Abstract
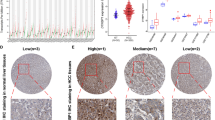
Hepatocellular carcinoma (HCC) is the fifth most common neoplasm in the world. Chronic inflammation of liver and associated wound healing processes collectively contribute to the development of cirrhosis which further progresses to dysplastic nodule and then to HCC. Etiological mediators and ongoing manipulations at cellular level in HCC are well established; however, key protein interactions and genetic alterations involved in stepwise hepatocarcinogenic pathways are seldom explored. This study aims to unravel novel targets of HCC and repurpose the FDA-approved drugs against the same. Genetic data pertinent to different stages of HCC were retrieved from GSE6764 dataset and analyzed via GEO2R. Subsequently, protein–protein interaction network analysis of differentially expressed genes was performed to identify the hub genes with significant interaction. Hub genes displaying higher interactions were considered as potential HCC targets and were validated thorough UALCAN and GEPIA databases. These targets were screened against FDA-approved drugs through molecular docking and dynamics simulation studies to capture the drugs with potential activity against HCC. Finally, cytotoxicity of the shortlisted drug was confirmed in vitro by MTT assay. CDC20 was identified as potential druggable target. Docking, binding energy calculations, and dynamic studies revealed significant interaction exhibited by Labetalol with CDC20. Further, in MTT assay, Labetalol demonstrated an IC50 of 200.29 µg/ml in inhibiting the cell growth of HepG2 cell line. In conclusion, this study discloses a series of key genetic underpinnings of HCC and recommends the pertinence of labetalol as a potential repurposable drug against HCC.










Similar content being viewed by others
Data availability
Not applicable.
Code availability
Not applicable.
Abbreviations
- rGyr:
-
Radius of Gyration
- ASPM:
-
Assembly Factor for Spindle Microtubules
- BCLC:
-
Barcelona Clinic Liver Cancer
- BUB1:
-
BUB1 Mitotic Checkpoint Serine/Threonine Kinase
- BUB1B:
-
BUB1 Mitotic Checkpoint Serine/Threonine Kinase B
- CCL19:
-
Chemokine (C–C Motif) Ligand 19
- CCNB1:
-
Cyclin B1
- CCNB1:
-
Cyclin B1
- CCNB2:
-
Cyclin B2
- CDC20:
-
Cell Division Cycle 20
- CDK1:
-
Cyclin-Dependent Kinase 1
- CDKN3:
-
Cyclin-Dependent Kinase Inhibitor 3
- CENPF:
-
Centromere Protein F
- CFTR:
-
Cystic Fibrosis Transmembrane Conductance Regulator
- CXCL11:
-
C-X-C Motif Chemokine 11
- DEG:
-
Differentially Expressed Genes
- DN:
-
Dysplastic Nodule
- FDA:
-
Food and Drug Administration
- GEPIA:
-
Gene Expression Profiling Interactive Analysis
- GO:
-
Gene Ontology
- HBV:
-
Hepatitis-B Virus
- HCC:
-
Hepatocellular Carcinoma
- HCV:
-
Hepatitis-C Virus
- HMMR:
-
Hyaluronan-Mediated Motility Receptor
- IGF1:
-
Insulin-Like Growth Factor 1
- IL7R:
-
Interleukin 7 Receptor
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- KIF20A:
-
Kinesin Family Member 20A
- KIF2C:
-
Kinesin Family Member 2C
- KIF4A:
-
Kinesin Family Member 4A
- KRT19:
-
Keratin 19
- MAD2L1:
-
Mitotic Arrest Deficient 2 Like 1
- MELK:
-
Maternal Embryonic Leucine Zipper Kinase
- MM/GBSA:
-
Molecular Mechanics energies combined with the Generalized Born and Surface Area continuum Solvation
- MX1:
-
MX Dynamin Like GTPase 1
- NCAPG:
-
Non-SMC Condensin I Complex Subunit G
- NEK2:
-
NIMA-Related Kinase 2
- PBK:
-
PDZ Binding Kinase
- PDB:
-
Protein Data Bank
- PROM1:
-
Prominin 1
- RMSD:
-
Root Mean Square Deviation
- RMSF:
-
Root-Mean-Square Fluctuation
- STAT1:
-
Signal Transducer and Activator of Transcription 1
- TCGA:
-
The Cancer Genome Atlas
- TOP2A:
-
DNA Topoisomerase II Alpha
- TPX2:
-
TPX2 Microtubule Nucleation Factor
References
McGlynn KA, Petrick JL, El-Serag HB. Epidemiology of hepatocellular carcinoma. Hepatology (Baltimore, MD). 2021;73(Suppl 1):4–13.
