Abstract
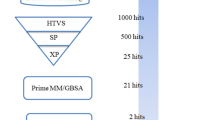
Lung cancer is a disease in which lung cells grow abnormally and uncontrollably, and the cause of it is direct smoking, secondhand smoke, radon, asbestos, and certain chemicals. The worldwide leading cause of death is lung cancer, which is responsible for more than 1.8 million deaths yearly and is expected to rise to 2.2 million by 2030. The most common type of lung cancer is non-small cell lung cancer (NSCLC), which accounts for about 80% and small cell lung cancer (SCLC), which is more aggressive than NSCLC and is often diagnosed later and accounts for 20% of cases. The global concern for lung cancer demands efficient drugs with the slightest chance of developing resistance, and the idea of multitargeted drug designing came up with the solution. In this study, we have performed multitargeted molecular docking studies of Drug Bank compounds with HTVS, SP and XP algorithms followed by MM\GBSA against the four proteins of lung cancer cellular survival and stress responses, which revealed Mitoglitazone as a multitargeted inhibitor with a docking and MM\GBSA score ranging from − 5.784 to − 7.739 kcal/mol and − 25.81 to − 47.65kcal/mol, respectively. Moreover, we performed pharmacokinetics studies and QM-based DFT analysis, showing suitable candidate and interaction pattern analysis revealed the most count of interacting residues was 4GLY, 5PHE, 6ASP, 6GLU, 6LYS, and 6THR. Further, the results were validated with SPC water model-based MD simulation for 100ns in neutralised condition, showing the cumulative deviation and fluctuation < 2Å with many intermolecular interactions. The whole analysis has suggested that Mitoglitazone can be used as a multitargeted inhibitor against lung cancer—however, experimental studies are needed before human use.








Similar content being viewed by others
Data availability
The manuscript provides all the data in figures and tables.
References
Rudin CM, et al. Small-cell lung cancer. Nat Rev Dis Primers. 2021;7(1):3.
Vanka KS, et al. Understanding the pathogenesis of occupational coal and silica dust-associated lung disease. Eur Respir Rev. 2022;31(165):210250.
Oser MG, et al. Transformation from non-small-cell lung cancer to small-cell lung cancer: molecular drivers and cells of origin. Lancet Oncol. 2015;16(4):e165–72.
Wistuba II, Gazdar AF, Minna JD. Molecular genetics of small cell lung carcinoma. Semin Oncol. 2001. https://doi.org/10.1016/S0093-7754(01)90072-7.
Molina JR, et al. Non-small cell lung cancer: epidemiology, risk factors, treatment, and survivorship. Mayo Clin Proc. 2008. https://doi.org/10.4065/83.5.584.
Birring S, Peake M. Symptoms and the early diagnosis of lung cancer. Thorax. 2005;60:268–9.
Kim HC, et al. Five-year overall survival and prognostic factors in patients with lung cancer: results from the Korean association of lung cancer registry (KALC-R) 2015. Cancer Res Treat. 2022;55(1):103–11.
Garinet S, et al. Updated prognostic factors in localized NSCLC. Cancers. 2022;14(6):1400.
Ahmad S, et al. Multitargeted molecular dynamic understanding of butoxypheser against SARS-CoV-2: an in silico study. Nat Product Commun. 2022;17(7):1934578221115499.
Ahmad S, et al. Molecular dynamics simulation and docking analysis of NF-κB protein binding with sulindac acid. Bioinformation. 2022;18(3):170–9.
Ahmad S, et al. Mobile technology solution for COVID-19: surveillance and prevention. In: Computational intelligence methods in COVID-19: surveillance, prevention, prediction and diagnosis. Singapore: Springer; 2021. p. 79–108.
Ahmad S, et al. Therapeutic protein-based vaccines. In: Protein-based therapeutics. Singapore: Springer; 2023. p. 355–84.
Abaza A, et al. Programmed cell death protein 1 (PD-1) and programmed cell death ligand 1 (PD-L1) immunotherapy: a promising breakthrough in cancer therapeutics. Cureus. 2023. https://doi.org/10.7759/cureus.44582.
Ahmad S, et al. Nanoinformatics and nanomodeling: recent developments in computational nanodrug design and delivery systems. In: Emerging nanotechnologies for medical applications. Amsterdam: Elsevier; 2023. p. 297–332.
