Abstract
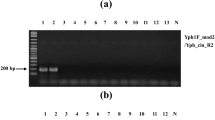
Collar rot disease caused by Sclerotium rolfsii is an economically important disease prevailing in all Amorphophallus growing areas. The pathogen propagules surviving in soil and planting material are the major sources of inoculum. A nested PCR assay has been developed for specific detection of S. rolfsii in soil and planting material. The PCR detection limit was 10 pg in conventional assay whereas 0.1 pg in nested assay. The primers designed were found to be highly specific and could be used for accurate identification of pathogen up to species level. The protocol was standardized for detection of the pathogen in artificially and naturally infected field samples.







Similar content being viewed by others
References
Hetterscheild, W. L. A., & Ittenbach, S. (1996). Everything you always wanted to know about Amorphophallus, but were afraid to stick your nose into. Aroideana, 19, 7–13.
Misra, R. S. (1997). Diseases of tuber crops in Northern and Eastern India. CTCRI technical series (Vol. 22, p. 27). Thiruvananthapuram: CTCRI.
Aycock, R. (1966). Stem rot and other diseases caused by Sclerotium rolfsii or the status of Rolf’s fungus after 70 years. North Carolina Agricultural Experiment Station Technical Bulletin, 174, 202.
Punja, Z. K. (1988). Sclerotium (Athelia) rolfsii, a pathogen of many plant species. Advances in Plant Pathology, 6, 523–534.
Misra, R. S., Sriram, S., Nedunchezhiyan, M., & Mohandas, C. (2003). Field and storage diseases of Amorphophallus and their management. Aroideana, 26, 101–112.
Leivens, B., Grauwet, T. J. M. A., Cammue, B. P. A., & Thomma, B. P. H. J. (2005). Recent developments in diagnostics of plant pathogens: a review. In: S.G. Pandalai (Ed.), Recent research developments in microbiology vol. 9 (pp. 57–79). Kerala, India.
Bonants, P., Weerdt, Hagenaar-de, Van Gent-Pelzer, M., Lacourt, I., Cooke, D., & Duncan, J. (1997). Detection and identification of Phytophthora fragarie Hickman by the polymerase chain reaction. European Journal of Plant Pathology, 103, 345–355.
McCartney, H. A., Foster, S. J., Fraaije, B. A., & Ward, E. (2003). Molecular diagnostics for fungal plant pathogens. Pest Management Science, 59, 129–1472. doi:10.1002/ps.575.
Henson, J. M., & French, R. (1993). The polymerase chain reaction and plant disease diagnosis. Annual Review of Phytopathology, 31, 81–109.
Ristaino, J. B., Madritch, M., & Trout, C. I. (1998). PCR amplification of ribosomal DNA for species identification in the plant pathogen genus Phytophthora. Applied and Environmental Microbiology, 64, 948–954.
Schaad, N. W., Frederick, R. D., Shaw, J., Schneider, W. L., Hickson, R., Petrillo, M. D., et al. (2003). Advances in molecular-based diagnostics in meeting crop biosecurity and phytosanitary issues. Annual Review of Phytopathology, 41, 305–324.
Willits, D. A., & Sherwood, J. E. (1999). Polymerase chain reaction detection of Ustilago hordei in leaves of susceptible and resistant barley varieties. Phytopathology, 89, 212–217.
Grote, D., Olmos, A., Kofoet, A., Tuser, J. J., Bertolini, E., & Cambra, M. (2002). Specific and sensitive detection of Phytophthora nicotianae by simple and nested PCR. European Journal of Plant Pathology, 108, 197–207.
Wallenhammer, A., & Arwidsson, O. (2001). Detection of Plasmodiophora brassicae by PCR in naturally infested soils. European Journal of Plant Pathology, 107, 313–321.
Mishra, A. K., Jeeva, M. L., Pravi, V., Misra, R. S., & Hegde, V. (2010). Rapid and sensitive detection of Phytophthora colacasiae associated with leaf blight of taro by species-specific polymerase chain reaction assay. Annals of Microbiology, 60, 209–215.
Mishra, A. K., Sharma, K., & Misra, R. S. (2008). Rapid and efficient method for the extraction of fungal and oomycetes genomic DNA. Gene, Genome and Genomics, 2, 57–59.
