Abstract
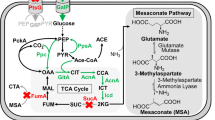
Cellular pool of malonyl-CoA in Escherichia coli is small, which impedes its utility for overproduction of natural products such as phenylpropanoids, polyketides, and flavonoids. In this study, we report the use of a new metabolic pathway to increase the malonyl-CoA concentration as a limiting metabolite in E. coli. For this purpose, the malonate/sodium symporter from Malonomonas rubra, and malonyl-CoA synthetase (MCS) from Bradyrhizobium japonicum were co-expressed in E. coli. This new pathway allows the cell to actively import malonate from the culture medium and to convert malonate and CoA to malonyl-CoA via an ATP-dependent ligation reaction. HPLC analysis confirmed elevated levels of malonyl-CoA and (2S)-naringenin as a malonyl-CoA-dependent metabolite, in E. coli. A 6.8-fold and more than 3.5-fold increase in (2S)-naringenin production were achieved in the engineered host in comparison with non-engineered E. coli and previously reported passive transport MatBMatC pathway, respectively. This observation suggests that using active transporters of malonate not only improves malonyl-CoA-dependent production but also makes it possible to harness low concentrations of malonate in culture media.
Graphical Abstract





Similar content being viewed by others
Data availability
The published article includes all data sets generated during this study.
Abbreviations
- CoA:
-
Coenzyme A
- MCoA:
-
Malonyl-coenzyme A
- MCS:
-
Malonyl-CoA synthetase
- ACC:
-
Acetyl-CoA carboxylase
References
Dixon, R. A., & Steele, C. L. (1999). Flavonoids and isoflavonoids–a gold mine for metabolic engineering. Trends in Plant Science, 4, 394–400.
Zha, W., Rubin-Pitel, S. B., Shao, Z., & Zhao, H. (2009). Improving cellular malonyl-CoA level in Escherichia coli via metabolic engineering. Metabolic Engineering, 11, 192–198.
Nielsen, J., & Keasling, J. D. (2011). Synergies between synthetic biology and metabolic engineering. Nature Biotechnology, 29, 693–695.
Koryakina, I., & Williams, G. J. (2011). Mutant malonyl-CoA synthetases with altered specificity for polyketide synthase extender unit generation. ChemBioChem, 12, 2289–2293.
Jang, M., Cai, L., Udeani, G. O., Slowing, K. V., Thomas, C. F., Beecher, C. W., Fong, H. H., Farnsworth, N. R., Kinghorn, A. D., & Mehta, R. G. (1997). Cancer chemopreventive activity of resveratrol, a natural product derived from grapes. Science, 275, 218–220.
Bradamante, S., Barenghi, L., & Villa, A. (2004). Cardiovascular protective effects of resveratrol. Cardiovascular Drug Reviews, 22, 169–188.
Lim, C. G., Fowler, Z. L., Hueller, T., Schaffer, S., & Koffas, M. A. (2011). High-yield resveratrol production in engineered Escherichia coli. Applied and Environmental Microbiology, 77, 3451–3460.
Wu, J., Zhou, T., Du, G., Zhou, J., & Chen, J. (2014). Modular optimization of heterologous pathways for de novo synthesis of (2S)-naringenin in Escherichia coli. PLoS ONE, 9, e101492.
Magnuson, K., Jackowski, S., Rock, C. O., & Cronan, J. E. (1993). Regulation of fatty acid biosynthesis in Escherichia coli. Microbiological Reviews, 57, 522–542.
Davis, M. S., Solbiati, J., & Cronan, J. E. (2000). Overproduction of acetyl-CoA carboxylase activity increases the rate of fatty acid biosynthesis in Escherichia coli. J BiolChem, 275, 28593–28598.
Liu, T., Vora, H., & Khosla, C. (2010). Quantitative analysis and engineering of fatty acid biosynthesis in E. coli. Metabolic Engineering, 12, 378–386.
