Abstract
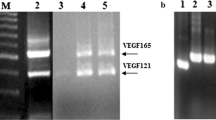
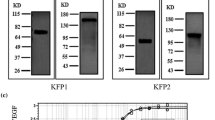
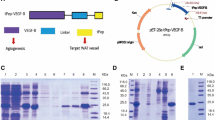
Vascular endothelial growth factor A165 (VEGF-A165) and VEGF receptor 2 (KDR) are important mediators of angiogenesis. We aimed to express the soluble KDR ligand-binding domain (sKDR1-3) and evaluate its interaction with the VEGF-A165 receptor-binding domain (VEGFA165-RBD). sKDR1-3 DNA was designed and subcloned into pPinkα-HC plasmid. The cassette was transfected into the Pichia pink™ 4 genome by homologous recombination. We optimized the expression of sKDR1-3 under the induction of different methanol concentrations. VEGFA165-RBD was expressed in E. coli BL21 harboring pET28a( +)─VEGFA165-RBD vector under induction with IPTG with/without lactose. Interaction and biological activity of sKDR1-3 and VEGFA165-RBD were investigated by ELISA and anti-proliferation tests. sKDR1-3 migrated on SDS-PAGE gel as a 35–180 kDa protein due to glycosylation. The relative expression level of sKDR1-3 under 1% methanol was higher than 0.5% and 4% methanol induction. IPTG and cysteine were suitable for induction and refolding of VEGFA165-RBD. 25 ng sKDR1-3 and 20 ng VEGFA165-RBD showed strong binding. sKDR1-3 bound to VEGFA165-RBD and VEGF-A165 with dissociation constants of 0.148 and 0.2 nM, respectively. 4–10 nM concentrations of sKDR1-3 inhibited the proliferation of HUVE cells induced by 5 nM VEGFA165-RBD. In consideration, sKDR1-3 in the nanomolar concentration range, is a promising anticancer drug to inhibit angiogenesis.







Similar content being viewed by others
Data Availability
The data that support the findings of this study are available on request from the corresponding author.
Abbreviations
- VEGF:
-
Vascular endothelial growth factor
- VEGFR-2:
-
VEGF receptor 2
- VEGFA165-RBD:
-
VEGF-A165 receptor binding domain
- VEGFA165-HBD:
-
VEGF-A165 heparin binding domain
- KDR:
-
Kinase domain receptor
- YPD:
-
Yeast Extract-Peptone-Dextrose Medium
- YNB:
-
Yeast nitrogen base
- Avastin:
-
An anti-VEGF-A165 monoclonal antibody
- AOX1 :
-
alcohol oxidase I
- ADE2:
-
Auxotrophic marker
- amp:
-
Ampicillin
- AmpR:
-
Ampicillin resistance
- A280 :
-
Absorbance at 280 nm
- BSA:
-
Bovine serum albumin
- PBS:
-
Phosphate buffered saline
- βME:
-
β-Mercaptoethanol
- BMGY:
-
Buffered glycerol complex medium
- BMMY:
-
Buffered methanol-complex medium
- CYC1 TT:
-
Transcription termination of CYC1
- DTT:
-
Dithiothreitol
- ELISA:
-
Enzyme-linked immunosorbent assay
- E. coli :
-
Escherichia coli
- EDTA:
-
Ethylenediaminetetraacetic acid
- G4S linker:
-
Gly4Ser
- HGF:
-
Hepatocyte growth factor
- His:
-
Histidine
- HRP:
-
Horse radish peroxidase
- HUVE:
-
Human umbilical vein endothelial cells
- Ig:
-
Immunoglobulin-like
- IgG:
-
Immunoglobulin class G
- IPTG:
-
Isopropyl β-D-1-thiogalactopyranoside
- IC50 :
-
Half-maximum inhibitory concentration
- Kaff :
-
Affinity constant
- KD:
-
Dissociation constant
- LBD:
-
Ligand binding domain
- LB:
-
Luria–Bertani
- Ni–NTA:
-
Nickel-nitrilotriaceticacid
- NP:
-
Normal physiological proteins
- nM:
-
Nanomolar
- ms:
-
Millisecond
- OD:
-
Optical density
- OD600 :
-
Optical density at 600 nm
- OD450 :
-
Optical density at 400 nm
- PBS:
-
Phosphate buffered saline
- PADE2 HC:
-
ADE2 promoter
