Abstract
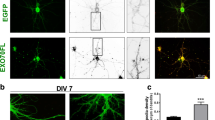
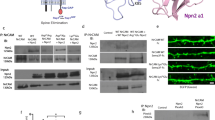
The Ring Finger Protein 10 [RNF10] is a novel synapse-to-nucleus signaling protein that specifically links activation of synaptic NMDA receptors to modulation of gene expression. RNF10 dissociation from the GluN2A subunit of the NMDA receptor represents the first step of its synaptonuclear transport and it is followed by an importin-dependent translocation into the nucleus. Here, we have identified protein kinase C [PKC]-dependent phosphorylation of RNF10 Ser31 as a key step for RNF10 detachment from NMDA receptor and its subsequent trafficking to the nucleus. We show that pSer31-RNF10 plays a role both in synaptonuclear signaling and in neuronal morphology. In particular, the prevention of Ser31 RNF10 phosphorylation induces a decrease in spine density, neuronal branching, and CREB signaling, while opposite effects are obtained by mimicking a stable RNF10 phosphorylation at Ser31. Overall, these results add novel information about the functional and structural role of synaptonuclear protein messengers in shaping dendritic architecture in hippocampal neurons.




Similar content being viewed by others
References
Fainzilber M, Budnik V, Segal RA, Kreutz MR (2011) From synapse to nucleus and back again--communication over distance within neurons. J Neurosci 31:16045–16048. https://doi.org/10.1523/JNEUROSCI.4006-11.2011
Ch’Ng TH, Uzgil B, Lin P, Avliyakulov NK, O’Dell TJ, Martin KC (2012) Activity-dependent transport of the transcriptional coactivator CRTC1 from synapse to nucleus. Cell 150:207–221. https://doi.org/10.1016/j.cell.2012.05.027
Herbst WA, Martin KC (2017) Regulated transport of signaling proteins from synapse to nucleus. Curr Opin Neurobiol 45:78–84. https://doi.org/10.1016/j.conb.2017.04.006
Lim AFY, Lim WL, Ch’ng TH (2017) Activity-dependent synapse to nucleus signaling. Neurobiol Learn Mem 138:78–84. https://doi.org/10.1016/j.nlm.2016.07.024
Karpova A, Mikhaylova M, Bera S, Bär J, Reddy PP, Behnisch T, Rankovic V, Spilker C et al (2013) Encoding and transducing the synaptic or extrasynaptic origin of NMDA receptor signals to the nucleus. Cell 152:1119–1133. https://doi.org/10.1016/j.cell.2013.02.002
Marcello E, Di Luca M, Gardoni F (2018) Synapse-to-nucleus communication: from developmental disorders to Alzheimer’s disease. Curr Opin Neurobiol 48:160–166. https://doi.org/10.1016/j.conb.2017.12.017
Dinamarca MC, Guzzetti F, Karpova A, Lim D, Mitro N, Musardo S, Mellone M, Marcello E et al (2016) Ring finger protein 10 is a novel synaptonuclear messenger encoding activation of NMDA receptors in hippocampus. Elife 5:e12430. https://doi.org/10.7554/eLife.12430
Lin J, Friesen MT, Bocangel P, Cheung D, Rawszer K, Wigle JT (2005) Characterization of mesenchyme homeobox 2 (MEOX2) transcription factor binding to RING finger protein 10. Mol Cell Biochem 275:75–84
Malik YS, Sheikh MA, Lai M, Cao R, Zhu X (2013) RING finger protein 10 regulates retinoic acid-induced neuronal differentiation and the cell cycle exit of P19 embryonic carcinoma cells. J Cell Biochem 114:2007–2015. https://doi.org/10.1002/jcb.24544
Piccoli G, Verpelli C, Tonna N, Romorini S, Alessio M, Nairn AC, Bachi A, Sala C (2007) Proteomic analysis of activity-dependent synaptic plasticity in hippocampal neurons. J Proteome Res 6:3203–3215. https://doi.org/10.1021/pr0701308
