Abstract
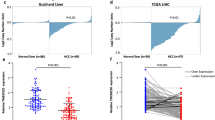
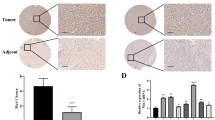
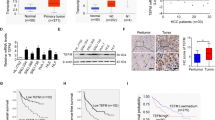
Tubulin γ-1 (TUBG1) is a highly conserved component of the centrosome and its deregulation is involved in the development of several types of cancer. However, the role of TUBG1 in hepatocellular carcinoma (HCC) remains unclear. In this study, we found that TUBG1 was upregulated in human HCC cells and tissues and that TUBG1 upregulation was associated with promoter hypomethylation in HCC tissues. TUBG1 knockdown suppressed the proliferation, invasion, and migration of HCC cells. While TUBG1 expression was positively correlated with CD4 + memory T lymphocyte infiltration, it was negatively correlated with CD4 + regulatory T-cell infiltration in human HCC tissues. Furthermore, TUBG1 expression was positively correlated with the expression of genes involved in cell division. Noticeably, high expression of TUBG1 was associated with poor prognosis in patients with HCC. Overall, our findings revealed that TUBG1 promotes hepatocarcinogenesis by increasing proliferation, invasion, and migration of HCC cells and may regulate T lymphocyte infiltration. The current findings provide important insights into TUBG1 regulation in HCC, which could provide new therapeutic targets for hepatocarcinoma which has a very high incidence and mortality rate worldwide.






Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Abbreviations
- BP:
-
Biological process
- CC:
-
Cellular component
- DFS:
-
Disease-free survival
- DMEM:
-
Dulbecco’s modified eagle medium
- DSS:
-
Disease-specific survival
- FBS:
-
Fetal bovine serum
- GO:
-
Gene Ontology
- GTEx:
-
Genotype-tissue expression
- HCC:
-
Hepatocellular carcinoma
- ICGC:
-
International cancer genome consortium
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- K-M:
-
Kaplan–Meier
- LIHC:
-
Liver hepatocellular carcinoma
- LIRI-JP:
-
Liver Cancer-RIKEN, Japan
- MF:
-
Molecular function
- OS:
-
Overall survival
- PFS:
-
Progression-free survival
- siRNA:
-
Small interfering RNA
- TCGA:
-
The cancer genome atlas
- TILs:
-
Tumor-infiltrating lymphocytes
- TUBG1 :
-
Tubulin gamma-1
- TUBG2 :
-
Tubulin γ-2
- UALCAN:
-
The University of Alabama at Birmingham Cancer data analysis Portal
References
Arnold M, Abnet CC, Neale RE, Vignat J, Giovannucci EL, McGlynn KA, et al. Global burden of 5 major types of gastrointestinal cancer. Gastroenterology. 2020;159(335–49):e15. https://doi.org/10.1053/j.gastro.2020.02.068.
Huang DQ, El-Serag HB, Loomba R. Global epidemiology of NAFLD-related HCC: trends, predictions, risk factors and prevention. Nat Rev Gastroenterol Hepatol. 2021;18:223–38. https://doi.org/10.1038/s41575-020-00381-6.
Ahn JC, Teng PC, Chen PJ, Posadas E, Tseng HR, Lu SC, et al. Detection of circulating tumor cells and their implications as a biomarker for diagnosis, prognostication, and therapeutic monitoring in hepatocellular carcinoma. Hepatology. 2021;73:422–36. https://doi.org/10.1002/hep.31165.
Dutcher SK. The tubulin fraternity alpha to eta. Curr Opin Cell Biol. 2001;13:49–54. https://doi.org/10.1016/s0955-0674(00)00173-3.
Stathatos GG, Dunleavy JEM, Zenker J, O’Bryan MK. Delta and epsilon tubulin in mammalian development. Trends Cell Biol. 2021;31:774–87. https://doi.org/10.1016/j.tcb.2021.03.010.
Akhmanova A, Steinmetz MO. Control of microtubule organization and dynamics: two ends in the limelight. Nat Rev Mol Cell Biol. 2015;16:711–26. https://doi.org/10.1038/nrm4084.