Ahmad S, et al. Reporting dinaciclib and theodrenaline as a multitargeted inhibitor against SARS-CoV-2: an in-silico study. J Biomol Struct Dyn. 2023;41(9):4013–23.
Ahmad S, Raza K. Identification of 5-nitroindazole as a multitargeted inhibitor for CDK and transferase kinase in lung cancer: a multisampling algorithm-based structural study. Mol Divers. 2023. https://doi.org/10.1007/s11030-023-10648-0.
Ahmad S, et al. In-silico analysis reveals Quinic acid as a multitargeted inhibitor against cervical cancer. J Biomol Struct Dyn. 2022;41:1–17.
Robertson AG, et al. Identification and structure of the nerve growth factor binding site on TrkA. Biochem Biophys Res Commun. 2001;282(1):131–41.
He X-L, Garcia KC. Structure of nerve growth factor complexed with the shared neurotrophin receptor p75. Science. 2004;304(5672):870–5.
Sun C, et al. Solution structure of human survivin and its binding interface with Smac/Diablo. Biochemistry. 2005;44(1):11–7.
Wisniewska M, et al. Crystal structures of the ATPase domains of four human Hsp70 isoforms: HSPA1L/Hsp70-hom, HSPA2/Hsp70-2, HSPA6/Hsp70B’, and HSPA5/BiP/GRP78. PLoS ONE. 2010;5(1): e8625.
Ahmad S, et al. Illustrious implications of nature-inspired computing methods in therapeutics and computer-aided drug design. In: Nature-inspired intelligent computing techniques in bioinformatics. Singapore: Springer; 2022. p. 293–308.
Ahmad S, et al. Multisampling-based docking reveals Imidazolidinyl urea as a multitargeted inhibitor for lung cancer: an optimisation followed multi-simulation and in-vitro study. J Biomol Struct Dyn. 2023;42:1–18.
Alghamdi YS, et al. Unveiling the multitargeted potential of N-(4-Aminobutanoyl)-S-(4-methoxybenzyl)-L-cysteinylglycine (NSL-CG) against SARS CoV-2: a virtual screening and molecular dynamics simulation study. J Biomol Struct Dyn. 2023;41(14):6633–42.
Alturki NA, et al. In-silico screening and molecular dynamics simulation of drug bank experimental compounds against SARS-CoV-2. Molecules. 2022;27(14):4391.
Alzamami A, et al. Hemi-Babim and fenoterol as potential inhibitors of MPro and papain-like protease against SARS-CoV-2: an in-silico study. Medicina. 2022;58(4):515.
Balasubramanian B, et al. Exosomes as an emerging nanoplatform for functional therapeutics. In: Handbook on nanobiomaterials for therapeutics and diagnostic applications. Amsterdam: Elsevier; 2021. p. 483–98.
Famuyiwa SO, et al. Comprehensive computational studies of naturally occurring kuguacins as antidiabetic agents by targeting visfatin. Chem Afr. 2023. https://doi.org/10.1007/s42250-023-00604-8.
Karwasra R, et al. Macrophage-targeted punicalagin nanoengineering to alleviate methotrexate-induced neutropenia: a molecular docking, DFT, and MD simulation analysis. Molecules. 2022;27(18):6034.
Ramlal A, et al. From molecules to patients: the clinical applications of biological databases and electronic health records. In: Translational bioinformatics in healthcare and medicine. Amsterdam: Academic Press; 2021. p. 107–25.
Berman, H., et al., RCSB PDB. 2012. https://academic.oup.com/nar/article/28/1/235/2384399
Yadav MK, et al. Predictive modeling and therapeutic repurposing of natural compounds against the receptor-binding domain of SARS-CoV-2. J Biomol Struct Dyn. 2022. https://doi.org/10.1080/07391102.2021.2021993.
Release S. Protein preparation Wizard, Epik, Schrödinge, LLC, New York, NY, 2022. New York: Schrödinger LLC; 2022.
Madhavi Sastry G, et al. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des. 2013;27:221–34.
Release, S., Maestro S, LLC, New York, NY, 2022. Maestro, schrödinger. Maestro, Schrödinger, 2022. http://schrodinger.com/
Shelley JC, et al. Epik: a software program for pK a prediction and protonation state generation for drug-like molecules. J Comput Aided Mol Des. 2007;21:681–91.
Li H, Robertson AD, Jensen JH. Very fast empirical prediction and rationalization of protein pKa values. Proteins. 2005;61(4):704–21.
