Abstract
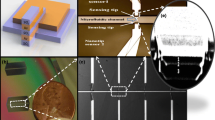
The development and critical evaluation of new technologies for identifying genetic polymorphisms will rapidly accelerate the discovery and diagnosis of disease-related genes. We report a novel way for distinguishing a new class of human DNA polymorphisms, short insertion/deletion polymorphisms (indels). A sensor with cylindrical pores named channel glass in combination with tandem hybridization, which uses a 5′-fluorescent labeled stacking probe and microarray-based short allele-specific oligonucleotide (capture probe) was investigated. This methodology allows indels to be detected individually and rapidly with small quantities of target DNA. This establishes a reliable quantitative test. Approaches for simultaneously hybridizing different targets to arrayed probes, designed to detect various indels in parallel, were examined. Five markers were consistently detected in a single hybridization. Possible factors impeding the hybridization reaction process are discussed.





Similar content being viewed by others
References
Ramsay, G. (1998) DNA chips: State-of-the art. Nature Biotechnology, 16, 40–44.
Dearlove, A. M. (2002). High throughput genotyping technologies. Briefings in Functional Genomic & Proteomics, 1, 139–150.
Matsuzaki1, H., Loi, H., Dong, S., Tsai, Y., Fang, J., Law, J., Di, X., Liu, W., Yang, G., Liu, G., Huang, J., Kennedy, G. C., Ryder, T. B., Marcus, G. A., Sean Walsh, P. S., Shriver, M. D., Puck, J. P., Jones, K. W., & Mei1, R. (2004). Parallel genotyping of over 10,000 SNPs using a one-primer assay on a high-density oligonucleotide array. Genome Research, 14, 414–425.
Stephens, M., Sloan, J. S., Robertson, P. D., Scheet, P., & Nickerson, D. A. (2006). Automating sequence-based detection and genotyping of SNPs from diploid samples. Nature Genetics, 38, 375–381.
Drmanac, S., Kita, D., Labat, I., Hauser, B., Schmidt, C., & Burczak, J. D. (1998). Accurate sequencing by hybridization for DNA diagnostics and individual genomics. Nature Biotechnology, 16, 54–58.
Hinds, D. A., Stuve, L. L., Nilsen, G. B., Halperin, E., Eskin, E., Ballinger, D. G., Frazer, K. A., & Cox, D.R. (2005). Whole-genome patterns of common DNA variation in three human populations. Science, 307, 1072–1079.
Pennisi, E. (2001). The human genome. Science, 291, 1177–1180.
Weber, J. L., David, D., Heil, J., Fan, Y., Zhao, C., & Marth, G. (2002). Human diallelic insertion/deletion polymorphisms. American Journal of Human Genetics, 71, 854–862.
Nickerson, D. A., Whitehurst, C., Boysen, C., Charmley, P., Kaiser, R., & Hood, L. (1992). Identification of clusters of bialleic polymorphic sequence-tagged sites (pSTSs) that generate highly informative and automatable markers for genetic linkage mapping. Genomics, 12, 377–387.
Kruglyak, L. (1997). What is significant in whole-genome linkage disequilibrium studies? American Journal of Human Genetics, 61, 810–812.
Lupski, J. R., Roth, J. R., & Weinstock, G. M. (1996). Chromosomal duplicationsin bacteria, fruit flies, and humans. American Journal of Human Genetics, 58, 21–27.
Cooper, D. N., & Krawczak, M. (1991). Mechanisms of insertional mutagenesis in human genes causing genetic disease. Human Genetics, 87, 409–415.
Guo, Z., Guifoyle, R. A., Thiel, A. J., Wang, R., & Smith, L. M. (1994). Direct fluorescence analysis of genetic polymorphisms by hybridization with oligonucleotide arrays on glass supports. Nucleic Acids Research, 22, 327–333.
Proudnikov, D., Timofeev, E., & Mirzabekov, A. (1998). Immobilization of DNA in polyacrylamide gel for the manufacture of DNA and DNA oligonucleotide microchips. Analytical Biochemistry, 259, 34–41.
Benoit, V., Steel, A., Torres, M., Yu, Y.-Y., Yang, H., & Cooper, J. (2001). Evaluation of three-dimensional microchannel glass biochips for multiplex nucleic acid fluorescence hybridization assays. Analytical Chemistry, 73, 2412–2420.
Pirrung, M. C. (2002). How to make a DNA chip. Angewandte Chemie International Edition, 41, 1276–1289.
Cheek, J., Steel, A. B., Torres, M. P., Yu, Y.-Y., & Yang, H. (2001). Chemiluminescence detection for hyrbidization assays on the flow-thru chip, a three-dimensional microchannel biochip. Analytical Chemistry, 73, 5777–5783.
Bessueille, F., Dugas, V., Vikulov, V., Cloarec, J. P., Souteyrand, E., & Martin, J. R. (2005). Assessment of porous silicon substrate for well-characterised sensitive DNA chip implement. Biosensors & Bioelectronics, 21, 908–916.
Wu, Y., de Kievit, P., Vahlkamp, L., Pijnenburg, D., Smit, M., Dankers, M., Melchers, D., Stax, M., Boender, P. J., Ingham, C., Bastiaensen, N., de Wijn, R., van Alewijk, D., van Damme, H., Raap, A. K, Chan, A. B., & van Beuningen R. (2004). Quantitative assessment of a novel flow-through porous microarray for the rapid analysis of gene expression profiles. Nucleic Acids Research, 32, e123.
Liu, Y., & Rauch, C. B. (2003). DNA probe attachment on plastic surfaces and microfluidic hybridization array channel devices with sample oscillation. Analytical Biochemistry, 317, 76–84.