Sáez-Sáez, J., Wang, G., Marella, E. R., Sudarsan, S., Pastor, M. C., & Borodina, I. (2020). Engineering the oleaginous yeast Yarrowia lipolytica for high-level resveratrol production. Metabolic Engineering, 62, 51–61.
Leonard, E., Lim, K.-H., Saw, P.-N., & Koffas, M. A. (2007). Engineering central metabolic pathways for high-level flavonoid production in Escherichia coli. Applied and Environmental Microbiology, 73, 3877–3886.
Wu, J., Zhou, L., Duan, X., Peng, H., Liu, S., Zhuang, Q., Pablo, C.-M., Fan, X., Ding, S., & Dong, M. (2021). Applied evolution: Dual dynamic regulations-based approaches in engineering intracellular malonyl-CoA availability. Metabolic Engineering, 67, 403–416.
Baba, T., Ara, T., Hasegawa, M., Takai, Y., Okumura, Y., Baba, M., Datsenko, K. A., Tomita, M., Wanner, B. L., & Mori, H. (2006). Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: The Keio collection. Molecular Systems Biology, 2(2006), 0008.
Kim, Y.-S. (2002). Malonate metabolism: Biochemistry, molecular biology, physiology, and industrial application. BMB Reports, 35, 443–451.
An, J. H., & Kim, Y. S. (1998). A gene cluster encoding malonyl-CoA decarboxylase (MatA), malonyl-CoA synthetase (MatB) and a putative dicarboxylate carrier protein (MatC) in Rhizobium trifolii. European Journal of Biochemistry, 257, 395–402.
Leonard, E., Yan, Y., Fowler, Z. L., Li, Z., Lim, C.-G., Lim, K.-H., & Koffas, M. A. (2008). Strain improvement of recombinant Escherichia coli for efficient production of plant flavonoids. Molecular Pharmaceutics, 5, 257–265.
Choi, O., Wu, C.-Z., Kang, S. Y., Ahn, J. S., Uhm, T.-B., & Hong, Y.-S. (2011). Biosynthesis of plant-specific phenylpropanoids by construction of an artificial biosynthetic pathway in Escherichia coli. Journal of Industrial Microbiology and Biotechnology, 38, 1657–1665.
Jeschek, M., Bahls, M. O., Schneider, V., Marlière, P., Ward, T. R., & Panke, S. (2017). Biotin-independent strains of Escherichia coli for enhanced streptavidin production. Metabolic Engineering, 40, 33–40.
Wu, J., Zhou, P., Zhang, X., & Dong, M. (2017). Efficient de novo synthesis of resveratrol by metabolically engineered Escherichia coli. Journal of Industrial Microbiology and Biotechnology, 44, 1083–1095.
Fang, Z., Jones, J. A., Zhou, J., & Koffas, M. A. (2018). Engineering Escherichia coli co-cultures for production of curcuminoids from glucose. Biotechnology Journal, 13, 1700576.
Liang, B., Sun, G., Wang, Z., Xiao, J., & Yang, J. (2019). Production of 3-hydroxypropionate using a novel malonyl-CoA-mediated biosynthetic pathway in genetically engineered E. coli strain. Green Chemistry, 21, 6103–6115.
Moore, S. J., Hleba, Y. B., Bischoff, S., Bell, D., Polizzi, K. M., & Freemont, P. S. (2021). Refactoring of a synthetic raspberry ketone pathway with EcoFlex. Microbial Cell Factories, 20, 1–11.
Zhou, S., Yuan, S.-F., Nair, P. H., Alper, H. S., Deng, Y., & Zhou, J. (2021). Development of a growth coupled and multi-layered dynamic regulation network balancing malonyl-CoA node to enhance (2S)-naringenin biosynthesis in Escherichia coli. Metabolic Engineering, 67, 41–52.
Schaffitzel, C., Berg, M., Dimroth, P., & Pos, K. M. (1998). Identification of an Na+-dependent malonate transporter of Malonomonas rubra and its dependence on two separate genes. Journal of Bacteriology, 180, 2689–2693.