Gardoni F, Schrama LH, Kamal A, Gispen WH, Cattabeni F, Di Luca M (2001) Hippocampal synaptic plasticity involves competi- 616 tion between ca2+/calmodulin-dependent protein kinase II and 617 postsynaptic density 95 for binding to the NR2A subunit of the 618 NMDA receptor. J Neurosci 21:1501–1509
Shaner NC, Campbell RE, Steinbach PA, Giepmans BN, Palmer AE, Tsien RY (2004) Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat Biotechnol 22:1567–1572. https://doi.org/10.1038/nbt1037
Toullec D, Pianetti P, Coste H, Bellevergue P, Grand-Perret T, Ajakane M, Baudet V, Boissin P et al (1991) The bisindolylmaleimide GF 109203X is a potent and selective inhibitor of protein kinase C. J Biol Chem 1266:15771–15781
Robinson PJ (1992) Differential stimulation of protein kinase C activity by phorbol ester or calcium/phosphatidylserine in vitro and in intact synaptosomes. J Biol Chem 267:21637–21344
Dash PK, Hochner B, Kandel ER (1990) Injection of the cAMP-responsive element into the nucleus of Aplysia sensory neurons blocks long-term facilitation. Nature 345:718–721
Hardingham GE, Fukunaga Y, Bading H (2002) Extrasynaptic NMDARs oppose synaptic NMDARs by triggering CREB shut-off and cell death pathways. Nat Neurosci 5:405–414. https://doi.org/10.1038/nn835
Sheng M, McFadden G, Greenberg ME (1990) Membrane depolarization and calcium induce c-fos transcription via phosphorylation of transcription factor CREB. Neuron 4:571–582
Roberson ED, English JE, Adams JP, Selcher JC, Kondratick C, Sweatt JD (1999) The mitogen-activated protein kinase cascade couples PKA and PKC to cAM P response element binding protein phosphorylationin area CA1 of hippocampus. J Neurosci 19:4337–4348
Kanterewicz BI, Urban NN, McMahon DB, Norman ED, Giffen LJ, Favata MF, Scherle PA, Trzskos JM et al (2000 May 1) The extracellular signal-regulated kinase cascade is required for NMDA receptor-independent LTP in area CA1 but not area CA3 of the hippocampus. J Neurosci 20(9):3057–3066
English JE, Sweatt JD (1997) A requirement for the mitogen-activatedprotein kinase cascade in hippocampal long-term potentiation. J Biol Chem 272:19103–19106
Hotte M, Thuault S, Dineley KT, Hemmings HC Jr, Nairn AC, Jay TM (2007) Phosphorylation of CREB and DARPP-32 during late LTP at hippocampal to prefrontal cortex synapses in vivo. Synapse. 61(1):24–28
Spilker C, Nullmeier S, Grochowska KM, Schumacher A, Butnaru I, Macharadze T, Gomes GM, Yuanxiang P et al (2016) A Jacob/Nsmf gene knockout results in hippocampal dysplasia and impaired BDNF signaling in dendritogenesis. PLoS Genet 12:e1005907. https://doi.org/10.1371/journal.pgen.1005907
Proepper C, Johannsen S, Siebau S, Dahl J, Vaida B, Bockmann J, Kreutz MR, Gundelfinger ED et al (2017) Abelson interacting protein 1 (Abi-1) is essential for dendrite morphogenesis and synapse formation. EMBO J 26:1397–1409. https://doi.org/10.1038/sj.emboj.7601569
Finsterwald C, Fiumelli H, Cardinaux JR, Martin JL (2010) Regulation of dendritic development by BDNF requires activation of CRTC1 by glutamate. J Biol Chem 285:28587–28595. https://doi.org/10.1074/jbc.M110.125740