Mukherjee A. Conduit PT. γ-TuRCs. Curr Biol. 2019;29:R398-400. https://doi.org/10.1016/j.cub.2019.04.013.
Hećimović H, Bosnjak J, Demarin V. Prevalence of mood dysfunction in epilepsy patients in Croatia. Coll Antropol. 2008;32(Suppl 1):65–8.
Stearns T, Evans L, Kirschner M. Gamma-tubulin is a highly conserved component of the centrosome. Cell. 1991;65:825–36. https://doi.org/10.1016/0092-8674(91)90390-k.
Wise DO, Krahe R, Oakley BR. The gamma-tubulin gene family in humans. Genomics. 2000;67:164–70. https://doi.org/10.1006/geno.2000.6247.
Yuba-Kubo A, Kubo A, Hata M, Tsukita S. Gene knockout analysis of two gamma-tubulin isoforms in mice. Dev Biol. 2005;282:361–73. https://doi.org/10.1016/j.ydbio.2005.03.031.
Draberova E, Sulimenko V, Vinopal S, Sulimenko T, Sladkova V, D’Agostino L, et al. Differential expression of human gamma-tubulin isotypes during neuronal development and oxidative stress points to a gamma-tubulin-2 prosurvival function. FASEB J. 2017;31:1828–46. https://doi.org/10.1096/fj.201600846RR.
Nigg EA, Raff JW. Centrioles, centrosomes, and cilia in health and disease. Cell. 2009;139:663–78. https://doi.org/10.1016/j.cell.2009.10.036.
Shao W, Yang J, He M, Yu XY, Lee CH, Yang Z, et al. Centrosome anchoring regulates progenitor properties and cortical formation. Nature. 2020;580:106–12. https://doi.org/10.1038/s41586-020-2139-6.
Camargo Ortega G, Falk S, Johansson PA, Peyre E, Broix L, Sahu SK, et al. The centrosome protein AKNA regulates neurogenesis via microtubule organization. Nature. 2019;567:113–7. https://doi.org/10.1038/s41586-019-0962-4.
Blanco I, Kuchenbaecker K, Cuadras D, Wang X, Barrowdale D, de Garibay GR, et al. Assessing associations between the AURKA-HMMR-TPX2-TUBG1 functional module and breast cancer risk in BRCA1/2 mutation carriers. PLoS One. 2015;10:e0120020. https://doi.org/10.1371/journal.pone.0120020.
Olson JE, Wang X, Pankratz VS, Fredericksen ZS, Vachon CM, Vierkant RA, et al. Centrosome-related genes, genetic variation, and risk of breast cancer. Breast Cancer Res Treat. 2011;125:221–8. https://doi.org/10.1007/s10549-010-0950-8.
Alvarado-Kristensson M. γ-tubulin as a signal-transducing molecule and meshwork with therapeutic potential. Signal Transduction Targeted Ther. 2018;3:24. https://doi.org/10.1038/s41392-018-0021-x.
Caracciolo V, D’Agostino L, Draberova E, Sladkova V, Crozier-Fitzgerald C, Agamanolis DP, et al. Differential expression and cellular distribution of gamma-tubulin and betaIII-tubulin in medulloblastomas and human medulloblastoma cell lines. J Cell Physiol. 2010;223:519–29. https://doi.org/10.1002/jcp.22077.
Chengcheng L, Raza SHA, Shengchen Y, Mohammedsaleh ZM, Shater AF, Saleh FM, et al. Bioinformatics role of the WGCNA analysis and co-expression network identifies of prognostic marker in lung cancer. Saudi J Biol Sci. 2022;29:3519–27. https://doi.org/10.1016/j.sjbs.2022.02.016.
Dementyeva E, Kryukov F, Kubiczkova L, Nemec P, Sevcikova S, Ihnatova I, et al. Clinical implication of centrosome amplification and expression of centrosomal functional genes in multiple myeloma. J Transl Med. 2013;11:77. https://doi.org/10.1186/1479-5876-11-77.
Maounis NF, Dráberová E, Mahera E, Chorti M, Caracciolo V, Sulimenko T, et al. Overexpression of γ-tubulin in non-small cell lung cancer. Histol Histopathol. 2012;27:1183–94. https://doi.org/10.14670/hh-27.1183.