Jeeva, M. L., Sharma, K., Mishra, A. K., & Misra, R. S. (2008). Rapid extraction of genomic DNA from Sclerotium rolfsii causing collar rot of Amorphophallus. Genes, Genomes and Genomics, 2(1), 60–62.
Sambrook, J., Fritsch, R. F., & Maniatis, T. (1989). Molecular cloning: A laboratory manual (2nd ed.). New York: Cold Spring Harbor Laboratory Press.
White, T. J., Bruns, T., Lee, S., & Taylor, J. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In M. A. Innis, D. H. Gelfand, & J. J. Sininsky (Eds.), PCR protocols: A guide to methods and applications (pp. 315–322). San Diego: Academic press.
Hall, T. A. (1999). Bioedit: A user friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41, 95–98.
Zhang, Z., Schwartz, S., Wagner, L., & Miller, W. (2000). A greedy algorithm for aligning DNA sequences. Journal of Computational Biology, 7, 203–214.
Wang, H., Qi, M., & Cutler, A. J. (1993). A simple method of preparing plant samples for PCR. Nucleic Acids Research, 21, 4153–4154.
Ippolito, A., Schena, L., & Nigro, F. (2002). Detection of Phytophthora nicotianae and P. citrophthora in citrus roots and soils by nested PCR. European Journal of Plant Pathology, 108, 855–868.
Schena, L., Nigro, F., & Ippolito, A. (2002). Identification and detection of Rosellinia necatrix by conventional and real-time Scorpio-PCR. European Journal of Plant Pathology, 108, 355–366.
Weller, S. A., Elphinstone, J. G., Smith, N. C., Boonham, N., & Stead, D. E. (2000). Detection of Ralstonia solanacearum with a quantitative, multiplex, real time, fluorogenic (TaqMan) assay. Applied and Environment Microbiology, 66, 2853–2858.
Liew, E. C. Y., MacLean, D. J., & Irwin, J. A. G. (1998). Specific PCR based detection of Phytophthora medicaginis using the intergenic spacer regions of the ribosomal DNA. Mycological Research, 102, 73–80.
Schubert, R., Bahnweg, G., & Nechwatal, J. (1999). Detection and quantification of Phytophthora species which are associated with root to rot diseases in European deciduous forests by species specific polymerase chain reaction. European Journal of Plant Pathology, 29, 169–188.
Cooke, D. E. L., & Duncan, J. M. (1997). Phylogenetics analysis of Phytophthora species based on ITS 1 and ITS 2 sequences of the ribosomal RNA gene repeat. Mycological Research, 101, 667–677.
Lee, S. B., & Taylor, J. W. (1990). Isolation of DNA from fungal mycelia and single spores. In M. A. Innis, D. H. Gelfand, J. J. Sininsky, & T. J. White (Eds.), OCR protocols: A guide to methods and applications (pp. 282–287). San Diego: Academic press.
Bruns, T. D., White, T. J., & Taylor, J. W. (1991). Fungal molecular systematic. Annual Review of Ecology and Systematics, 22, 525–564.
Yao, C., Frederiksen, R. A., & Magill, C. W. (1992). Length heterogeneity in ITS 2 and the methylation status of CCGG and GCGC sites in the rRNA genes of the genus Peronosclerospora. Current Genetics, 22, 415–420.
Cullen, D. W., Lees, A. K., Toth, I. K., & Duncan, J. M. (2002). Detection of Colletotrichum coccodes from soil and potato tubers by conventional and quantitative real-time PCR. Plant Pathology, 51, 281–292.
Acknowledgments
The funding provided for research work by the National Fund for Basic Strategic and Frontier Application Research in Agricultural Sciences (NFBSFARA), ICAR, New Delhi, India, is gratefully acknowledged. The authors thank The Director, Central Tuber Crops Research Institute, Thiruvananthapuram for providing the infrastructure facilities. We are also grateful to the Indian Institute of Spices Research (Calicut, India) for providing the Phytophthora cultures and the College of Agriculture (Vellayani, India) and CTCRI (Sreekariyam, India) for providing the other fungal and bacterial cultures.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pravi, V., Jeeva, M.L. & Archana, P.V. Rapid and Sensitive Detection of Sclerotium rolfsii Associated with Collar Rot Disease of Amorphophallus paeoniifolius by Species-Specific Polymerase Chain Reaction Assay. Mol Biotechnol 56, 787–794 (2014). https://doi.org/10.1007/s12033-014-9757-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-014-9757-x