Li S, Zhang C, Takemori H, Zhou Y, Xiong ZQ (2009) TORC1 regulates activity-dependent CREB-target gene transcription and dendritic growth of developing cortical neurons. J Neurosci 29:2334–2343. https://doi.org/10.1523/JNEUROSCI.2296-08.2009
Li Z, Van Aelst L, Cline HT (2000) Rho GTPases regulate distinct aspects of dendritic arbor growth in Xenopus central neurons in vivo. Nat Neurosci 3:217–225. https://doi.org/10.1038/72920
Sin WC, Haas K, Ruthazer ES, Cline HT (2002) Dendrite growth increased by visual activity requires NMDA receptor and rho GTPases. Nature 419:475–480. https://doi.org/10.1038/nature00987
Mauceri D, Freitag HE, Oliveira AM, Bengtson CP, Bading H (2011) Nuclear calcium-VEGFD signaling controls maintenance of dendrite arborization necessary for memory formation. Neuron 71:117–130. https://doi.org/10.1016/j.neuron.2011.04.022
Kuo TY, Chen CY, Hsueh YP (2010) Bcl11A/CTIP1 mediates the effect of the glutamate receptor on axon branching and dendrite outgrowth. J Neurochem 114:1381–1392. https://doi.org/10.1111/j.1471-4159.2010.06852.x
Ewald RC, Van Keuren-Jensen KR, Aizenman CD, Cline HT (2008) Roles of NR2A and NR2B in the development of dendritic arbor morphology in vivo. J Neurosci 28:850–861. https://doi.org/10.1523/JNEUROSCI.5078-07.2008
Kannangara TS, Bostrom CA, Ratzlaff A, Thompson L, Cater RM, Gil-Mohapel J, Christie BR (2014) Deletion of the NMDA receptor GluN2A subunit significantly decreases dendritic growth in maturing dentate granule neurons. PLoS One 9:e103155. https://doi.org/10.1371/journal.pone.0103155
Balu D, Larson JR, Schmidt JV, Wirtshafter D, Yevtodiyenko A, Leonard AS (2016) Behavioral and physiological characterization of PKC-dependent phosphorylation in the Grin2a∆PKC mouse. Brain Res 1646(15–326):315–326. https://doi.org/10.1016/j.brainres.2016.06.022
Leonard AS, Hell JW (1997) Cyclic AMP-dependent protein kinase and protein kinase C phosphorylate N-methyl-D-aspartate receptors at different sites. J Biol Chem 272:12107–12115
Lin Y, Jover-Mengual T, Wong J, Bennett MV, Zukin RS (2006) PSD-95 and PKC converge in regulating NMDA receptor trafficking and gating. Proc Natl Acad Sci U S A 103:9902–19907. https://doi.org/10.1073/pnas.0609924104
Zheng X, Zhang L, Wang AP, Bennett MV, Zukin RS (1999) 611 Protein kinase C potentiation of N-methyl-D-aspartate receptor ac- 612 tivity is not mediated by phosphorylation of N-methyl-D-aspartate 613 receptor subunits. Proc Natl Acad Sci U S A 96:15262–15267
Acknowledgments
The authors are thankful to Anna Grassi, Francesco Tonolini, and Marta Ornaghi for their technical support.
Funding
DM is supported by DFG grants SFB1158 and FOR2325. DM is a member of the Excellence Cluster Cell Networks at Heidelberg University. This work was supported by the Italian Ministry of University and Research [PRIN 2015N4FKJ4 to MDL], Joint programme—Neurodegenerative disease research project STAD and by MIUR Progetto Eccellenza.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
ESM 1
(PDF 2243 kb)
Rights and permissions
About this article
Cite this article
Carrano, N., Samaddar, T., Brunialti, E. et al. The Synaptonuclear Messenger RNF10 Acts as an Architect of Neuronal Morphology. Mol Neurobiol 56, 7583–7593 (2019). https://doi.org/10.1007/s12035-019-1631-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-019-1631-1