- Pad:
-
Pichia Adenine Dropout
- PBST:
-
PBS containing tween 20
- PAOX1 :
-
AOX1 promoter
- PCR:
-
Polymerase chain reaction
- PDB:
-
Protein data bank
- pUC ori:
-
PUC origin of replication
- PIGF:
-
Placental growth factor
- RBD:
-
Receptor-binding domain
- SDS-PAGE:
-
Sodium dodecyl sulfatepolyacrylamide gel electrophoresis
- sKDR1-3:
-
Solube KDR domains 1–3
- TB:
-
Terrific broth
- TMB:
-
Tetramentylbenzidine
- TBST:
-
TBS containing tween 20 and BSA
- TRP2 gene:
-
Integration locus
- TBS:
-
Tris-buffered saline
- SXA :
-
β-D-Xylosidase/α-ʟ-arabinofuranosidase from the ruminal anaerobic bacterium Selenomonas ruminantium
References
Fairbrother, W. J., Champe, M. A., Christinger, H. W., Keyt, B. A., & Starovasnik, M. A. (1998). Solution structure of the heparin-binding domain of vascular endothelial growth factor. Structure, 6(5), 637–648.
Muller, Y. A., Christinger, H. W., Keyt, B. A., & de Vos, A. M. (1997). The crystal structure of vascular endothelial growth factor (VEGF) refined to 1.93 Å resolution: multiple copy flexibility and receptor binding. Structure, 5(10), 1325–1338.
Muller, Y. A., Li, B., Christinger, H. W., Wells, J. A., Cunningham, B. C., & De Vos, A. M. (1997). Vascular endothelial growth factor: Crystal structure and functional mapping of the kinase domain receptor binding site. Proceedings of the National Academy of Sciences, 94(14), 7192–7197.
Uciechowska-Kaczmarzyk, U., Babik, S., Zsila, F., Bojarski, K. K., Beke-Somfai, T., & Samsonov, S. A. (2018). Molecular dynamics-based model of VEGF-A and its heparin interactions. Journal of Molecular Graphics and Modelling, 82, 157–166.
Zajkowska, M., Lubowicka, E., Malinowski, P., Szmitkowski, M., & Ławicki, S. (2018). Plasma levels of VEGF-A, VEGF B, and VEGFR-1 and applicability of these parameters as tumor markers in diagnosis of breast cancer. Acta Biochimica Polonica, 65(4), 621–628.
Brozzo, M. S., Bjelić, S., Kisko, K., Schleier, T., Leppänen, V.-M., Alitalo, K., et al. (2012). Thermodynamic and structural description of allosterically regulated VEGFR-2 dimerization. Blood, The Journal of the American Society of Hematology, 119(7), 1781–1788.
Leppänen, V.-M., Prota, A. E., Jeltsch, M., Anisimov, A., Kalkkinen, N., Strandin, T., et al. (2010). Structural determinants of growth factor binding and specificity by VEGF receptor 2. Proceedings of the National Academy of Sciences, 107(6), 2425–2430.
Cao, W., Li, H., Zhang, J., Li, D., Acheampong, D. O., Chen, Z., et al. (2013). Periplasmic expression optimization of VEGFR2 D3 adopting response surface methodology: Antiangiogenic activity study. Protein Expression and Purification, 90(2), 55–66.
Huang, X., Gottstein, C., Brekken, R. A., & Thorpe, P. E. (1998). Expression of soluble VEGF receptor 2 and characterization of its binding by surface plasmon resonance. Biochemical and Biophysical Research Communications, 252(3), 643–648.
Kou, B., Li, Y., Zhang, L., Zhu, G., Wang, X., Li, Y., et al. (2004). In vivo inhibition of tumor angiogenesis by a soluble VEGFR-2 fragment. Experimental and Molecular Pathology, 76(2), 129–137.
Li, H., Cao, W., Chen, Z., Acheampong, D. O., Jin, H., Li, D., et al. (2013). The antiangiogenic activity of a soluble fragment of the VEGFR extracellular domain. Biomedicine & Pharmacotherapy, 67(7), 599–606.