Acharya SK. Epidemiology of hepatocellular carcinoma in India. J Clin Exp Hepatol. 2014;4:S27-33.
von Felden J. New systemic agents for hepatocellular carcinoma: an update 2020. Curr Opin Gastroenterol. 2020;36:177–83.
Kondo M, Numata K, Hara K, Nozaki A, Fukuda H, Chuma M, et al. Treatment of advanced hepatocellular carcinoma after failure of sorafenib treatment: subsequent or additional treatment interventions contribute to prolonged survival postprogression. Gastroenterol Res Pract. 2017;2017:5728946.
He B, Yin J, Gong S, Gu J, Xiao J, Shi W, et al. Bioinformatics analysis of key genes and pathways for hepatocellular carcinoma transformed from cirrhosis. Medicine. 2017;96:e6938.
Gns HS, Gr S, Murahari M, Krishnamurthy M. An update on drug repurposing: re-written saga of the drug’s fate. Biomed Pharmacother Biomed Pharmacother. 2019;110:700–16.
Wurmbach E, Chen YB, Khitrov G, Zhang W, Roayaie S, Schwartz M, et al. Genome-wide molecular profiles of HCV-induced dysplasia and hepatocellular carcinoma. Hepatology (Baltimore, MD). 2007;45:938–47.
Meissl K, Macho-Maschler S, Müller M, Strobl B. The good and the bad faces of STAT1 in solid tumours. Cytokine. 2017;89:12–20.
Gao B, Wang H, Lafdil F, Feng D. STAT proteins—key regulators of anti-viral responses, inflammation, and tumorigenesis in the liver. J Hepatol. 2012;57:430–41.
Sung PS, Shin E-C, Yoon SK. Interferon response in Hepatitis C Virus (HCV) infection: lessons from cell culture systems of HCV infection. Int J Mol Sci. 2015;16:23683–94.
Nazari A, Ahmadi Z, Hassanshahi G, Abbasifard M, Taghipour Z, Falahati-Pour SK, et al. Effective treatments for Bladder cancer affecting CXCL9/CXCL10/CXCL11/CXCR3 Axis: a review. Oman Med Spec Board. 2020;35:e103.
Chalin A, Lefevre B, Devisme C, Barget N, Amiot L, Samson M. Circulating levels of CXCL11 and CXCL12 are biomarkers of cirrhosis in patients with chronic hepatitis C infection. Cytokine. 2019;117:72–8.
Hillinger S, Yang SC, Batra RK, Strieter RM, Weder W, Dubinett SM, et al. CCL19 reduces tumour burden in a model of advanced lung cancer. Br J Cancer. 2006;94:1029–34.
Al-Mossawi H, Yager N, Taylor CA, Lau E, Danielli S, de Wit J, et al. Context-specific regulation of surface and soluble IL7R expression by an autoimmune risk allele. Nat Commun. 2019;10:4575.
Willis CR, Seamons A, Maxwell J, Treuting PM, Nelson L, Chen G, et al. Interleukin-7 receptor blockade suppresses adaptive and innate inflammatory responses in experimental colitis. J Inflamm. 2012;9:39.
Liu C, Song C, Li J, Sun Q. CFTR functions as a tumor suppressor and is regulated by DNA methylation in colorectal cancer. Cancer Manag Res. 2020;12:4261–70.
Saha SK, Islam SMR, Kwak KS, Rahman MS, Cho SG. PROM1 and PROM2 expression differentially modulates clinical prognosis of cancer: a multiomics analysis. Cancer Gene Ther. 2020;27:147–67.
Kawai T, Yasuchika K, Ishii T, Katayama H, Yoshitoshi EY, Ogiso S, et al. Keratin 19, a cancer stem cell marker in human hepatocellular carcinoma. Clin Cancer Res Off J Am Assoc Cancer Res. 2015;21:3081–91.
Lu D, Bai X, Zou Q, Gan Z, Lv Y. Identification of the association between HMMR expression and progression of hepatocellular carcinoma via construction of a co-expression network. Oncol Lett. 2020;20:2645–54.
