Abstract
Targeting the extracellular matrix (ECM) is considered as a promising strategy in cancer therapeutics. This study was designed to identify the potential ECM modulators for gastric cancer therapeutics. Exploration of the expression profiles of gastric tumors revealed the elevated expression of ECM genes in gastric tumor tissues compared to the adjacent normal tissues with increased expression in diffuse subtype gastric tumors and specifically in epithelial to mesenchymal transition (EMT) molecular subtype tumors. Consensus ECM gene set was derived from the expression profiles of gastric tumors. The correlative analysis was performed between the expression pattern of the ECM gene set and the drug sensitivity pattern of a panel of drugs across gastric cancer cell lines. Negative correlation between the expression of ECM genes and sensitivity of a number of drugs targeting PI3K/mTOR signaling, chromatin histone acetylation and ABL signaling was observed. These pathways are known for their role in cell-mediated adhesion, differentiation and epithelial to mesenchymal transition. The current results reveal the possibility of using PI3K/AKT/mTOR modulators for targeted gastric cancer therapy in patients with dysregulated ECM.






Similar content being viewed by others
Data availability
Not applicable.
Abbreviations
- ACRG:
-
Asian Cancer Research Group
- ATCC:
-
American Type Culture Collection
- CCLE:
-
The cancer cell line encyclopedia
- ECM:
-
Extracellular matrix
- EMT:
-
Epithelial to mesenchymal transition
- GDSC:
-
Genomics of drug sensitivity in cancer
- GEO:
-
Gene expression omnibus
- MMP:
-
Matrix metalloproteinase
- MSigDB:
-
Molecular signatures database
- TCGA:
-
The cancer genome atlas
References
Andreuzzi E, Capuano A, Poletto E, et al. Role of extracellular matrix in gastrointestinal cancer-associated angiogenesis. Int J Mol Sci. 2020;21:3686.
Henke E, Nandigama R, Ergün S. Extracellular matrix in the tumor microenvironment and its impact on cancer therapy. Front Mol Biosci. 2020;6:160.
Huang J, Zhang L, Wan D, et al. Extracellular matrix and its therapeutic potential for cancer treatment. Signal Transduct Target Ther. 2021;6:153.
Elgundi Z, Papanicolaou M, Major G, et al. Cancer metastasis: the role of the extracellular matrix and the heparan sulfate proteoglycan perlecan. Front Oncol. 2020;9:1482.
Bass AJ, Thorsson V, Shmulevich I, et al. Comprehensive molecular characterization of gastric adenocarcinoma. Nature. 2014;513:202–9. https://doi.org/10.1038/nature13480.
Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50. https://doi.org/10.1073/pnas.0506580102.
Shao X, Taha IN, Clauser KR, et al. MatrisomeDB: the ECM-protein knowledge database. Nucl Acids Res. 2020. https://doi.org/10.1093/nar/gkz849.
Levine DM, Haynor DR, Castle JC, et al. Pathway and gene-set activation measurement from mRNA expression data: the tissue distribution of human pathways. Genome Biol. 2006. https://doi.org/10.1186/gb-2006-7-10-r93.
Ooi CH, Ivanova T, Wu J, et al. Oncogenic pathway combinations predict clinical prognosis in gastric cancer. PLoS Genet. 2009. https://doi.org/10.1371/journal.pgen.1000676.
Ashburner M, Ball CA, Blake JA, et al. Gene ontology: tool for the unification of biology. Nat Genet. 2000;25:25–9.
Chen EY, Tan CM, Kou Y, et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 2013. https://doi.org/10.1186/1471-2105-14-128.
Szász AM, Lánczky A, Nagy Á et al (2016) Cross-validation of survival associated biomarkers in gastric cancer using transcriptomic data of 1065 patients. Oncotarget 7:49322–49333. https://doi.org/10.18632/oncotarget.10337
Gotea V, Ovcharenko I. DiRE: identifying distant regulatory elements of co-expressed genes. Nucl Acids Res. 2008. https://doi.org/10.1093/nar/gkn300.