Chen, H., Kim, H. U., & Weng, H. (2011). Malonyl-CoA synthetase, encoded by acyl activating enzyme13, is essential for growth and development of Arabidopsis. The Plant Cell, 23, 2247–2262.
Laemmli, U. K. (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227, 680–685.
Alishah, K., Asad, S., Khajeh, K., & Akbari, N. (2016). Utilizing intein-mediated protein cleaving for purification of uricase, a multimeric enzyme. Enzyme and Microbial Technology, 93, 92–98.
Kim, Y. S., & Bang, S. K. (1988). Assays for malonyl-coenzyme A synthase. Analytical Biochemistry, 170, 45–49.
Fowler, Z. L., Gikandi, W. W., & Koffas, M. A. (2009). Increased malonyl coenzyme A biosynthesis by tuning the Escherichia coli metabolic network and its application to flavanone production. Applied and Environmental Microbiology, 75, 5831–5839.
Asad, S., Dabirmanesh, B., & Khajeh, K. (2014). Phenol removal from refinery wastewater by mutant recombinant horseradish peroxidase. Biotechnology and Applied Biochemistry, 61, 226–229.
Korkina, L., Kostyuk, V., De Luca, C., & Pastore, S. (2011). Plant phenylpropanoids as emerging anti-inflammatory agents. Mini Reviews in Medicinal Chemistry, 11, 823–835.
Yu, O., & Jez, J. M. (2008). Nature’s assembly line: Biosynthesis of simple phenylpropanoids and polyketides. The Plant Journal, 54, 750–762.
Pandey, R. P., Parajuli, P., Koffas, M. A., & Sohng, J. K. (2016). Microbial production of natural and non-natural flavonoids: Pathway engineering, directed evolution and systems/synthetic biology. Biotechnology Advances, 34, 634–662.
Xu, P., Gu, Q., Wang, W., Wong, L., Bower, A. G., Collins, C. H., & Koffas, M. A. (2013). Modular optimization of multi-gene pathways for fatty acids production in E coli. Nature Communications, 4, 1409.
Galen, J. E., Wang, J. Y., Chinchilla, M., Vindurampulle, C., Vogel, J. E., Levy, H., Blackwelder, W. C., Pasetti, M. F., & Levine, M. M. (2010). A new generation of stable, nonantibiotic, low-copy-number plasmids improves immune responses to foreign antigens in Salmonella enterica serovar Typhi live vectors. Infection and Immunity, 78, 337–347.
Freitas, P., da Silva, D., Beluomini, M., da Silva, J., & Stradiotto, N. (2018). Determination of phenolic acids in sugarcane vinasse by HPLC with pulse amperometry. Applied and Environmental Microbiology. https://doi.org/10.1155/2018/4869487
Benke, M., Mermut, A., & Chatson, B. (1998). Carbon-13 CP/MAS NMR and DR-FTIR spectroscopic studies of sugarcane distillery waste. Canadian Journal of Soil Science, 78, 227–236.
Carrier, D. J., Bergeron, C., & Ramaswamy, S. (2012). Biorefinery co-products: Phytochemicals, primary metabolites and value-added biomass processing. Wiley.
Acknowledgements
We gratefully appreciate the Iran National Science Foundation (INSF) for financial support of this research (Project No. 94017574). Additionally, we would like to thank Prof. K.M. Pos (Goethe University Frankfurt, Germany) and Prof. J. Chen (Jiangnan University, Jiangsu, China) for providing us with the recombinant vectors and Mohammad Ranjbar for his constructive feedbacks over the manuscript.
Funding
This study was supported by Iran National Science Foundation (Grant No. 94017574).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Moteallehi-Ardakani, M.H., Asad, S., Marashi, SA. et al. Engineering a Novel Metabolic Pathway for Improving Cellular Malonyl-CoA Levels in Escherichia coli. Mol Biotechnol 65, 1508–1517 (2023). https://doi.org/10.1007/s12033-022-00635-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-022-00635-5