Kirov, G., Nikolov, I., Georgieva, L., Moskvina, L., Owen, M. J., & O’Donovan, M. C. (2006). Pooled DNA genotyping on affymetrix SNP genotyping arrays. BMC Genomics, 7, 27.
Beattie, K. L., Beattie, W.G., Meng, L., Turner, S., Bishop, C., Dao, D., Coral, R., Smith, D., & McIntyre, P. (1995). Advances in genosensor research. Clinical Chemistry, 41, 700–706.
Doktycz, M. J., & Beattie, K. L. (1997). Genosensors and model hybridization studies. In A. Beugelsdiik (Ed.), Automation technologies for genome characterization (pp. 205–225). New York: John Wiley and Sons.
Matsumoto, F., Nishio, K., & Masuda, H. (2004). Flow-through-type DNA array based on ideally ordered anodic porous alumina substrate. Advanced Materials Published Online: 17 December. doi: 10.1002/adma.200400360.
Maldonado, R., Espinosa, M., Loyola, P., Beattie, W. G., & Beattie, K. L. (1999). Mutation detection by stacking hybridization on genosensor arrays. Molecular Biotechnology, 11, 13–25.
Maldonado, R., & Beattie, K. L. (2001). Analysis of nucleic acids by tandem hybridization on oligonucleotide microarrays. In J. B. Rampal (Ed.), DNA arrays. Methods and protocols (pp. 157–171). Totowa, NJ: Humana Press.
Maldonado-Rodríguez, R., Espinosa-Lara, M., Barrera-León, O., Colin-Tovar, C., González-Yebra, B., Salcedo-Vargas, M., Santiago-Hernández, J. C., Méndez-Tenorio, A., & Beattie, K. L. (2003). Detection of mutations in RET proto-oncogene codon 634 through double tandem hybridization. Molecular Biotechnology, 25, 13–29.
Rangel-López, A., Maldonado-Rodríguez, R., Salcedo-Vargas, M., Espinosa-Lara, J. M., Méndez-Tenorio, A., & Beattie, K. L. (2005). Low density DNA microarray for detection of most frequent TP53 missense point mutations. BMC Biotechnology, 5, 8.
Sambrook, J., Fritsh, E. F., & Maniatis, T. (2001). Molecular cloning: A laboratory manual (3rd ed.). Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.
Hicks, J. S., Harker, B. W., Beattie, K. L., & Doktycz, M. J. (2001). Modification of an automated liquid-handling system for reagent-jet, nanoliter-level dispensing. Biotechniques, 30, 878–885.
Weber, J. L., & May, P. E. (1989). Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. American Journal of Human Genetics, 44, 388–396.
Betanzos-Cabrera, G., Harker, B. W., Doktycz, M. J., Weber, J. L., & Beattie, K. L. (2007). A comparison of hybridization efficiency between flat glass and channel glass solid supports. Molecular Biotechnology. doi: 10.1007/s12033-007-9001-z.
Dietrich, W. F., Weber, J. L., Kwok, P.-Y., & Nickerson, D. A. (1998). Isolation and analysis of DNA polymorphisms. In R. Myers et al. (Eds.), Genome analysis: A laboratory manual. New York: Cold Spring Harbor Laboratory Press.
Twyman, R. M., & Primrose, S. B. (2003). Techniques patents for SNP genotyping. Pharmacogenomics, 4, 67–79.
Iwasaki, H., Ezura, Y., Ishida, R., Kajita, M., Kodaira, M., Knight, J., Daniel, S., Shi, M., & Emi, M. (2002). Accuracy of genotyping for single nucleotide polymorphisms by a microarray-based single nucleotide polymorphism typing method involving hybridization of short allele-specific oligonucleotides. DNA Research, 30, 59–62.
Wirtenberger, M., Hemminki, K., Chen, B., & Burwinkel, B. (2005). SNP microarray analysis for genome-wide detection of crossover regions. Human Genetics, 117, 389–397.
Allawi, H. T., & SantaLucia, J. (1998). Nearest-neighbor thermodynamics of internal A–C mismatches in DNA. Sequence dependence and pH effects. Biochemistry, 37, 9435–9444.
Doktycz, M. J., Morris, M. D., Dormady, S. J, Beattie, K. L., & Jacobson, K. B. (1995). Optical melting of 128 octamer DNA duplexes. Effects of base pair location and nearest neighbors on thermal stability. The Journal of Biological Chemistry, 270, 8439–8445.
Pyshnyi, D. V., Pyshnaya, I. A., Levina, A. S., Goldberg, E. L., Zarytova, V. F., Knorre, D. G., & Ivanova, E. M. (2001). Thermodynamic analysis of stacking hybridization of oligonucleotides with DNA template. Journal of Biomolecular Structure Dynamics, 19, 555–570.
Southern, E., Mir, K., & Shchepinov, M. (1999). Molecular interactions on microarrays. Nature Genetics, 21, S5–S9.
Lane, M. J., Paner, T., Kashin, I., Faldasz, B. D., Li, B., Gallo, F. J., & Benight, A. S. (1997). The thermodynamic advantage of DNA oligonucleotide “stacking hybridization” reactions: energetics of a DNA nick. Nucleic Acids Research, 25, 611–616.
Acknowledgments
Financial support for this work was provided by PROMEP/103.5/03/1130, Fondo Sectorial Área de Ciencia Básica SEP-CONACYT 2004-C01-46537, PA144B-2007 and by NIH grant R01 HL62681-01.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Betanzos-Cabrera, G., Harker, B.W., Doktycz, M.J. et al. Channel Glass-based Detection of Human Short Insertion/Deletion Polymorphisms by Tandem Hybridization. Mol Biotechnol 38, 145–153 (2008). https://doi.org/10.1007/s12033-007-9004-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-007-9004-9