Abstract
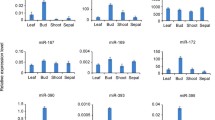
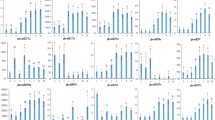
MicroRNAs (miRNAs) are a class of newly identified non-protein-coding small RNAs. miRNAs post-transcriptionally regulate the expression of more than 30% genes, which control many biological and metabolic progresses, including plant growth, development, and response to environmental stresses. However, no studies have been performed on miRNA expression in cotton, one of the most important fiber and economic crops. In this study, we employed quantitative real-time PCR (qRT-PCR) to detect and compare miRNA expressions in eight different cotton organs at different developmental stages. Our results showed that miRNAs were differentially expressed in different cotton organs, with certain classes expressed preferentially in an organ-specific manner. The miR-156 was highly expressed in cotyledon, whereas the miR-172 was highly expressed in young leaves at fruit branch, young flower buds, 0 day post-anthesis (DPA) ovules, and 0 DPA petals. We also found that miR-172 was not highly expressed in all parts of flowers. In contrast, miR-172 was highly expressed in petal but not in stamen and carpel. Interestingly, miR-162 was highly expressed in immature fiber, 2 DPA ovules, and mixtures of 0 DPA stamen and carpel, suggesting miR-162 may play a role in cotton fiber differentiation and development. Our previous study showed that miR-396 may target a fiber-related gene the callose synthase catalytic subunit. In this study, the miR-396 expression was observed in all eight cotton organs. This study identified the expression of miRNAs that may regulate the development of cotton and cotton fiber.

Similar content being viewed by others
References
Zhang, B. H., & Feng, R. (2000). Cotton resistance to insect and pest-resistant cotton. Beijing: Chinese Agricultural Science and Technology Press.
Stephens, S. G., & Mosley, M. E. (1974). Early domesticated cottons from archaeological sites in central coastal Peru. American Antiquity, 39, 109–122. doi:10.2307/279225.
IAC (1996). Cotton: Review of world situation, Monogram by International Advisory Committee, (Washington, DC).
Lee, J. J., Woodward, A. W., & Chen, Z. J. (2007). Gene expression changes and early events in cotton fibre development. Annals of Botany, 100, 1391–1401. doi:10.1093/aob/mcm232.
Taliercio, E. W., & Boykin, D. (2007). Analysis of gene expression in cotton fiber initials. BMC Plant Biology, 7, 22.
Wilkins, T. A., & Arpat, A. B. (2005). The cotton fiber transcriptome. Physiologia Plantarum, 124, 295–300. doi:10.1111/j.1399-3054.2005.00514.x.
Zhang, B. H., Pan, X. P., Guo, T. L., Wang, Q. L., & Anderson, T. A. (2005). Measuring gene flow in the cultivation of transgenic cotton (Gossypium hirsutum L.). Molecular Biotechnology, 31, 11–20. doi:10.1385/MB:31:1:011.
Zhang, B. H., Liu, F., Yao, C. B., & Wang, K. B. (2000). Recent progress in cotton biotechnology and genetic engineering in China. Current Science, 79, 37–44.
Haigler, C. H., Zhang, D., & Wilkerson, C. G. (2005). Biotechnological improvement of cotton fibre maturity. Physiologia Plantarum, 124, 285–294. doi:10.1111/j.1399-3054.2005.00480.x.
Zhang, B. H., Pan, X. P., Cobb, G. P., & Anderson, T. A. (2006). Plant microRNA: A small regulatory molecule with big impact. Developmental Biology, 289, 3–16. doi:10.1016/j.ydbio.2005.10.036.
Ambros, V., & Chen, X. M. (2007). The regulation of genes and genomes by small RNAs. Development, 134, 1635–1641. doi:10.1242/dev.002006.
Kidner, C. A., & Martienssen, R. A. (2005). The developmental role of microRNA in plants. Current Opinion in Plant Biology, 8, 38–44. doi:10.1016/j.pbi.2004.11.008.
Chen, X. M. (2005). MicroRNA biogenesis and function in plants. FEBS Letters, 579, 5923–5931. doi:10.1016/j.febslet.2005.07.071.
Chen, X. M. (2004). A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science, 303, 2022–2025. doi:10.1126/science.1088060.
Aukerman, M. J., & Sakai, H. (2003). Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. The Plant Cell, 15, 2730–2741. doi:10.1105/tpc.016238.
Griffiths-Jones, S., Saini, H. K., van Dongen, S., & Enright, A. J. (2008). miRBase: Tools for microRNA genomics. Nucleic Acids Research, 36, D154–D158. doi:10.1093/nar/gkm952.
Sunkar, R., Girke, T., Jain, P. K., & Zhu, J. K. (2005). Cloning and characterization of microRNAs from rice. The Plant Cell, 17, 1397–1411. doi:10.1105/tpc.105.031682.
