’s disease patients. J Alzheimers Dis 66:1483–1495. https://doi.org/10.3233/JAD-170205
Dickson DW, Baker MC, Jackson JL et al (2019) Extensive transcriptomic study emphasizes importance of vesicular transport in C9orf72 expansion carriers. Acta Neuropathol Commun 7:1–21. https://doi.org/10.1186/s40478-019-0797-0
Brummer T, Müller SA, Pan-Montojo F et al (2019) NrCAM is a marker for substrate-selective activation of ADAM 10 in Alzheimer’s disease. EMBO Mol Med 11:1–20. https://doi.org/10.15252/emmm.201809695
Palluzzi F, Ferrari R, Graziano F et al (2017) A novel network analysis approach reveals DNA damage, oxidative stress and calcium/cAMP homeostasis-associated biomarkers in frontotemporal dementia. PLoS One 12:1–27. https://doi.org/10.1371/journal.pone.0185797
Santiago JA, Bottero V, Potashkin JA (2020) Transcriptomic and network analysis identifies shared and unique pathways across dementia spectrum disorders. Int J Mol Sci 21. https://doi.org/10.3390/ijms21062050
Elahi FM, Casaletto KB, La JR et al (2020) Plasma biomarkers of astrocytic and neuronal dysfunction in early- and late-onset Alzheimer’s disease. Alzheimers Dement 16:681–695. https://doi.org/10.1016/j.jalz.2019.09.004
Benussi A, Ashton NJ, Karikari TK et al (2020) Serum glial fibrillary acidic protein (GFAP) is a marker of disease severity in frontotemporal lobar degeneration. J Alzheimers Dis 77:1129–1141. https://doi.org/10.3233/JAD-200608
Zhu N, Santos-Santos M, Illán-Gala I et al (2021) Plasma glial fibrillary acidic protein and neurofilament light chain for the diagnostic and prognostic evaluation of frontotemporal dementia. Transl Neurodegener 10:50. https://doi.org/10.1186/s40035-021-00275-w
Humphries CE, Kohli MA, Nathanson L et al (2015) Integrated whole transcriptome and DNA methylation analysis identifies gene networks specific to late-onset Alzheimer’s disease. J Alzheimers Dis 44:977–987. https://doi.org/10.3233/JAD-141989
Birdsill AC, Walker DG, Lue LF et al (2011) Postmortem interval effect on RNA and gene expression in human brain tissue. Cell Tissue Bank 12:311–318. https://doi.org/10.1007/s10561-010-9210-8
Copois V, Bibeau F, Bascoul-Mollevi C et al (2007) Impact of RNA degradation on gene expression profiles: Assessment of different methods to reliably determine RNA quality. J Biotechnol 127:549–559. https://doi.org/10.1016/j.jbiotec.2006.07.032
Trabzuni D, Ryten M, Walker R et al (2011) Quality control parameters on a large dataset of regionally dissected human control brains for whole genome expression studies. J Neurochem 119:275–282. https://doi.org/10.1111/j.1471-4159.2011.07432.x
Durrenberger PF, Fernando S, Kashefi SN et al (2010) Effects of antemortem and postmortem variables on human brain mRNA quality: a brainNet Europe study. J Neuropathol Exp Neurol 69:70–81. https://doi.org/10.1097/NEN.0b013e3181c7e32f
Acknowledgements
The authors thank patients and their relatives for their participation in research.
Funding
This work was supported by Instituto de Salud Carlos III and Fondo Europeo de Desarrollo Regional (FEDER), Unión Europea, “Una manera de hacer Europa” (PI17/00670 to Dr Antonell, PI20/00448 to Dr Sánchez-Valle and PFIS grant (FI18/00121) to O. Ramos-Campoy); Departament de Salut de la Generalitat de Catalunya (PERIS 2016-2020, SLT002/16/00329 and PERIS 2019-2021, SLT008/18/00061).
Author information
Authors and Affiliations
Contributions
A. Antonell and R. Sánchez-Valle contributed to the study conception and design, data analysis, and manuscript writing. Material preparation, data collection, and analysis were performed by O. Ramos-Campoy. M. Ferrer, R. Gonzalo, and A. Pérez-Millán contributed to data analysis. The first draft of the manuscript was written by O. Ramos-Campoy, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics Approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the local Ethics Committee of the Hospital Clínic and Ethics committee of Neurological Tissue Bank-IDIBAPS-Hospital Clínic and Basque Biobank.
Consent to Participate
Informed consent was obtained from all individual participants included in the study.
Consent to Publish
The authors affirm that human research participants provided informed consent for publication of the images and results obtained.
Conflict of Interest
The authors declare no conflicts of interest related to this manuscript. RSV reports personal fees from Wave pharmaceuticals for attending Advisory board meetings, personal fees from Roche diagnostics, Janssen and Neuraxpharm for educational activities, and research grants to her institution from Biogen and Sage Therapeutics outside the submitted work.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Raquel Sánchez-Valle and Anna Antonell share senior and corresponding authorship.
Supplementary information
Online resource 1
(PDF 294 kb)
Online resource 2
(XLSX 40 kb)
Online resource 3
(XLSX 58 kb)
Online resource 4
(PDF 156 kb)
Online resource 5
(XLSX 59 kb)
Online resource 6
(XLSX 100 kb)
Online resource 7
(XLSX 32 kb)
Online resource 8
(XLSX 33 kb)
Online resource 9
(PDF 358 kb)

Online resource 10
(PNG 864 kb)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ramos-Campoy, O., Lladó, A., Bosch, B. et al. Differential Gene Expression in Sporadic and Genetic Forms of Alzheimer’s Disease and Frontotemporal Dementia in Brain Tissue and Lymphoblastoid Cell Lines. Mol Neurobiol 59, 6411–6428 (2022). https://doi.org/10.1007/s12035-022-02969-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-022-02969-2