Zhang H, Zhang X, Li X, Meng WB, Bai ZT, Rui SZ, et al. Effect of CCNB1 silencing on cell cycle, senescence, and apoptosis through the p53 signaling pathway in pancreatic cancer. J Cell Physiol. 2018;234:619–31.
Weroha SJ, Haluska P. The insulin-like growth factor system in cancer. Endocrinol Metab Clin North Am. 2012;41:335.
Kubiak JZ, El Dika M. Canonical and alternative pathways in cyclin-dependent kinase 1/cyclin B inactivation upon M-phase exit in Xenopus laevis cell-free extracts. Enzym Res. 2011;2011:523420.
Peyressatre M, Prével C, Pellerano M, Morris MC. Targeting cyclin-dependent kinases in human cancers: from small molecules to peptide inhibitors. Cancers. 2015;7:179–237.
Kidokoro T, Tanikawa C, Furukawa Y, Katagiri T, Nakamura Y, Matsuda K. CDC20, a potential cancer therapeutic target, is negatively regulated by p53. Oncogene. 2008;27:1562–71.
Gao Y, Zhang B, Wang Y, Shang G. Cdc20 inhibitor apcin inhibits the growth and invasion of osteosarcoma cells. Oncol Rep. 2018;40:841–8.
Percy MJ, Myrie KA, Neeley CK, Azim JN, Ethier SP, Petty EM. Expression and mutational analyses of the human MAD2L1 gene in breast cancer cells. Genes Chromosom Cancer. 2000;29:356–62.
Kokuryo T, Yokoyama Y, Yamaguchi J, Tsunoda N, Ebata T, Nagino M. NEK2 is an effective target for cancer therapy with potential to induce regression of multiple human malignancies. Anticancer Res. 2019;39:2251–8.
Ma W, Wang B, Zhang Y, Wang Z, Niu D, Chen S, et al. Prognostic significance of TOP2A in non-small cell lung cancer revealed by bioinformatic analysis. Cancer Cell Int. 2019;19:239.
Li T-F, Zeng H-J, Shan Z, Ye R-Y, Cheang T-Y, Zhang Y-J, et al. Overexpression of kinesin superfamily members as prognostic biomarkers of breast cancer. Cancer Cell Int. 2020;20:123.
Huang H, Lee M-H, Liu K, Dong Z, Ryoo Z, Kim MO. PBK/TOPK: an effective drug target with diverse therapeutic potential. Cancers. 2021;13:2232.
Liu W, Liang B, Liu H, Huang Y, Yin X, Zhou F, et al. Overexpression of non-SMC condensin I complex subunit G serves as a promising prognostic marker and therapeutic target for hepatocellular carcinoma. Int J Mol Med. 2017;40:731–8.
Janostiak R, Rauniyar N, Lam TT, Ou J, Zhu LJ, Green MR, et al. MELK promotes melanoma growth by stimulating the NF-κB pathway. Cell Rep. 2017;21:2829–41.
Dou QP, Zonder JA. Overview of proteasome inhibitor-based anti-cancer therapies: perspective on bortezomib and second generation proteasome inhibitors versus future generation inhibitors of ubiquitin-proteasome system. Curr Cancer Drug Targets. 2014;14:517–36.
Rico M, Baglioni M, Bondarenko M, Laluce NC, Rozados V, André N, et al. Metformin and propranolol combination prevents cancer progression and metastasis in different breast cancer models. Oncotarget. 2017;8:2874–89.
Ford SJ, Obeidy P, Lovejoy DB, Bedford M, Nichols L, Chadwick C, et al. Deferasirox (ICL670A) effectively inhibits oesophageal cancer growth in vitro and in vivo. Br J Pharmacol. 2013;168:1316–28.
Pullarkat V, Meng Z, Donohue C, Yamamoto VN, Tomassetti S, Bhatia R, et al. Iron chelators induce autophagic cell death in multiple myeloma cells. Leuk Res. 2014;38:988–96.
Pascale F, Bedouet L, Baylatry M, Namur J, Laurent A. Comparative chemosensitivity of VX2 and HCC cell lines to drugs used in TACE. Anticancer Res. 2015;35:6497–503.
Funding
Not applicable.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Ethical approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
About this article
Cite this article
Nair, G., Hema Sree, G.N.S., Saraswathy, G.R. et al. Application of comprehensive bioinformatics approaches to reconnoiter crucial genes and pathways underpinning hepatocellular carcinoma: a drug repurposing endeavor. Med Oncol 38, 145 (2021). https://doi.org/10.1007/s12032-021-01576-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-021-01576-w