Yao, Y. Y., Guo, G. G., Ni, Z. F., Sunkar, R., Du, J. K., Zhu, J. K., et al. (2007). Cloning and characterization of microRNAs from wheat (Triticum aestivum L.). Genome Biology, 8, R96.
Zhang, B. H., Pan, X. P., & Anderson, T. A. (2006). Identification of 188 conserved maize microRNAs and their targets. FEBS Letters, 580, 3753–3762. doi:10.1016/j.febslet.2006.05.063.
Zhang, B. H., Pan, X. P., & Stellwag, E. J. (2008). Identification of soybean microRNAs and their targets. Planta, 229, 161–182. doi:10.1007/s00425-008-0818-x.
Zhang, B. H., Wang, Q. L., & Pan, X. P. (2007). MicroRNAs and their regulatory roles in animals and plants. Journal of Cellular Physiology, 210, 279–289. doi:10.1002/jcp.20869.
Zhang, B. H., Pan, X. P., Wang, Q. L., Cobb, G. P., & Anderson, T. A. (2006). Computational identification of microRNAs and their targets. Computational Biology and Chemistry, 30, 395–407. doi:10.1016/j.compbiolchem.2006.08.006.
Floyd, S. K., & Bowman, J. L. (2004). Gene regulation: Ancient microRNA target sequences in plants. Nature, 428, 485–486. doi:10.1038/428485a.
Zhang, B. H., Pan, X. P., Cannon, C. H., Cobb, G. P., & Anderson, T. A. (2006). Conservation and divergence of plant microRNA genes. The Plant Journal, 46, 243–259. doi:10.1111/j.1365-313X.2006.02697.x.
Zhang, B. H., Wang, Q. L., Wang, K. B., Pan, X. P., Liu, F., Guo, T. L., et al. (2007). Identification of cotton microRNAs and their targets. Gene, 397, 26–37. doi:10.1016/j.gene.2007.03.020.
Zhang, B. H., Pan, X. P., Wang, Q. L., Cobb, G. P., & Anderson, T. A. (2005). Identification and characterization of new plant microRNAs using EST analysis. Cell Research, 15, 336–360. doi:10.1038/sj.cr.7290302.
Lauter, N., Kampani, A., Carlson, S., Goebel, M., & Moose, S. P. (2005). microRNA172 down-regulates glossy15 to promote vegetative phase change in maize. Proceedings of the National Academy of Sciences of the United States of America, 102, 9412–9417. doi:10.1073/pnas.0503927102.
Chen, C. F., Ridzon, D. A., Broomer, A. J., Zhou, Z. H., Lee, D. H., Nguyen, J. T., et al. (2005). Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Research, 33, e179. doi:10.1093/nar/gni178.
Schwab, R., Palatnik, J. F., Riester, M., Schommer, C., Schmid, M., & Weigel, D. (2005). Specific effects of microRNAs on the plant transcriptome. Developmental Cell, 8, 517–527. doi:10.1016/j.devcel.2005.01.018.
Kurihara, Y., & Watanabe, Y. (2004). Arabidopsis micro-RNA biogenesis through Dicer-like 1 protein functions. Proceedings of the National Academy of Sciences of the United States of America, 101, 12753–12758. doi:10.1073/pnas.0403115101.
Tang, G. L., Reinhart, B. J., Bartel, D. P., & Zamore, P. D. (2003). A biochemical framework for RNA silencing in plants. Genes & Development, 17, 49–63. doi:10.1101/gad.1048103.
Park, W., Li, J. J., Song, R. T., Messing, J., & Chen, X. M. (2002). CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana. Current Biology, 12, 1484–1495. doi:10.1016/S0960-9822(02)01017-5.
Xie, Z. X., Kasschau, K. D., & Carrington, J. C. (2003). Negative feedback regulation of Dicer-like1 in Arabidopsis by microRNA-guided mRNA degradation. Current Biology, 13, 784–789. doi:10.1016/S0960-9822(03)00281-1.
Liu, B., Li, P. C., Li, X., Liu, C. Y., Cao, S. Y., Chu, C. C., et al. (2005). Loss of function of OsDCL1 affects microRNA accumulation and causes developmental defects in rice. Plant Physiology, 139, 296–305. doi:10.1104/pp.105.063420.
Cui, X. J., Shin, H. S., Song, C., Laosinchai, W., Amano, Y., & Brown, R. M. (2001). A putative plant homolog of the yeast beta-1, 3-glucan synthase subunit FKS1 from cotton (Gossypium hirsutum L.) fibers. Planta, 213, 223–230. doi:10.1007/s004250000496.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, B., Pan, X. Expression of MicroRNAs in Cotton. Mol Biotechnol 42, 269–274 (2009). https://doi.org/10.1007/s12033-009-9163-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-009-9163-y