Donaldson JG. Immunofluorescence staining. Curr Protoc Cell Biol. 2015;69:431–7. https://doi.org/10.1002/0471143030.cb0403s69.
Tamilzhalagan S, Rathinam D, Ganesan K. Amplified 7q21-22 gene MCM7 and its intronic miR-25 suppress COL1A2 associated genes to sustain intestinal gastric cancer features. Mol Carcinog. 2017;56:1590–602. https://doi.org/10.1002/mc.22614.
Pickup MW, Mouw JK, Weaver VM (2014) The extracellular matrix modulates the hallmarks of cancer. EMBO Rep. https://doi.org/10.15252/embr.201439246
Moreira AM, Pereira J, Melo S, et al. The extracellular matrix: an accomplice in gastric cancer development and progression. Cells. 2020;9:394.
Jinawath N, Furukawa Y, Hasegawa S, et al. Comparison of gene-expression profiles between diffuse- and intestinal-type gastric cancers using a genome-wide cDNA microarray. Oncogene. 2004. https://doi.org/10.1038/sj.onc.1207886.
Yap YS, McPherson JR, Ong CK et al (2014) The NF1 gene revisited—from bench to bedside. Oncotarget. https://doi.org/10.18632/oncotarget.2194
Yang W, Soares J, Greninger P, et al. Genomics of drug sensitivity in cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucl Acids Res. 2013. https://doi.org/10.1093/nar/gks1111.
Barretina J, Caponigro G, Stransky N, et al. The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7. https://doi.org/10.1038/nature11003.
Malik R, Lelkes PI, Cukierman E. Biomechanical and biochemical remodeling of stromal extracellular matrix in cancer. Trends Biotechnol. 2015;33:230–6.
Walker C, Mojares E, Del Río HA. Role of extracellular matrix in development and cancer progression. Int J Mol Sci. 2018;19:3028.
Winkler J, Abisoye-Ogunniyan A, Metcalf KJ, Werb Z. Concepts of extracellular matrix remodelling in tumour progression and metastasis. Nat Commun. 2020;11:5120.
Aktar R, Peiris M, Fikree A, et al. A novel role for the extracellular matrix glycoprotein-Tenascin-X in gastric function. J Physiol. 2019. https://doi.org/10.1113/JP277195.
Li Z, Liu Z, Shao Z, et al. Identifying multiple collagen gene family members as potential gastric cancer biomarkers using integrated bioinformatics analysis. PeerJ. 2020. https://doi.org/10.7717/peerj.9123.
Järveläinen H, Sainio A, Koulu M, et al. Extracellular matrix molecules: potential targets in pharmacotherapy. Pharmacol Rev. 2009;61:198–223.
Holle AW, Young JL, Spatz JP. In vitro cancer cell-ECM interactions inform in vivo cancer treatment. Adv Drug Deliv Rev. 2016;97:270–9.
Harisi R, Jeney A. Extracellular matrix as target for antitumor therapy. Onco Targets Ther. 2015. https://doi.org/10.2147/OTT.S48883.
Subrahmanyam N, Ghandehari H. Harnessing extracellular matrix biology for tumor drug delivery. J Pers Med. 2021;11:88.
Nallanthighal S, Heiserman JP, Cheon DJ. The role of the extracellular matrix in cancer stemness. Front Cell Dev Biol. 2019;7:86.
Morris JC, Tan AR, Olencki TE, et al. Phase I study of GC1008 (Fresolimumab): a human anti-transforming growth factor-beta (TGFβ) monoclonal antibody in patients with advanced malignant melanoma or renal cell carcinoma. PLoS ONE. 2014. https://doi.org/10.1371/journal.pone.0090353.
Élez E, Kocáková I, Höhler T, et al. Abituzumab combined with cetuximab plus irinotecan versus cetuximab plus irinotecan alone for patients with KRAS wild-type metastatic colorectal cancer: the randomised phase I/II POSEIDON trial. Ann Oncol. 2015. https://doi.org/10.1093/annonc/mdu474.