Roeckl, W., Hecht, D., Sztajer, H., Waltenberger, J., Yayon, A., & Weich, H. A. (1998). Differential binding characteristics and cellular inhibition by soluble VEGF receptors 1 and 2. Experimental Cell Research, 241(1), 161–170.
Zhang, J., Li, H., Chen, W., Cao, P., & Wang, M. (2009). Preparation of extracellular domain 3 of human VEGF receptor-2 and the monitoring of its real-time binding to VEGF by biosensors. Biotechnology Progress, 25(6), 1703–1708.
Chandler, K. B., Leon, D. R., Meyer, R. D., Rahimi, N., & Costello, C. E. (2017). Site-specific N-glycosylation of endothelial cell receptor tyrosine kinase VEGFR-2. Journal of Proteome Research, 16(2), 677–688.
Ghavamipour, F., Shahangian, S. S., Sajedi, R. H., Arab, S. S., Mansouri, K., & Aghamaali, M. R. (2014). Development of a highly-potent anti-angiogenic VEGF 8–109 heterodimer by directed blocking of its VEGFR-2 binding site. The FEBS Journal, 281(19), 4479–4494.
Shahangian, S. S., Sajedi, R. H., Hasannia, S., Jalili, S., Mohammadi, M., Taghdir, M., et al. (2015). A conformation-based phage-display panning to screen neutralizing anti-VEGF VHHs with VEGFR2 mimicry behavior. International Journal of Biological Macromolecules, 77, 222–234.
Cao, Y., Sun, C., Huo, G., Wang, H., Wu, Y., Wang, F., et al. (2023). Novel hKDR mouse model depicts the antiangiogenesis and apoptosis-promoting effects of neutralizing antibodies targeting vascular endothelial growth factor receptor 2. Cancer Science, 114(1), 115–128.
Dehnavi, E., Siadat, S. O. R., Roudsari, M. F., & Khajeh, K. (2016). Cloning and high-level expression of β-xylosidase from Selenomonas ruminantium in Pichia pastoris by optimizing of pH, methanol concentration and temperature conditions. Protein Expression and Purification, 124, 55–61.
Beatty, J. D., Beatty, B. G., & Vlahos, W. G. (1987). Measurement of monoclonal antibody affinity by non-competitive enzyme immunoassay. Journal of Immunological Methods, 100(1–2), 173–179.
Sarabipour, S., Ballmer-Hofer, K., & Hristova, K. (2016). VEGFR-2 conformational switch in response to ligand binding. eLife, 5, e13876.
Povolotskaya, I. S., Kondrashov, F. A., Ledda, A., & Vlasov, P. K. (2012). Stop codons in bacteria are not selectively equivalent. Biology Direct, 7, 1–13.
Gruenberg, M. C., & TerAvest, M. A. (2023). A common inducer molecule enhances sugar utilization by Shewanella oneidensis MR-1. Journal of Industrial Microbiology and Biotechnology. https://doi.org/10.1093/jimb/kuad018
Kilikian, B. V., Suárez, I. D., Liria, C. W., & Gombert, A. K. (2000). Process strategies to improve heterologous protein production in Escherichia coli under lactose or IPTG induction. Process Biochemistry, 35, 1019–1025.
Leppänenc, K. A., Winklerb, F. K., & Ballmer-Hofera, K. (2011). Thermodynamic and structural description of allosterically regulated VEGF receptor 2 dimerization. Blood, the Journal of the American Society of Hematology, 119, 1781–1788.
Acknowledgements
The authors thank the universities of Kharazmi, Maragheh, and Tarbiat Modares of Iran.
Author information
Authors and Affiliations
Contributions
ZJ, ZF, MMAB, and RHS conceived the idea and developed the theory. ZF and ED performed the experiments, verified the methods, supervised the findings, and analyzed the data. ZJ and ZF designed the manuscript and provided critical feedback to help shape the manuscript. All authors approved the final version.
Corresponding author
Ethics declarations
Competing Interests
The authors have not disclosed any competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fathi, Z., Boojar, M.M.A., Sajedi, R.H. et al. Expression of VEGFR2 Ligand Binding Domain in Pichia pink™ 4 Cells and Evaluation of Its Interactions with VEGF-A165 Receptor Binding Domain. Mol Biotechnol (2024). https://doi.org/10.1007/s12033-024-01057-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-024-01057-1