Tang Z, Kang B, Li C, Chen T, Zhang Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019;47:W556–60. https://doi.org/10.1093/nar/gkz430.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1. https://doi.org/10.1126/scisignal.2004088.
Chandrashekar DS, Karthikeyan SK, Korla PK, Patel H, Shovon AR, Athar M, et al. UALCAN: an update to the integrated cancer data analysis platform. Neoplasia. 2022;25:18–27. https://doi.org/10.1016/j.neo.2022.01.001.
Newman AM, Steen CB, Liu CL, Gentles AJ, Chaudhuri AA, Scherer F, et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat Biotechnol. 2019;37:773–82. https://doi.org/10.1038/s41587-019-0114-2.
Henrichsen CN, Chaignat E, Reymond A. Copy number variants, diseases and gene expression. Hum Mol Genet. 2009;18:R1-8. https://doi.org/10.1093/hmg/ddp011.
Zarrei M, MacDonald JR, Merico D, Scherer SW. A copy number variation map of the human genome. Nat Rev Genet. 2015;16:172–83. https://doi.org/10.1038/nrg3871.
Gromley A, Jurczyk A, Sillibourne J, Halilovic E, Mogensen M, Groisman I, et al. A novel human protein of the maternal centriole is required for the final stages of cytokinesis and entry into S phase. J Cell Biol. 2003;161:535–45. https://doi.org/10.1083/jcb.200301105.
Müller H, Fogeron ML, Lehmann V, Lehrach H, Lange BM. A centrosome-independent role for gamma-TuRC proteins in the spindle assembly checkpoint. Science. 2006;314:654–7. https://doi.org/10.1126/science.1132834.
Nayak T, Edgerton-Morgan H, Horio T, Xiong Y, De Souza CP, Osmani SA, et al. Gamma-tubulin regulates the anaphase-promoting complex/cyclosome during interphase. J Cell Biol. 2010;190:317–30. https://doi.org/10.1083/jcb.201002105.
Ferguson RL, Maller JL. Centrosomal localization of cyclin E-Cdk2 is required for initiation of DNA synthesis. Curr Biol. 2010;20:856–60. https://doi.org/10.1016/j.cub.2010.03.028.
Hopkins JL, Lan L, Zou L. DNA repair defects in cancer and therapeutic opportunities. Genes Dev. 2022;36:278–93. https://doi.org/10.1101/gad.349431.122.
Zhao Y, Simon M, Seluanov A, Gorbunova V. DNA damage and repair in age-related inflammation. Nat Rev Immunol. 2022. https://doi.org/10.1038/s41577-022-00751-y.
Huang R, Zhou PK. DNA damage repair: historical perspectives, mechanistic pathways and clinical translation for targeted cancer therapy. Signal Transduct Target Ther. 2021;6:254. https://doi.org/10.1038/s41392-021-00648-7.
Hojyo S, Tumes D, Murata A, Tokoyoda K. Multiple developmental pathways lead to the generation of CD4 T-cell memory. Int Immunol. 2020;32:589–95. https://doi.org/10.1093/intimm/dxaa051.
Sakaguchi S, Mikami N, Wing JB, Tanaka A, Ichiyama K, Ohkura N. Regulatory T Cells and human disease. Annu Rev Immunol. 2020;38:541–66. https://doi.org/10.1146/annurev-immunol-042718-041717.
Acknowledgements
The authors wish to thank Editage (www.editage.cn) for English language editing.
Funding
This work was supported by the National Natural Science Foundation of China (Grant Numbers 81972648, 82172915, and 81773011) and Chongqing Medical University Program for Youth Innovation in Future Medicine (Grant Number W0084).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. K-FT conceived and designed the study. Z-jW, Z-zD, M-zH, J-nL, HL, M-mS, S-jZ, H-jS, and JG performed the experiments and analyzed the data. Z-jW and Z-zD drafted the manuscript. K-FT reviewed the manuscript. K-FT and A-LH supervised the study. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
We declare no relevant information regarding the ethical conduct in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Zj., Dai, Zz., Hu, Mz. et al. Upregulation of TUBG1 expression promotes hepatocellular carcinoma development. Med Oncol 40, 96 (2023). https://doi.org/10.1007/s12032-023-01966-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-023-01966-2