Jorgensen WL, Tirado-Rives J. The OPLS [optimized potentials for liquid simulations] potential functions for proteins, energy minimizations for crystals of cyclic peptides and crambin. J Am Chem Soc. 1988;110(6):1657–66.
Singh AP, et al. Computational screening and MM/GBSA-based MD simulation studies reveal the high binding potential of FDA-approved drugs against Cutibacterium acnes sialidase. J Biomol Struct Dyn. 2023. https://doi.org/10.1080/07391102.2023.2242950.
Tarique M, et al. Novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV2) and other coronaviruses: a genome-wide comparative annotation and analysis. Mol Cell Biochem. 2021;476:2203–17.
Tripathi MK, et al. Fundamentals of molecular modeling in drug design. In: Computer aided drug design (CADD): from ligand-based methods to structure-based approaches. Amsterdam: Elsevier; 2022. p. 125–55.
Wishart DS, et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 2018;46(1):D1074–82.
Release, S., LigPrep, Schrödinger, LLC, New York, NY, 2022. New York, NY, 2022. http://schrodinger.com/
Rasul HO, et al. Screening the possible anti-cancer constituents of Hibiscus rosa-sinensis flower to address mammalian target of rapamycin: an in silico molecular docking, HYDE scoring, dynamic studies, and pharmacokinetic prediction. Mol Divers. 2023;27(5):2273–96.
Rana M, et al. Synthesis, single crystal, TD-DFT, molecular dynamics simulation and DNA binding studies of carbothioamide analog. J Mol Struct. 2023;1287: 135701.
Shah AA, et al. Structure-based virtual screening, molecular docking, molecular dynamics simulation, and metabolic reactivity studies of quinazoline derivatives for their anti-EGFR activity against tumor angiogenesis. Curr Med Chem. 2023. https://doi.org/10.2174/0929867330666230309143711.
Sheikh K, et al. Consequential innovations in nature-inspired intelligent computing techniques for biomarkers and potential therapeutics identification. In: Nature-inspired intelligent computing techniques in bioinformatics. Singapore: Springer; 2022. p. 247–74.
Glide, S., LLC. New York, NY, 2022. 2022. http://schrodinger.com/
Friesner RA, et al. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein—ligand complexes. J Med Chem. 2006;49(21):6177–96.
QikProp, S., LLC. New York, NY, 2022. http://schrodinger.com/
Lipinski CA. Lead-and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol. 2004;1(4):337–41.
Bochevarov AD, et al. Jaguar: a high-performance quantum chemistry software program with strengths in life and materials sciences. Int J Quantum Chem. 2013;113(18):2110–42.
Bowers, K.J., et al. Scalable algorithms for molecular dynamics simulations on commodity clusters. In: Proceedings of the 2006 ACM/IEEE Conference on Supercomputing. 2006.
Jorgensen WL. Convergence of Monte Carlo simulations of liquid water in the NPT ensemble. Chem Phys Lett. 1982;92(4):405–10.
Rasul HO, et al. Discovery of potential mTOR inhibitors from Cichorium intybus to find new candidate drugs targeting the pathological protein related to the breast cancer: an integrated computational approach. Mol Divers. 2023;27(3):1141–62.
Rasul HO, et al. In silico molecular docking and dynamic simulation of eugenol compounds against breast cancer. J Mol Model. 2022;28(1):17.
Acknowledgements
The authors extend their appreciation to the Deputyship for Research & Innovation, Ministry of Education in Saudi Arabia, for funding this research work through the project number (IF2/PSAU/2022/03/22763).
Funding
The authors are thankful to the Deanship of Scientific Research at Prince Sattam bin Abdulaziz University for funding this work under the General Research Funding program Grant Code Number (IF2/PSAU/2022/03/22763).
Author information
Authors and Affiliations
Contributions
ASB: Conceptualisation, Supervision, Data curation, writing—first draft, FMA: Data collection, analysis, reviewing and editing, OSA: Supervision, reviewing and editing, AH: extensive editing of the first draft, HHA: Supervision, visualisation and editing of the first draft.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare no competing or conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Binshaya, A.S., Alkahtani, O.S., Aldakheel, F.M. et al. Structure-based multitargeted docking screening, pharmacokinetics, DFT, and dynamics simulation studies reveal mitoglitazone as a potent inhibitor of cellular survival and stress response proteins of lung cancer. Med Oncol 41, 101 (2024). https://doi.org/10.1007/s12032-024-02342-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-024-02342-4