Maden CH, Fairman D, Chalker M, et al. Safety, tolerability and pharmacokinetics of GSK3008348, a novel integrin αvβ6 inhibitor, in healthy participants. Eur J Clin Pharmacol. 2018. https://doi.org/10.1007/s00228-018-2435-3.
Jang M, Koh I, Lee SJ, et al. Droplet-based microtumor model to assess cell-ECM interactions and drug resistance of gastric cancer cells. Sci Rep. 2017. https://doi.org/10.1038/srep41541.
Funding
This work was supported by the Department of Biotechnology (DBT), Government of India, through the Unit of Excellence (UOE) in Cancer Genetics Grant BT/MED/30/SP11290/2015 and MKU-RUSA supported grant 014/ RUSA/MKU/2020-2021 to Dr. Kumaresan Ganesan, Madurai Kamaraj University. Senior Research Fellowship to Ponmathi Panneerpandian from Lady Tata Memorial Trust, Mumbai, India, is acknowledged. MKU-RUSA, DST-FIST, UGC-NRCBS, UGC-CAS, and DST-PURSE programe supported central facilities of SBS, MKU are acknowledged.
Author information
Authors and Affiliations
Contributions
GK and PP conceptualized the study. PP performed the experiments. GK and PP wrote the paper. PP and GK analyzed and interpreted the data.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Ethics approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary Fig. S1
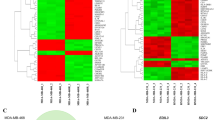
: Extracellular matrix gene sets are highly expressed in gastric tumors. A Heatmap showing differential expression of ECM gene sets across the gastric tumor profiles GSE29272 (A) and GSE54129 (B) containing normal and gastric tumor samples. The ECM genes are highly expressed in gastric tumors compared to the normal tissues. B Pathway activation scoring for the 21 ECM gene sets across the gastric tumor profiles GSE3809 (C), GSE62254 (D) and TCGA (E) reveal the enrichment of ECM gene sets in diffuse subtype gastric tumors compared to intestinal subtype gastric tumors. The derived 141 ECM genes obtained through overlap among 21 ECM gene sets show the higher expression of ECM genes in diffuse subtype gastric tumors in the gastric tumor profiles GSE62254 (F), GSE35809 (G), GSE22377 (H), and TCGA (I). For the clear visibility of the ECM gene sets comprising a list of 21 signatures is provided in Supplementary Table S1. Fig. S2: The table represents the previously annotated drug probabilities for diffuse, epithelial to mesenchymal transition, and genomically stable subtypes of gastric cancers for TCGA and ACRG cohort based studies. Fig. S3: Analysis of the potential ECM targetable drugs. A Expression of ECM genes across a panel of gastric cancer cell lines. B, C The sensitivity pattern of the corresponding gastric cancer cell lines to dactolisib,the PI3K inhibitor (B) and dasatinib—an ABL inhibitor (C), show striking negative correlation. D The genes downregulated upon treatment with dasatinib also overlap with the signatures associated with EGFR and vasculature from MsigDB indicating its potential role in inhibiting angiogenesis. E Gene ontology based functional analysis of the genes downregulated by dasatinib show the enrichment of TGF-β gene set. (PPTX 1801 kb)
Supplementary Table S1
: List of ECM related signatures collected from MSigDB and used for pathway activation scoring in gastric tumors. Table S2: ECM gene list (n = 141) derived by overlap among the collected 21 ECM related gene sets. Table S3: The drugs negatively correlated for their expression of ECM with sensitivity to drugs belonging to the annotated target pathways along with their respective correlation coefficient values. (XLSX 58 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Panneerpandian, P., Ganesan, K. PI3K/AKT/mTOR inhibitors as potential extracellular matrix modulators for targeting EMT subtype gastric tumors. Med Oncol 40, 120 (2023). https://doi.org/10.1007/s12032-023-01984-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-023-01984-0